EMICSS
EMICSS stands for EMDB Integration with Complexes, Structures and Sequences. This service provides cross-referenced annotations for all EMDB entries, including both entry-level annotations (e.g., publication, corresponding PDB and EMPIAR entries, etc.) and sample-level (e.g., UniProt identifiers, AlphaFold DB models, etc.) annotations. The information from EMICSS is used on the EMDB website to provide relevant links and annotation for individual entries and sample components. The search system also takes advantage of this data to enable advanced queries not otherwise possible.
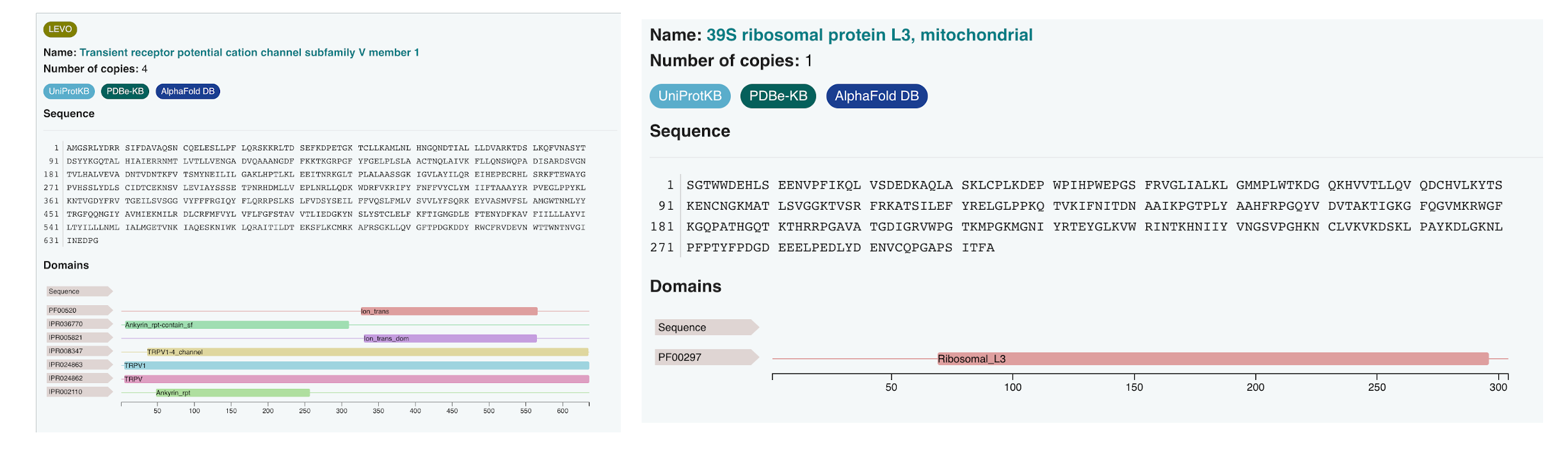
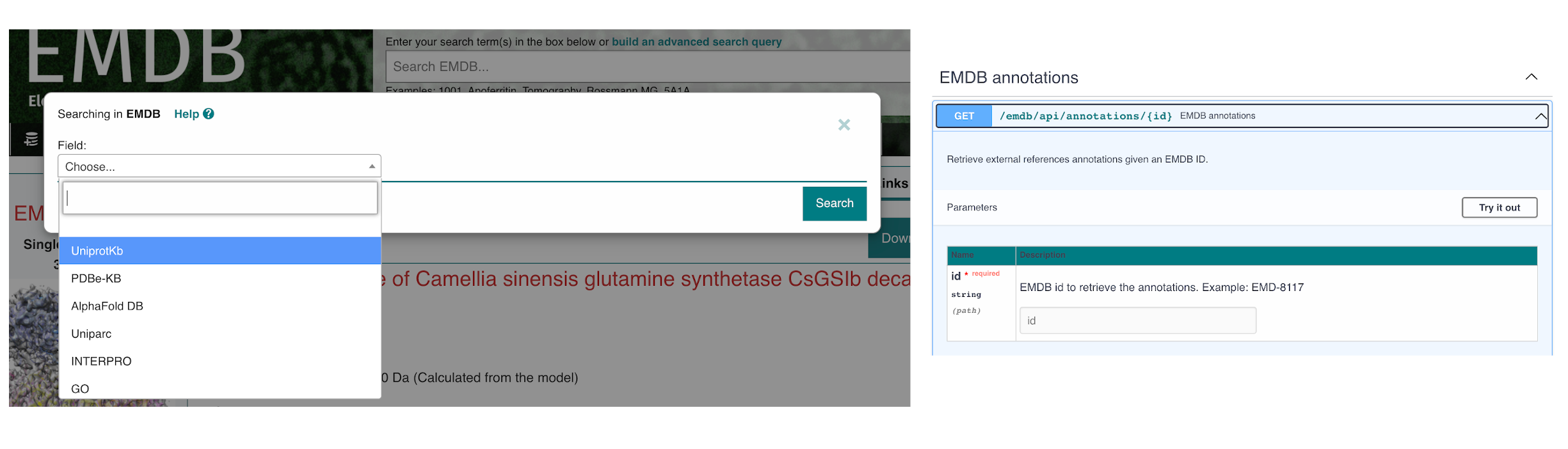
EMICSS annotations are produced by an automated pipeline that produces XML files (one for every EMDB entry) and TSV files (one for every resource that is used by EMICSS). EMICSS annotations are generated with the weekly release of new archive entries, and the whole archive is updated on a monthly basis. The collection date of the annotation is captured and stored by the EMICSS system in the EMICSS XML/TSV files. The structure and content of the EMICSS XML files is described by an XSD data model (available from https://ftp.ebi.ac.uk/pub/databases/em_ebi/emdb_related/emicss/emicss-schema/current/emdb_emicss.xsd). The EMICSS XML file for an EMDB entry can be accessed through the corresponding entry page on the EMDB website, either via the 'Links' tab or using the ‘Download’ dropdown menu. Sample-level annotations are also shown in the sample tab of every EMDB entry page. The EMICSS FTP area contains both the XML and TSV files. EMICSS can also be accessed through the EMDB annotation API.
If you use EMICSS, please cite: A. Duraisamy, N. Fonseca, G. J. Kleywegt and A. Patwardhan, "EMICSS: Value-added annotations for EMDB entries", manuscript in preparation (2023).
All EMICSS data is free to download and use; we encourage resources that index or expose EMDB data to make use of them. If you have any questions or suggestions regarding EMICSS, please contact the EMDB helpdesk.
The table below shows the provenance of the various information items collected by EMICSS. Note that much of the data about individual molecules that have been modelled into the EM volume are obtained from the PDBe/UniProt SIFTS resource.
| Information | Source |
|---|---|
| EMPIAR id(s) | EMPIAR |
| PDB id(s) | EMDB |
| Sample Weight | EMDB/PDBe |
| DOI | EMDB/Europe PMC |
| PubMed id | EMDB/Europe PMC |
| PMC id | EMDB/Europe PMC |
| ISSN | EMDB |
| ORCID identifiers | Europe PMC |
| UniProt id(s) | EMDB/UniProt |
| PDBe-KB links | UniProt |
| Complex Portal id(s) | Complex Portal/UniProt |
| Gene Ontology terms | PDBe/UniProt/QuickGO/EMDB |
| InterPro mappings | PDBe/UniProt/InterPro/EMDB |
| Pfam domains | PDBe/UniProt/Pfam/EMDB |
| CATH domains | PDBe/UniProt |
| SCOP domains | PDBe/UniProt |
| SCOP2 domains | PDBe/UniProt |
| ChEMBL id(s) | PDBe CCD |
| ChEBI id(s) | PDBe CCD |
| DrugBank id(s) | PDBe CCD |
| AlphaFold DB links | UniProt/AlphaFold DB |
Downloads
| Directory | Description |
| https://ftp.ebi.ac.uk/pub/databases/em_ebi/emdb_related/emicss/ | EMICSS FTP area |
| https://ftp.ebi.ac.uk/pub/databases/em_ebi/emdb_related/emicss/emicss-schema/current/ | EMICSS data model |
| https://ftp.ebi.ac.uk/pub/databases/em_ebi/emdb_related/emicss/entries/ | EMICSS XML files for EMDB entries |
| https://ftp.ebi.ac.uk/pub/databases/em_ebi/emdb_related/emicss/entries.tar.gz | Whole-archive EMICSS XML files for EMDB entries compressed (tar.gz) |
| https://ftp.ebi.ac.uk/pub/databases/em_ebi/emdb_related/emicss/resources/ | EMICSS TSV files for external resources |
| https://ftp.ebi.ac.uk/pub/databases/em_ebi/emdb_related/emicss/resources.tar.gz | Whole-archive EMICSS TSV files for external resources compressed (tar.gz) |
Organisation of the per-entry information
For EMDB entries with a 4-digit identifier (e.g., EMD-8117), the directories are grouped by the first two digits (in the example, /81/). The next level of the directory tree then consists of the entire 4-digit code. In this example, the full directory path will thus be: https://ftp.ebi.ac.uk/pub/databases/em_ebi/emdb_related/emicss/entries/81/8117/. The file name in that directory will be emd_8117_emicss.xml
For EMDB entries with a 5-digit identifier (e.g., EMD-28754), there is an additional level. The first level will consist of the first two digits (/28/), the second level of the third digit (/7/) and the lowest level of the entire 5-digit code. In this example, the full directory path will thus be: https://ftp.ebi.ac.uk/pub/databases/em_ebi/emdb_related/emicss/entries/28/7/28754/. The filename in that directory will be emd_28754_emicss.xml
Note: if you want to download the entire set of EMICSS XML files for all entries it will be significantly faster to download the tarball (see table above). Note that EMICSS mappings are transient. We therefore regenerate all EMICSS files every week, thus every file will differ from the same file a week earlier (even if the mapping may not have changed).
Statistics
The graph directly below provides information regarding the EMICSS coverage for the most recent release. Each bar indicates how many EMDB entries have one or more references to that resource (hover over a bar to see the exact count). The bottom graph tracks the development of this coverage over time.
Funding
The work on EMDB and EMICSS is funded by the Wellcome Trust (grant 212977/Z/18/Z) and EMBL-EBI.