 |
PDBsum entry 1ig9

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Transferase/DNA
|
PDB id
|
|

|

|
1ig9
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
 |
Contents |
 |
|
|

|

|
|
|
|
|
|
|
|
|
|
|
|
* Residue conservation analysis
|
|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 1:
|
 |
E.C.2.7.7.7
- DNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
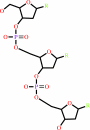 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 2:
|
 |
E.C.3.1.11.-
- ?????
|
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
DOI no:
|
Cell
105:657-667
(2001)
|
|
PubMed id:
|
|
|
|

|
| |
|
Structure of the replicating complex of a pol alpha family DNA polymerase.
|
|
M.C.Franklin,
J.Wang,
T.A.Steitz.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
We describe the 2.6 A resolution crystal structure of RB69 DNA polymerase with
primer-template DNA and dTTP, capturing the step just before primer extension.
This ternary complex structure in the human DNA polymerase alpha family shows a
60 degrees rotation of the fingers domain relative to the apo-protein structure,
similar to the fingers movement in pol I family polymerases. Minor groove
interactions near the primer 3' terminus suggest a common fidelity mechanism for
pol I and pol alpha family polymerases. The duplex product DNA orientation
differs by 40 degrees between the polymerizing mode and editing mode structures.
The role of the thumb in this DNA motion provides a model for editing in the pol
alpha family.

|

|
|
|

|
| |
Selected figure(s)
|
|

|
| |
 |
 |

|
 |

|
 |
Figure 2.
Figure 2. The Polymerase Active Site(A) The backbones of
the polymerase palm and fingers domains are shown as magenta and
blue ribbons. The carbon atoms of the polymerase side chains
shown in stick form are white, while those of the DNA template
are dark gray, and those of the primer DNA and dTTP are gold.
The bound calcium atoms are shown as light blue mesh spheres,
labeled A and B according to their catalytic function. Hydrogen
bonds are shown as green lines, while interactions with the
metal ions are blue lines. This view looks down the length of
the fingers, roughly 180° away from the orientation of
Figure 1A.(B) The KKRY motif (residues 705–708) interacts with
the primer-template duplex next to the polymerase active site.
Residues 703–708 are shown; Thr 703 and Gly 704 are not
conserved in the pol α family. Asp 621 from Figure 2A is shown
as an isolated side chain. Only the last two bases of the primer
strand are shown for clarity. The orientation of this figure can
be related to that of Figure 2A by comparing the primer-template
DNA
|
 |
Figure 4.
Figure 4. A Tunnel into the Polymerase Active SiteThe
molecular surface of RB69 pol in its polymerizing mode is
colored by domain according to the scheme of Figure 1A. The dTTP
and associated metal ions are at the center of the image, with
the primer-template DNA, shown semitransparently behind the
surface, extending off to the left. The terminal amine of Lys
560 protrudes into the tunnel, as indicated. This view is
roughly the same as that of Figure 2 and Figure 3
|
 |
|
|

|
| |
The above figures are
reprinted
by permission from Cell Press:
Cell
(2001,
105,
657-667)
copyright 2001.
|
|
| |
Figures were
selected
by an automated process.
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Literature references that cite this PDB file's key reference
|
|
 |
| |
PubMed id
|
 |
Reference
|
 |
|
|
|
 |
T.Nakamura,
Y.Zhao,
Y.Yamagata,
Y.J.Hua,
and
W.Yang
(2012).
Watching DNA polymerase η make a phosphodiester bond.
|
| |
Nature,
487,
196-201.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
K.Mayanagi,
S.Kiyonari,
H.Nishida,
M.Saito,
D.Kohda,
Y.Ishino,
T.Shirai,
and
K.Morikawa
(2011).
Architecture of the DNA polymerase B-proliferating cell nuclear antigen (PCNA)-DNA ternary complex.
|
| |
Proc Natl Acad Sci U S A,
108,
1845-1849.
|
 |
|
|
|
|
 |
S.K.Jozwiakowski,
and
B.A.Connolly
(2011).
A modified family-B archaeal DNA polymerase with reverse transcriptase activity.
|
| |
Chembiochem,
12,
35-37.
|
 |
|
|
|
|
 |
Y.W.Yin
(2011).
Structural insight on processivity, human disease and antiviral drug toxicity.
|
| |
Curr Opin Struct Biol,
21,
83-91.
|
 |
|
|
|
|
 |
A.A.Golosov,
J.J.Warren,
L.S.Beese,
and
M.Karplus
(2010).
The mechanism of the translocation step in DNA replication by DNA polymerase I: a computer simulation analysis.
|
| |
Structure,
18,
83-93.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
E.Johansson,
and
S.A.Macneill
(2010).
The eukaryotic replicative DNA polymerases take shape.
|
| |
Trends Biochem Sci,
35,
339-347.
|
 |
|
|
|
|
 |
G.Stengel,
M.Urban,
B.W.Purse,
and
R.D.Kuchta
(2010).
Incorporation of the fluorescent ribonucleotide analogue tCTP by T7 RNA polymerase.
|
| |
Anal Chem,
82,
1082-1089.
|
 |
|
|
|
|
 |
H.Zhang,
and
F.P.Guengerich
(2010).
Effect of N2-guanyl modifications on early steps in catalysis of polymerization by Sulfolobus solfataricus P2 DNA polymerase Dpo4 T239W.
|
| |
J Mol Biol,
395,
1007-1018.
|
 |
|
|
|
|
 |
J.Beckman,
M.Wang,
G.Blaha,
J.Wang,
and
W.H.Konigsberg
(2010).
Substitution of Ala for Tyr567 in RB69 DNA polymerase allows dAMP to be inserted opposite 7,8-dihydro-8-oxoguanine .
|
| |
Biochemistry,
49,
4116-4125.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
J.D.Pata
(2010).
Structural diversity of the Y-family DNA polymerases.
|
| |
Biochim Biophys Acta,
1804,
1124-1135.
|
 |
|
|
|
|
 |
M.E.Arana,
and
T.A.Kunkel
(2010).
Mutator phenotypes due to DNA replication infidelity.
|
| |
Semin Cancer Biol,
20,
304-311.
|
 |
|
|
|
|
 |
M.Hogg,
J.Rudnicki,
J.Midkiff,
L.Reha-Krantz,
S.Doublié,
and
S.S.Wallace
(2010).
Kinetics of mismatch formation opposite lesions by the replicative DNA polymerase from bacteriophage RB69.
|
| |
Biochemistry,
49,
2317-2325.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
M.T.Washington,
K.D.Carlson,
B.D.Freudenthal,
and
J.M.Pryor
(2010).
Variations on a theme: eukaryotic Y-family DNA polymerases.
|
| |
Biochim Biophys Acta,
1804,
1113-1123.
|
 |
|
|
|
|
 |
M.Uzan,
and
E.S.Miller
(2010).
Post-transcriptional control by bacteriophage T4: mRNA decay and inhibition of translation initiation.
|
| |
Virol J,
7,
360.
|
 |
|
|
|
|
 |
N.Ramsay,
A.S.Jemth,
A.Brown,
N.Crampton,
P.Dear,
and
P.Holliger
(2010).
CyDNA: synthesis and replication of highly Cy-Dye substituted DNA by an evolved polymerase.
|
| |
J Am Chem Soc,
132,
5096-5104.
|
 |
|
|
|
|
 |
P.Aller,
Y.Ye,
S.S.Wallace,
C.J.Burrows,
and
S.Doublié
(2010).
Crystal structure of a replicative DNA polymerase bound to the oxidized guanine lesion guanidinohydantoin.
|
| |
Biochemistry,
49,
2502-2509.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
S.K.Perumal,
H.Yue,
Z.Hu,
M.M.Spiering,
and
S.J.Benkovic
(2010).
Single-molecule studies of DNA replisome function.
|
| |
Biochim Biophys Acta,
1804,
1094-1112.
|
 |
|
|
|
|
 |
T.C.Mueser,
J.M.Hinerman,
J.M.Devos,
R.A.Boyer,
and
K.J.Williams
(2010).
Structural analysis of bacteriophage T4 DNA replication: a review in the Virology Journal series on bacteriophage T4 and its relatives.
|
| |
Virol J,
7,
359.
|
 |
|
|
|
|
 |
T.Ogino,
K.Sato,
and
A.Matsuda
(2010).
Incorporation of 2'-deoxy-2'-isonucleoside 5'-triphosphates (iNTPs) into DNA by A- and B-family DNA polymerases with different recognition mechanisms.
|
| |
Chembiochem,
11,
2597-2605.
|
 |
|
|
|
|
 |
X.Meng,
Y.Zhou,
E.Y.Lee,
M.Y.Lee,
and
D.N.Frick
(2010).
The p12 subunit of human polymerase delta modulates the rate and fidelity of DNA synthesis.
|
| |
Biochemistry,
49,
3545-3554.
|
 |
|
|
|
|
 |
Y.Santoso,
C.M.Joyce,
O.Potapova,
L.Le Reste,
J.Hohlbein,
J.P.Torella,
N.D.Grindley,
and
A.N.Kapanidis
(2010).
Conformational transitions in DNA polymerase I revealed by single-molecule FRET.
|
| |
Proc Natl Acad Sci U S A,
107,
715-720.
|
 |
|
|
|
|
 |
Y.Santoso,
J.P.Torella,
and
A.N.Kapanidis
(2010).
Characterizing single-molecule FRET dynamics with probability distribution analysis.
|
| |
Chemphyschem,
11,
2209-2219.
|
 |
|
|
|
|
 |
Z.Zhuang,
and
Y.Ai
(2010).
Processivity factor of DNA polymerase and its expanding role in normal and translesion DNA synthesis.
|
| |
Biochim Biophys Acta,
1804,
1081-1093.
|
 |
|
|
|
|
 |
C.Castro,
E.D.Smidansky,
J.J.Arnold,
K.R.Maksimchuk,
I.Moustafa,
A.Uchida,
M.Götte,
W.Konigsberg,
and
C.E.Cameron
(2009).
Nucleic acid polymerases use a general acid for nucleotidyl transfer.
|
| |
Nat Struct Mol Biol,
16,
212-218.
|
 |
|
|
|
|
 |
C.Xu,
B.A.Maxwell,
J.A.Brown,
L.Zhang,
and
Z.Suo
(2009).
Global conformational dynamics of a Y-family DNA polymerase during catalysis.
|
| |
PLoS Biol,
7,
e1000225.
|
 |
|
|
|
|
 |
E.P.Tchesnokov,
A.Obikhod,
R.F.Schinazi,
and
M.Götte
(2009).
Engineering of a chimeric RB69 DNA polymerase sensitive to drugs targeting the cytomegalovirus enzyme.
|
| |
J Biol Chem,
284,
26439-26446.
|
 |
|
|
|
|
 |
F.Wang,
and
W.Yang
(2009).
Structural insight into translesion synthesis by DNA Pol II.
|
| |
Cell,
139,
1279-1289.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
H.J.Russell,
T.T.Richardson,
K.Emptage,
and
B.A.Connolly
(2009).
The 3'-5' proofreading exonuclease of archaeal family-B DNA polymerase hinders the copying of template strand deaminated bases.
|
| |
Nucleic Acids Res,
37,
7603-7611.
|
 |
|
|
|
|
 |
H.Nishida,
K.Mayanagi,
S.Kiyonari,
Y.Sato,
T.Oyama,
Y.Ishino,
and
K.Morikawa
(2009).
Structural determinant for switching between the polymerase and exonuclease modes in the PCNA-replicative DNA polymerase complex.
|
| |
Proc Natl Acad Sci U S A,
106,
20693-20698.
|
 |
|
|
|
|
 |
H.Zhang,
J.Beckman,
J.Wang,
and
W.Konigsberg
(2009).
RB69 DNA polymerase mutants with expanded nascent base-pair-binding pockets are highly efficient but have reduced base selectivity.
|
| |
Biochemistry,
48,
6940-6950.
|
 |
|
|
|
|
 |
I.Rodríguez,
J.M.Lázaro,
M.Salas,
and
M.de Vega
(2009).
Involvement of the TPR2 subdomain movement in the activities of phi29 DNA polymerase.
|
| |
Nucleic Acids Res,
37,
193-203.
|
 |
|
|
|
|
 |
J.E.Stone,
G.E.Kissling,
S.A.Lujan,
I.B.Rogozin,
C.M.Stith,
P.M.Burgers,
and
T.A.Kunkel
(2009).
Low-fidelity DNA synthesis by the L979F mutator derivative of Saccharomyces cerevisiae DNA polymerase zeta.
|
| |
Nucleic Acids Res,
37,
3774-3787.
|
 |
|
|
|
|
 |
J.L.Baltz,
D.J.Filman,
M.Ciustea,
J.E.Silverman,
C.L.Lautenschlager,
D.M.Coen,
R.P.Ricciardi,
and
J.M.Hogle
(2009).
The crystal structure of PF-8, the DNA polymerase accessory subunit from Kaposi's sarcoma-associated herpesvirus.
|
| |
J Virol,
83,
12215-12228.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
K.Datta,
N.P.Johnson,
V.J.Licata,
and
P.H.von Hippel
(2009).
Local Conformations and Competitive Binding Affinities of Single- and Double-stranded Primer-Template DNA at the Polymerization and Editing Active Sites of DNA Polymerases.
|
| |
J Biol Chem,
284,
17180-17193.
|
 |
|
|
|
|
 |
M.G.Pence,
P.Blans,
C.N.Zink,
T.Hollis,
J.C.Fishbein,
and
F.W.Perrino
(2009).
Lesion bypass of N2-ethylguanine by human DNA polymerase iota.
|
| |
J Biol Chem,
284,
1732-1740.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
M.K.Swan,
R.E.Johnson,
L.Prakash,
S.Prakash,
and
A.K.Aggarwal
(2009).
Structural basis of high-fidelity DNA synthesis by yeast DNA polymerase delta.
|
| |
Nat Struct Mol Biol,
16,
979-986.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
M.Renders,
M.Abramov,
M.Froeyen,
and
P.Herdewijn
(2009).
Polymerase-catalysed incorporation of glucose nucleotides into a DNA duplex.
|
| |
Chemistry,
15,
5463-5470.
|
 |
|
|
|
|
 |
M.Wang,
H.R.Lee,
and
W.Konigsberg
(2009).
Effect of A and B metal ion site occupancy on conformational changes in an RB69 DNA polymerase ternary complex.
|
| |
Biochemistry,
48,
2075-2086.
|
 |
|
|
|
|
 |
N.A.Cavanaugh,
M.Urban,
J.Beckman,
T.E.Spratt,
and
R.D.Kuchta
(2009).
Identifying the features of purine dNTPs that allow accurate and efficient DNA replication by herpes simplex virus I DNA polymerase.
|
| |
Biochemistry,
48,
3554-3564.
|
 |
|
|
|
|
 |
P.B.Balbo,
and
A.Bohm
(2009).
Proton transfer in the mechanism of polyadenylate polymerase.
|
| |
Biochem J,
420,
229-238.
|
 |
|
|
|
|
 |
S.M.Hamdan,
and
C.C.Richardson
(2009).
Motors, switches, and contacts in the replisome.
|
| |
Annu Rev Biochem,
78,
205-243.
|
 |
|
|
|
|
 |
S.Ogata,
M.Takahashi,
N.Minakawa,
and
A.Matsuda
(2009).
Unnatural imidazopyridopyrimidine:naphthyridine base pairs: selective incorporation and extension reaction by Deep Vent (exo- ) DNA polymerase.
|
| |
Nucleic Acids Res,
37,
5602-5609.
|
 |
|
|
|
|
 |
V.Borsenberger,
M.Kukwikila,
and
S.Howorka
(2009).
Synthesis and enzymatic incorporation of modified deoxyuridine triphosphates.
|
| |
Org Biomol Chem,
7,
3826-3835.
|
 |
|
|
|
|
 |
A.G.Baranovskiy,
N.D.Babayeva,
V.G.Liston,
I.B.Rogozin,
E.V.Koonin,
Y.I.Pavlov,
D.G.Vassylyev,
and
T.H.Tahirov
(2008).
X-ray structure of the complex of regulatory subunits of human DNA polymerase delta.
|
| |
Cell Cycle,
7,
3026-3036.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
A.Sheriff,
E.Motea,
I.Lee,
and
A.J.Berdis
(2008).
Mechanism and dynamics of translesion DNA synthesis catalyzed by the Escherichia coli Klenow fragment.
|
| |
Biochemistry,
47,
8527-8537.
|
 |
|
|
|
|
 |
C.A.Howell,
C.M.Kondratick,
and
M.T.Washington
(2008).
Substitution of a residue contacting the triphosphate moiety of the incoming nucleotide increases the fidelity of yeast DNA polymerase zeta.
|
| |
Nucleic Acids Res,
36,
1731-1740.
|
 |
|
|
|
|
 |
D.B.Gammon,
R.Snoeck,
P.Fiten,
M.Krecmerová,
A.Holý,
E.De Clercq,
G.Opdenakker,
D.H.Evans,
and
G.Andrei
(2008).
Mechanism of antiviral drug resistance of vaccinia virus: identification of residues in the viral DNA polymerase conferring differential resistance to antipoxvirus drugs.
|
| |
J Virol,
82,
12520-12534.
|
 |
|
|
|
|
 |
G.T.Hwang,
and
F.E.Romesberg
(2008).
Unnatural substrate repertoire of A, B, and X family DNA polymerases.
|
| |
J Am Chem Soc,
130,
14872-14882.
|
 |
|
|
|
|
 |
J.Cramer,
G.Rangam,
A.Marx,
and
T.Restle
(2008).
Varied active-site constraints in the klenow fragment of E. coli DNA polymerase I and the lesion-bypass Dbh DNA polymerase.
|
| |
Chembiochem,
9,
1243-1250.
|
 |
|
|
|
|
 |
K.F.Bryant,
and
D.M.Coen
(2008).
Inhibition of translation by a short element in the 5' leader of the herpes simplex virus 1 DNA polymerase transcript.
|
| |
J Virol,
82,
77-85.
|
 |
|
|
|
|
 |
K.K.Ng,
J.J.Arnold,
and
C.E.Cameron
(2008).
Structure-function relationships among RNA-dependent RNA polymerases.
|
| |
Curr Top Microbiol Immunol,
320,
137-156.
|
 |
|
|
|
|
 |
K.Ozawa,
S.Jergic,
A.Y.Park,
N.E.Dixon,
and
G.Otting
(2008).
The proofreading exonuclease subunit epsilon of Escherichia coli DNA polymerase III is tethered to the polymerase subunit alpha via a flexible linker.
|
| |
Nucleic Acids Res,
36,
5074-5082.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
L.A.Loeb,
and
R.J.Monnat
(2008).
DNA polymerases and human disease.
|
| |
Nat Rev Genet,
9,
594-604.
|
 |
|
|
|
|
 |
M.Renders,
R.Lievrouw,
M.Krecmerová,
A.Holý,
and
P.Herdewijn
(2008).
Enzymatic polymerization of phosphonate nucleosides.
|
| |
Chembiochem,
9,
2883-2888.
|
 |
|
|
|
|
 |
P.Kukreti,
K.Singh,
A.Ketkar,
and
M.J.Modak
(2008).
Identification of a new motif required for the 3'-5' exonuclease activity of Escherichia coli DNA polymerase I (Klenow fragment): the RRRY motif is necessary for the binding of single-stranded DNA substrate and the template strand of the mismatched duplex.
|
| |
J Biol Chem,
283,
17979-17990.
|
 |
|
|
|
|
 |
R.A.Wing,
S.Bailey,
and
T.A.Steitz
(2008).
Insights into the replisome from the structure of a ternary complex of the DNA polymerase III alpha-subunit.
|
| |
J Mol Biol,
382,
859-869.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
S.Broyde,
L.Wang,
O.Rechkoblit,
N.E.Geacintov,
and
D.J.Patel
(2008).
Lesion processing: high-fidelity versus lesion-bypass DNA polymerases.
|
| |
Trends Biochem Sci,
33,
209-219.
|
 |
|
|
|
|
 |
S.J.Garforth,
M.A.Parniak,
and
V.R.Prasad
(2008).
Utilization of a deoxynucleoside diphosphate substrate by HIV reverse transcriptase.
|
| |
PLoS ONE,
3,
e2074.
|
 |
|
|
|
|
 |
T.A.Kunkel,
and
P.M.Burgers
(2008).
Dividing the workload at a eukaryotic replication fork.
|
| |
Trends Cell Biol,
18,
521-527.
|
 |
|
|
|
|
 |
W.J.Allen,
P.J.Rothwell,
and
G.Waksman
(2008).
An intramolecular FRET system monitors fingers subdomain opening in Klentaq1.
|
| |
Protein Sci,
17,
401-408.
|
 |
|
|
|
|
 |
W.Tian,
Y.T.Hwang,
and
C.B.Hwang
(2008).
The enhanced DNA replication fidelity of a mutant herpes simplex virus type 1 DNA polymerase is mediated by an improved nucleotide selectivity and reduced mismatch extension ability.
|
| |
J Virol,
82,
8937-8941.
|
 |
|
|
|
|
 |
X.Zhong,
L.C.Pedersen,
and
T.A.Kunkel
(2008).
Characterization of a replicative DNA polymerase mutant with reduced fidelity and increased translesion synthesis capacity.
|
| |
Nucleic Acids Res,
36,
3892-3904.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
A.J.Berman,
S.Kamtekar,
J.L.Goodman,
J.M.Lázaro,
M.de Vega,
L.Blanco,
M.Salas,
and
T.A.Steitz
(2007).
Structures of phi29 DNA polymerase complexed with substrate: the mechanism of translocation in B-family polymerases.
|
| |
EMBO J,
26,
3494-3505.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
A.Jacewicz,
K.Makiela,
A.Kierzek,
J.W.Drake,
and
A.Bebenek
(2007).
The roles of Tyr391 and Tyr619 in RB69 DNA polymerase replication fidelity.
|
| |
J Mol Biol,
368,
18-29.
|
 |
|
|
|
|
 |
A.N.Sakamoto,
J.E.Stone,
G.E.Kissling,
S.D.McCulloch,
Y.I.Pavlov,
and
T.A.Kunkel
(2007).
Mutator alleles of yeast DNA polymerase zeta.
|
| |
DNA Repair (Amst),
6,
1829-1838.
|
 |
|
|
|
|
 |
A.P.Silverman,
Q.Jiang,
M.F.Goodman,
and
E.T.Kool
(2007).
Steric and electrostatic effects in DNA synthesis by the SOS-induced DNA polymerases II and IV of Escherichia coli.
|
| |
Biochemistry,
46,
13874-13881.
|
 |
|
|
|
|
 |
A.Wood,
P.Garg,
and
P.M.Burgers
(2007).
A ubiquitin-binding motif in the translesion DNA polymerase Rev1 mediates its essential functional interaction with ubiquitinated proliferating cell nuclear antigen in response to DNA damage.
|
| |
J Biol Chem,
282,
20256-20263.
|
 |
|
|
|
|
 |
C.Castro,
E.Smidansky,
K.R.Maksimchuk,
J.J.Arnold,
V.S.Korneeva,
M.Götte,
W.Konigsberg,
and
C.E.Cameron
(2007).
Two proton transfers in the transition state for nucleotidyl transfer catalyzed by RNA- and DNA-dependent RNA and DNA polymerases.
|
| |
Proc Natl Acad Sci U S A,
104,
4267-4272.
|
 |
|
|
|
|
 |
E.Fidalgo da Silva,
and
L.J.Reha-Krantz
(2007).
DNA polymerase proofreading: active site switching catalyzed by the bacteriophage T4 DNA polymerase.
|
| |
Nucleic Acids Res,
35,
5452-5463.
|
 |
|
|
|
|
 |
H.Choo,
J.R.Beadle,
Y.Chong,
J.Trahan,
and
K.Y.Hostetler
(2007).
Synthesis of the 5-phosphono-pent-2-en-1-yl nucleosides: a new class of antiviral acyclic nucleoside phosphonates.
|
| |
Bioorg Med Chem,
15,
1771-1779.
|
 |
|
|
|
|
 |
H.Zhang,
W.Cao,
E.Zakharova,
W.Konigsberg,
and
E.M.De La Cruz
(2007).
Fluorescence of 2-aminopurine reveals rapid conformational changes in the RB69 DNA polymerase-primer/template complexes upon binding and incorporation of matched deoxynucleoside triphosphates.
|
| |
Nucleic Acids Res,
35,
6052-6062.
|
 |
|
|
|
|
 |
J.Beckman,
K.Kincaid,
M.Hocek,
T.Spratt,
J.Engels,
R.Cosstick,
and
R.D.Kuchta
(2007).
Human DNA polymerase alpha uses a combination of positive and negative selectivity to polymerize purine dNTPs with high fidelity.
|
| |
Biochemistry,
46,
448-460.
|
 |
|
|
|
|
 |
J.Bischerour,
and
R.Chalmers
(2007).
Base-flipping dynamics in a DNA hairpin processing reaction.
|
| |
Nucleic Acids Res,
35,
2584-2595.
|
 |
|
|
|
|
 |
K.Singh,
A.Srivastava,
S.S.Patel,
and
M.J.Modak
(2007).
Participation of the fingers subdomain of Escherichia coli DNA polymerase I in the strand displacement synthesis of DNA.
|
| |
J Biol Chem,
282,
10594-10604.
|
 |
|
|
|
|
 |
L.B.Goodman,
A.Loregian,
G.A.Perkins,
J.Nugent,
E.L.Buckles,
B.Mercorelli,
J.H.Kydd,
G.Palù,
K.C.Smith,
N.Osterrieder,
and
N.Davis-Poynter
(2007).
A point mutation in a herpesvirus polymerase determines neuropathogenicity.
|
| |
PLoS Pathog,
3,
e160.
|
 |
|
|
|
|
 |
M.D.Hamilton,
A.A.Nuara,
D.B.Gammon,
R.M.Buller,
and
D.H.Evans
(2007).
Duplex strand joining reactions catalyzed by vaccinia virus DNA polymerase.
|
| |
Nucleic Acids Res,
35,
143-151.
|
 |
|
|
|
|
 |
M.Hogg,
P.Aller,
W.Konigsberg,
S.S.Wallace,
and
S.Doublié
(2007).
Structural and biochemical investigation of the role in proofreading of a beta hairpin loop found in the exonuclease domain of a replicative DNA polymerase of the B family.
|
| |
J Biol Chem,
282,
1432-1444.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
M.Strerath,
C.Gloeckner,
D.Liu,
A.Schnur,
and
A.Marx
(2007).
Directed DNA polymerase evolution: effects of mutations in motif C on the mismatch-extension selectivity of thermus aquaticus DNA polymerase.
|
| |
Chembiochem,
8,
395-401.
|
 |
|
|
|
|
 |
M.de Vega,
and
M.Salas
(2007).
A highly conserved Tyrosine residue of family B DNA polymerases contributes to dictate translesion synthesis past 8-oxo-7,8-dihydro-2'-deoxyguanosine.
|
| |
Nucleic Acids Res,
35,
5096-5107.
|
 |
|
|
|
|
 |
N.Z.Rudinger,
R.Kranaster,
and
A.Marx
(2007).
Hydrophobic amino acid and single-atom substitutions increase DNA polymerase selectivity.
|
| |
Chem Biol,
14,
185-194.
|
 |
|
|
|
|
 |
P.Aller,
M.A.Rould,
M.Hogg,
S.S.Wallace,
and
S.Doublié
(2007).
A structural rationale for stalling of a replicative DNA polymerase at the most common oxidative thymine lesion, thymine glycol.
|
| |
Proc Natl Acad Sci U S A,
104,
814-818.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
P.R.Meyer,
W.Rutvisuttinunt,
S.E.Matsuura,
A.G.So,
and
W.A.Scott
(2007).
Stable complexes formed by HIV-1 reverse transcriptase at distinct positions on the primer-template controlled by binding deoxynucleoside triphosphates or foscarnet.
|
| |
J Mol Biol,
369,
41-54.
|
 |
|
|
|
|
 |
R.A.Perlow-Poehnelt,
I.Likhterov,
L.Wang,
D.A.Scicchitano,
N.E.Geacintov,
and
S.Broyde
(2007).
Increased flexibility enhances misincorporation: temperature effects on nucleotide incorporation opposite a bulky carcinogen-DNA adduct by a Y-family DNA polymerase.
|
| |
J Biol Chem,
282,
1397-1408.
|
 |
|
|
|
|
 |
R.N.Venkatesan,
P.M.Treuting,
E.D.Fuller,
R.E.Goldsby,
T.H.Norwood,
T.A.Gooley,
W.C.Ladiges,
B.D.Preston,
and
L.A.Loeb
(2007).
Mutation at the polymerase active site of mouse DNA polymerase delta increases genomic instability and accelerates tumorigenesis.
|
| |
Mol Cell Biol,
27,
7669-7682.
|
 |
|
|
|
|
 |
S.A.Nick McElhinny,
C.M.Stith,
P.M.Burgers,
and
T.A.Kunkel
(2007).
Inefficient proofreading and biased error rates during inaccurate DNA synthesis by a mutant derivative of Saccharomyces cerevisiae DNA polymerase delta.
|
| |
J Biol Chem,
282,
2324-2332.
|
 |
|
|
|
|
 |
Z.F.Pursell,
I.Isoz,
E.B.Lundström,
E.Johansson,
and
T.A.Kunkel
(2007).
Regulation of B family DNA polymerase fidelity by a conserved active site residue: characterization of M644W, M644L and M644F mutants of yeast DNA polymerase epsilon.
|
| |
Nucleic Acids Res,
35,
3076-3086.
|
 |
|
|
|
|
 |
A.M.DeLucia,
S.Chaudhuri,
O.Potapova,
N.D.Grindley,
and
C.M.Joyce
(2006).
The properties of steric gate mutants reveal different constraints within the active sites of Y-family and A-family DNA polymerases.
|
| |
J Biol Chem,
281,
27286-27291.
|
 |
|
|
|
|
 |
A.Wieczorek,
and
C.S.McHenry
(2006).
The NH2-terminal php domain of the alpha subunit of the Escherichia coli replicase binds the epsilon proofreading subunit.
|
| |
J Biol Chem,
281,
12561-12567.
|
 |
|
|
|
|
 |
C.Hariharan,
L.B.Bloom,
S.A.Helquist,
E.T.Kool,
and
L.J.Reha-Krantz
(2006).
Dynamics of nucleotide incorporation: snapshots revealed by 2-aminopurine fluorescence studies.
|
| |
Biochemistry,
45,
2836-2844.
|
 |
|
|
|
|
 |
E.Longás,
M.de Vega,
J.M.Lázaro,
and
M.Salas
(2006).
Functional characterization of highly processive protein-primed DNA polymerases from phages Nf and GA-1, endowed with a potent strand displacement capacity.
|
| |
Nucleic Acids Res,
34,
6051-6063.
|
 |
|
|
|
|
 |
E.P.Tchesnokov,
C.Gilbert,
G.Boivin,
and
M.Götte
(2006).
Role of helix P of the human cytomegalovirus DNA polymerase in resistance and hypersusceptibility to the antiviral drug foscarnet.
|
| |
J Virol,
80,
1440-1450.
|
 |
|
|
|
|
 |
G.Andrei,
D.B.Gammon,
P.Fiten,
E.De Clercq,
G.Opdenakker,
R.Snoeck,
and
D.H.Evans
(2006).
Cidofovir resistance in vaccinia virus is linked to diminished virulence in mice.
|
| |
J Virol,
80,
9391-9401.
|
 |
|
|
|
|
 |
K.L.Damm,
and
H.A.Carlson
(2006).
Gaussian-weighted RMSD superposition of proteins: a structural comparison for flexible proteins and predicted protein structures.
|
| |
Biophys J,
90,
4558-4573.
|
 |
|
|
|
|
 |
L.Zhang,
O.Rechkoblit,
L.Wang,
D.J.Patel,
R.Shapiro,
and
S.Broyde
(2006).
Mutagenic nucleotide incorporation and hindered translocation by a food carcinogen C8-dG adduct in Sulfolobus solfataricus P2 DNA polymerase IV (Dpo4): modeling and dynamics studies.
|
| |
Nucleic Acids Res,
34,
3326-3337.
|
 |
|
|
|
|
 |
M.Garcia-Diaz,
K.Bebenek,
J.M.Krahn,
L.C.Pedersen,
and
T.A.Kunkel
(2006).
Structural analysis of strand misalignment during DNA synthesis by a human DNA polymerase.
|
| |
Cell,
124,
331-342.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
M.Garcia-Diaz,
and
T.A.Kunkel
(2006).
Mechanism of a genetic glissando: structural biology of indel mutations.
|
| |
Trends Biochem Sci,
31,
206-214.
|
 |
|
|
|
|
 |
M.H.Lamers,
R.E.Georgescu,
S.G.Lee,
M.O'Donnell,
and
J.Kuriyan
(2006).
Crystal structure of the catalytic alpha subunit of E. coli replicative DNA polymerase III.
|
| |
Cell,
126,
881-892.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
M.Hogg,
W.Cooper,
L.Reha-Krantz,
and
S.S.Wallace
(2006).
Kinetics of error generation in homologous B-family DNA polymerases.
|
| |
Nucleic Acids Res,
34,
2528-2535.
|
 |
|
|
|
|
 |
O.Potapova,
C.Chan,
A.M.DeLucia,
S.A.Helquist,
E.T.Kool,
N.D.Grindley,
and
C.M.Joyce
(2006).
DNA polymerase catalysis in the absence of Watson-Crick hydrogen bonds: analysis by single-turnover kinetics.
|
| |
Biochemistry,
45,
890-898.
|
 |
|
|
|
|
 |
P.Pérez-Arnaiz,
J.M.Lázaro,
M.Salas,
and
M.de Vega
(2006).
Involvement of phi29 DNA polymerase thumb subdomain in the proper coordination of synthesis and degradation during DNA replication.
|
| |
Nucleic Acids Res,
34,
3107-3115.
|
 |
|
|
|
|
 |
R.D.Smiley,
Z.Zhuang,
S.J.Benkovic,
and
G.G.Hammes
(2006).
Single-molecule investigation of the T4 bacteriophage DNA polymerase holoenzyme: multiple pathways of holoenzyme formation.
|
| |
Biochemistry,
45,
7990-7997.
|
 |
|
|
|
|
 |
R.N.Venkatesan,
J.J.Hsu,
N.A.Lawrence,
B.D.Preston,
and
L.A.Loeb
(2006).
Mutator phenotypes caused by substitution at a conserved motif A residue in eukaryotic DNA polymerase delta.
|
| |
J Biol Chem,
281,
4486-4494.
|
 |
|
|
|
|
 |
R.Shi,
A.Azzi,
C.Gilbert,
G.Boivin,
and
S.X.Lin
(2006).
Three-dimensional modeling of cytomegalovirus DNA polymerase and preliminary analysis of drug resistance.
|
| |
Proteins,
64,
301-307.
|
 |
|
|
|
|
 |
S.Kamtekar,
A.J.Berman,
J.Wang,
J.M.Lázaro,
M.de Vega,
L.Blanco,
M.Salas,
and
T.A.Steitz
(2006).
The phi29 DNA polymerase:protein-primer structure suggests a model for the initiation to elongation transition.
|
| |
EMBO J,
25,
1335-1343.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
S.Liu,
J.D.Knafels,
J.S.Chang,
G.A.Waszak,
E.T.Baldwin,
M.R.Deibel,
D.R.Thomsen,
F.L.Homa,
P.A.Wells,
M.C.Tory,
R.A.Poorman,
H.Gao,
X.Qiu,
and
A.P.Seddon
(2006).
Crystal structure of the herpes simplex virus 1 DNA polymerase.
|
| |
J Biol Chem,
281,
18193-18200.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
S.Sun,
L.Geng,
and
Y.Shamoo
(2006).
Structure and enzymatic properties of a chimeric bacteriophage RB69 DNA polymerase and single-stranded DNA binding protein with increased processivity.
|
| |
Proteins,
65,
231-238.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
T.A.Steitz
(2006).
Visualizing polynucleotide polymerase machines at work.
|
| |
EMBO J,
25,
3458-3468.
|
 |
|
|
|
|
 |
V.K.Batra,
W.A.Beard,
D.D.Shock,
J.M.Krahn,
L.C.Pedersen,
and
S.H.Wilson
(2006).
Magnesium-induced assembly of a complete DNA polymerase catalytic complex.
|
| |
Structure,
14,
757-766.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
A.Johnson,
and
M.O'Donnell
(2005).
Cellular DNA replicases: components and dynamics at the replication fork.
|
| |
Annu Rev Biochem,
74,
283-315.
|
 |
|
|
|
|
 |
A.Vaisman,
H.Ling,
R.Woodgate,
and
W.Yang
(2005).
Fidelity of Dpo4: effect of metal ions, nucleotide selection and pyrophosphorolysis.
|
| |
EMBO J,
24,
2957-2967.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
D.Summerer,
N.Z.Rudinger,
I.Detmer,
and
A.Marx
(2005).
Enhanced fidelity in mismatch extension by DNA polymerase through directed combinatorial enzyme design.
|
| |
Angew Chem Int Ed Engl,
44,
4712-4715.
|
 |
|
|
|
|
 |
H.D.Cho,
C.L.Verlinde,
and
A.M.Weiner
(2005).
Archaeal CCA-adding enzymes: central role of a highly conserved beta-turn motif in RNA polymerization without translocation.
|
| |
J Biol Chem,
280,
9555-9566.
|
 |
|
|
|
|
 |
I.Rodríguez,
J.M.Lázaro,
L.Blanco,
S.Kamtekar,
A.J.Berman,
J.Wang,
T.A.Steitz,
M.Salas,
and
M.de Vega
(2005).
A specific subdomain in phi29 DNA polymerase confers both processivity and strand-displacement capacity.
|
| |
Proc Natl Acad Sci U S A,
102,
6407-6412.
|
 |
|
|
|
|
 |
J.M.Fortune,
Y.I.Pavlov,
C.M.Welch,
E.Johansson,
P.M.Burgers,
and
T.A.Kunkel
(2005).
Saccharomyces cerevisiae DNA polymerase delta: high fidelity for base substitutions but lower fidelity for single- and multi-base deletions.
|
| |
J Biol Chem,
280,
29980-29987.
|
 |
|
|
|
|
 |
P.Garg,
and
P.M.Burgers
(2005).
Ubiquitinated proliferating cell nuclear antigen activates translesion DNA polymerases eta and REV1.
|
| |
Proc Natl Acad Sci U S A,
102,
18361-18366.
|
 |
|
|
|
|
 |
P.J.Rothwell,
V.Mitaksov,
and
G.Waksman
(2005).
Motions of the fingers subdomain of klentaq1 are fast and not rate limiting: implications for the molecular basis of fidelity in DNA polymerases.
|
| |
Mol Cell,
19,
345-355.
|
 |
|
|
|
|
 |
S.Prakash,
R.E.Johnson,
and
L.Prakash
(2005).
Eukaryotic translesion synthesis DNA polymerases: specificity of structure and function.
|
| |
Annu Rev Biochem,
74,
317-353.
|
 |
|
|
|
|
 |
V.K.Batra,
W.A.Beard,
D.D.Shock,
L.C.Pedersen,
and
S.H.Wilson
(2005).
Nucleotide-induced DNA polymerase active site motions accommodating a mutagenic DNA intermediate.
|
| |
Structure,
13,
1225-1233.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
V.Kempeneers,
M.Renders,
M.Froeyen,
and
P.Herdewijn
(2005).
Investigation of the DNA-dependent cyclohexenyl nucleic acid polymerization and the cyclohexenyl nucleic acid-dependent DNA polymerization.
|
| |
Nucleic Acids Res,
33,
3828-3836.
|
 |
|
|
|
|
 |
W.T.Wolfle,
M.T.Washington,
E.T.Kool,
T.E.Spratt,
S.A.Helquist,
L.Prakash,
and
S.Prakash
(2005).
Evidence for a Watson-Crick hydrogen bonding requirement in DNA synthesis by human DNA polymerase kappa.
|
| |
Mol Cell Biol,
25,
7137-7143.
|
 |
|
|
|
|
 |
Y.H.Jin,
P.Garg,
C.M.Stith,
H.Al-Refai,
J.F.Sterling,
L.J.Murray,
T.A.Kunkel,
M.A.Resnick,
P.M.Burgers,
and
D.A.Gordenin
(2005).
The multiple biological roles of the 3'-->5' exonuclease of Saccharomyces cerevisiae DNA polymerase delta require switching between the polymerase and exonuclease domains.
|
| |
Mol Cell Biol,
25,
461-471.
|
 |
|
|
|
|
 |
A.F.Gardner,
C.M.Joyce,
and
W.E.Jack
(2004).
Comparative kinetics of nucleotide analog incorporation by vent DNA polymerase.
|
| |
J Biol Chem,
279,
11834-11842.
|
 |
|
|
|
|
 |
A.Niimi,
S.Limsirichaikul,
S.Yoshida,
S.Iwai,
C.Masutani,
F.Hanaoka,
E.T.Kool,
Y.Nishiyama,
and
M.Suzuki
(2004).
Palm mutants in DNA polymerases alpha and eta alter DNA replication fidelity and translesion activity.
|
| |
Mol Cell Biol,
24,
2734-2746.
|
 |
|
|
|
|
 |
B.D.Biles,
and
B.A.Connolly
(2004).
Low-fidelity Pyrococcus furiosus DNA polymerase mutants useful in error-prone PCR.
|
| |
Nucleic Acids Res,
32,
e176.
|
 |
|
|
|
|
 |
C.L.Hendrickson,
K.G.Devine,
and
S.A.Benner
(2004).
Probing minor groove recognition contacts by DNA polymerases and reverse transcriptases using 3-deaza-2'-deoxyadenosine.
|
| |
Nucleic Acids Res,
32,
2241-2250.
|
 |
|
|
|
|
 |
E.Freisinger,
A.P.Grollman,
H.Miller,
and
C.Kisker
(2004).
Lesion (in)tolerance reveals insights into DNA replication fidelity.
|
| |
EMBO J,
23,
1494-1505.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
I.Andricioaei,
A.Goel,
D.Herschbach,
and
M.Karplus
(2004).
Dependence of DNA polymerase replication rate on external forces: a model based on molecular dynamics simulations.
|
| |
Biophys J,
87,
1478-1497.
|
 |
|
|
|
|
 |
L.G.Brieba,
B.F.Eichman,
R.J.Kokoska,
S.Doublié,
T.A.Kunkel,
and
T.Ellenberger
(2004).
Structural basis for the dual coding potential of 8-oxoguanosine by a high-fidelity DNA polymerase.
|
| |
EMBO J,
23,
3452-3461.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
M.E.Mysiak,
C.Wyman,
P.E.Holthuizen,
and
P.C.van der Vliet
(2004).
NFI and Oct-1 bend the Ad5 origin in the same direction leading to optimal DNA replication.
|
| |
Nucleic Acids Res,
32,
6218-6225.
|
 |
|
|
|
|
 |
M.Garcia-Diaz,
K.Bebenek,
J.M.Krahn,
L.Blanco,
T.A.Kunkel,
and
L.C.Pedersen
(2004).
A structural solution for the DNA polymerase lambda-dependent repair of DNA gaps with minimal homology.
|
| |
Mol Cell,
13,
561-572.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
M.Hogg,
S.S.Wallace,
and
S.Doublié
(2004).
Crystallographic snapshots of a replicative DNA polymerase encountering an abasic site.
|
| |
EMBO J,
23,
1483-1493.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
M.T.Washington,
I.G.Minko,
R.E.Johnson,
W.T.Wolfle,
T.M.Harris,
R.S.Lloyd,
S.Prakash,
and
L.Prakash
(2004).
Efficient and error-free replication past a minor-groove DNA adduct by the sequential action of human DNA polymerases iota and kappa.
|
| |
Mol Cell Biol,
24,
5687-5693.
|
 |
|
|
|
|
 |
N.Andraos,
S.Tabor,
and
C.C.Richardson
(2004).
The highly processive DNA polymerase of bacteriophage T5. Role of the unique N and C termini.
|
| |
J Biol Chem,
279,
50609-50618.
|
 |
|
|
|
|
 |
P.Garg,
C.M.Stith,
N.Sabouri,
E.Johansson,
and
P.M.Burgers
(2004).
Idling by DNA polymerase delta maintains a ligatable nick during lagging-strand DNA replication.
|
| |
Genes Dev,
18,
2764-2773.
|
 |
|
|
|
|
 |
Q.Guo,
Y.Shen,
N.L.Zhukovskaya,
J.Florián,
and
W.J.Tang
(2004).
Structural and kinetic analyses of the interaction of anthrax adenylyl cyclase toxin with reaction products cAMP and pyrophosphate.
|
| |
J Biol Chem,
279,
29427-29435.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
R.A.Perlow-Poehnelt,
I.Likhterov,
D.A.Scicchitano,
N.E.Geacintov,
and
S.Broyde
(2004).
The spacious active site of a Y-family DNA polymerase facilitates promiscuous nucleotide incorporation opposite a bulky carcinogen-DNA adduct: elucidating the structure-function relationship through experimental and computational approaches.
|
| |
J Biol Chem,
279,
36951-36961.
|
 |
|
|
|
|
 |
T.A.Steitz,
and
Y.W.Yin
(2004).
Accuracy, lesion bypass, strand displacement and translocation by DNA polymerases.
|
| |
Philos Trans R Soc Lond B Biol Sci,
359,
17-23.
|
 |
|
|
|
|
 |
V.M.Petrov,
and
J.D.Karam
(2004).
Diversity of structure and function of DNA polymerase (gp43) of T4-related bacteriophages.
|
| |
Biochemistry (Mosc),
69,
1213-1218.
|
 |
|
|
|
|
 |
V.Truniger,
J.M.Lázaro,
and
M.Salas
(2004).
Function of the C-terminus of phi29 DNA polymerase in DNA and terminal protein binding.
|
| |
Nucleic Acids Res,
32,
361-370.
|
 |
|
|
|
|
 |
Y.T.Hwang,
H.J.Zuccola,
Q.Lu,
and
C.B.Hwang
(2004).
A point mutation within conserved region VI of herpes simplex virus type 1 DNA polymerase confers altered drug sensitivity and enhances replication fidelity.
|
| |
J Virol,
78,
650-657.
|
 |
|
|
|
|
 |
Y.W.Yin,
and
T.A.Steitz
(2004).
The structural mechanism of translocation and helicase activity in T7 RNA polymerase.
|
| |
Cell,
116,
393-404.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
Y.Zhou,
and
T.S.Wang
(2004).
A coordinated temporal interplay of nucleosome reorganization factor, sister chromatin cohesion factor, and DNA polymerase alpha facilitates DNA replication.
|
| |
Mol Cell Biol,
24,
9568-9579.
|
 |
|
|
|
|
 |
A.M.DeLucia,
N.D.Grindley,
and
C.M.Joyce
(2003).
An error-prone family Y DNA polymerase (DinB homolog from Sulfolobus solfataricus) uses a 'steric gate' residue for discrimination against ribonucleotides.
|
| |
Nucleic Acids Res,
31,
4129-4137.
|
 |
|
|
|
|
 |
B.Marchand,
and
M.Götte
(2003).
Site-specific footprinting reveals differences in the translocation status of HIV-1 reverse transcriptase. Implications for polymerase translocation and drug resistance.
|
| |
J Biol Chem,
278,
35362-35372.
|
 |
|
|
|
|
 |
D.R.Thomsen,
N.L.Oien,
T.A.Hopkins,
M.L.Knechtel,
R.J.Brideau,
M.W.Wathen,
and
F.L.Homa
(2003).
Amino acid changes within conserved region III of the herpes simplex virus and human cytomegalovirus DNA polymerases confer resistance to 4-oxo-dihydroquinolines, a novel class of herpesvirus antiviral agents.
|
| |
J Virol,
77,
1868-1876.
|
 |
|
|
|
|
 |
E.Delagoutte,
and
P.H.Von Hippel
(2003).
Function and assembly of the bacteriophage T4 DNA replication complex: interactions of the T4 polymerase with various model DNA constructs.
|
| |
J Biol Chem,
278,
25435-25447.
|
 |
|
|
|
|
 |
E.S.Miller,
E.Kutter,
G.Mosig,
F.Arisaka,
T.Kunisawa,
and
W.Rüger
(2003).
Bacteriophage T4 genome.
|
| |
Microbiol Mol Biol Rev,
67,
86.
|
 |
|
|
|
|
 |
J.Bestman-Smith,
and
G.Boivin
(2003).
Drug resistance patterns of recombinant herpes simplex virus DNA polymerase mutants generated with a set of overlapping cosmids and plasmids.
|
| |
J Virol,
77,
7820-7829.
|
 |
|
|
|
|
 |
J.M.Krahn,
W.A.Beard,
H.Miller,
A.P.Grollman,
and
S.H.Wilson
(2003).
Structure of DNA polymerase beta with the mutagenic DNA lesion 8-oxodeoxyguanine reveals structural insights into its coding potential.
|
| |
Structure,
11,
121-127.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
K.E.McGinness,
and
G.F.Joyce
(2003).
In search of an RNA replicase ribozyme.
|
| |
Chem Biol,
10,
5.
|
 |
|
|
|
|
 |
K.Singh,
and
M.J.Modak
(2003).
Presence of 18-A long hydrogen bond track in the active site of Escherichia coli DNA polymerase I (Klenow fragment). Its requirement in the stabilization of enzyme-template-primer complex.
|
| |
J Biol Chem,
278,
11289-11302.
|
 |
|
|
|
|
 |
L.Tsujikawa,
M.Weinfield,
and
L.J.Reha-Krantz
(2003).
Differences in replication of a DNA template containing an ethyl phosphotriester by T4 DNA polymerase and Escherichia coli DNA polymerase I.
|
| |
Nucleic Acids Res,
31,
4965-4972.
|
 |
|
|
|
|
 |
M.Ogawa,
S.Limsirichaikul,
A.Niimi,
S.Iwai,
S.Yoshida,
and
M.Suzuki
(2003).
Distinct function of conserved amino acids in the fingers of Saccharomyces cerevisiae DNA polymerase alpha.
|
| |
J Biol Chem,
278,
19071-19078.
|
 |
|
|
|
|
 |
M.T.Washington,
W.T.Wolfle,
T.E.Spratt,
L.Prakash,
and
S.Prakash
(2003).
Yeast DNA polymerase eta makes functional contacts with the DNA minor groove only at the incoming nucleoside triphosphate.
|
| |
Proc Natl Acad Sci U S A,
100,
5113-5118.
|
 |
|
|
|
|
 |
N.Ito,
O.Nureki,
M.Shirouzu,
S.Yokoyama,
and
F.Hanaoka
(2003).
Crystal structure of the Pyrococcus horikoshii DNA primase-UTP complex: implications for the mechanism of primer synthesis.
|
| |
Genes Cells,
8,
913-923.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
P.J.Gutiérrez,
and
T.S.Wang
(2003).
Genomic instability induced by mutations in Saccharomyces cerevisiae POL1.
|
| |
Genetics,
165,
65-81.
|
 |
|
|
|
|
 |
P.V.Shcherbakova,
Y.I.Pavlov,
O.Chilkova,
I.B.Rogozin,
E.Johansson,
and
T.A.Kunkel
(2003).
Unique error signature of the four-subunit yeast DNA polymerase epsilon.
|
| |
J Biol Chem,
278,
43770-43780.
|
 |
|
|
|
|
 |
S.J.Johnson,
J.S.Taylor,
and
L.S.Beese
(2003).
Processive DNA synthesis observed in a polymerase crystal suggests a mechanism for the prevention of frameshift mutations.
|
| |
Proc Natl Acad Sci U S A,
100,
3895-3900.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
S.Limsirichaikul,
M.Ogawa,
A.Niimi,
S.Iwai,
T.Murate,
S.Yoshida,
and
M.Suzuki
(2003).
The Gly-952 residue of Saccharomyces cerevisiae DNA polymerase alpha is important in discriminating correct deoxyribonucleotides from incorrect ones.
|
| |
J Biol Chem,
278,
19079-19086.
|
 |
|
|
|
|
 |
S.Sun,
and
Y.Shamoo
(2003).
Biochemical characterization of interactions between DNA polymerase and single-stranded DNA-binding protein in bacteriophage RB69.
|
| |
J Biol Chem,
278,
3876-3881.
|
 |
|
|
|
|
 |
V.Truniger,
J.M.Lázaro,
M.de Vega,
L.Blanco,
and
M.Salas
(2003).
phi 29 DNA polymerase residue Leu384, highly conserved in motif B of eukaryotic type DNA replicases, is involved in nucleotide insertion fidelity.
|
| |
J Biol Chem,
278,
33482-33491.
|
 |
|
|
|
|
 |
W.A.Beard,
and
S.H.Wilson
(2003).
Structural insights into the origins of DNA polymerase fidelity.
|
| |
Structure,
11,
489-496.
|
 |
|
|
|
|
 |
W.Yang
(2003).
Damage repair DNA polymerases Y.
|
| |
Curr Opin Struct Biol,
13,
23-30.
|
 |
|
|
|
|
 |
A.B.Brenkman,
E.C.Breure,
and
P.C.van der Vliet
(2002).
Molecular architecture of adenovirus DNA polymerase and location of the protein primer.
|
| |
J Virol,
76,
8200-8207.
|
 |
|
|
|
|
 |
A.F.Gardner,
and
W.E.Jack
(2002).
Acyclic and dideoxy terminator preferences denote divergent sugar recognition by archaeon and Taq DNA polymerases.
|
| |
Nucleic Acids Res,
30,
605-613.
|
 |
|
|
|
|
 |
A.V.Cherepanov,
and
S.de Vries
(2002).
Dynamic mechanism of nick recognition by DNA ligase.
|
| |
Eur J Biochem,
269,
5993-5999.
|
 |
|
|
|
|
 |
E.Fidalgo da Silva,
S.S.Mandal,
and
L.J.Reha-Krantz
(2002).
Using 2-aminopurine fluorescence to measure incorporation of incorrect nucleotides by wild type and mutant bacteriophage T4 DNA polymerases.
|
| |
J Biol Chem,
277,
40640-40649.
|
 |
|
|
|
|
 |
E.T.Kool
(2002).
Active site tightness and substrate fit in DNA replication.
|
| |
Annu Rev Biochem,
71,
191-219.
|
 |
|
|
|
|
 |
G.Yang,
M.Franklin,
J.Li,
T.C.Lin,
and
W.Konigsberg
(2002).
A conserved Tyr residue is required for sugar selectivity in a Pol alpha DNA polymerase.
|
| |
Biochemistry,
41,
10256-10261.
|
 |
|
|
|
|
 |
H.A.Held,
and
S.A.Benner
(2002).
Challenging artificial genetic systems: thymidine analogs with 5-position sulfur functionality.
|
| |
Nucleic Acids Res,
30,
3857-3869.
|
 |
|
|
|
|
 |
I.V.Shevelev,
and
U.Hübscher
(2002).
The 3' 5' exonucleases.
|
| |
Nat Rev Mol Cell Biol,
3,
364-376.
|
 |
|
|
|
|
 |
K.Shimizu,
K.Hashimoto,
J.M.Kirchner,
W.Nakai,
H.Nishikawa,
M.A.Resnick,
and
A.Sugino
(2002).
Fidelity of DNA polymerase epsilon holoenzyme from budding yeast Saccharomyces cerevisiae.
|
| |
J Biol Chem,
277,
37422-37429.
|
 |
|
|
|
|
 |
M.J.Fogg,
L.H.Pearl,
and
B.A.Connolly
(2002).
Structural basis for uracil recognition by archaeal family B DNA polymerases.
|
| |
Nat Struct Biol,
9,
922-927.
|
 |
|
|
|
|
 |
O.Potapova,
N.D.Grindley,
and
C.M.Joyce
(2002).
The mutational specificity of the Dbh lesion bypass polymerase and its implications.
|
| |
J Biol Chem,
277,
28157-28166.
|
 |
|
|
|
|
 |
P.Cramer
(2002).
Common structural features of nucleic acid polymerases.
|
| |
Bioessays,
24,
724-729.
|
 |
|
|
|
|
 |
P.Cramer
(2002).
Multisubunit RNA polymerases.
|
| |
Curr Opin Struct Biol,
12,
89-97.
|
 |
|
|
|
|
 |
S.S.Mandal,
E.Fidalgo da Silva,
and
L.J.Reha-Krantz
(2002).
Using 2-aminopurine fluorescence to detect base unstacking in the template strand during nucleotide incorporation by the bacteriophage T4 DNA polymerase.
|
| |
Biochemistry,
41,
4399-4406.
|
 |
|
|
|
|
 |
U.Hubscher,
G.Maga,
and
S.Spadari
(2002).
Eukaryotic DNA polymerases.
|
| |
Annu Rev Biochem,
71,
133-163.
|
 |
|
|
|
|
 |
V.M.Petrov,
S.S.Ng,
and
J.D.Karam
(2002).
Protein determinants of RNA binding by DNA polymerase of the T4-related bacteriophage RB69.
|
| |
J Biol Chem,
277,
33041-33048.
|
 |
|
|
|
|
 |
V.Truniger,
J.M.Lázaro,
F.J.Esteban,
L.Blanco,
and
M.Salas
(2002).
A positively charged residue of phi29 DNA polymerase, highly conserved in DNA polymerases from families A and B, is involved in binding the incoming nucleotide.
|
| |
Nucleic Acids Res,
30,
1483-1492.
|
 |
|
|
|
|
 |
W.A.Beard,
D.D.Shock,
X.P.Yang,
S.F.DeLauder,
and
S.H.Wilson
(2002).
Loss of DNA polymerase beta stacking interactions with templating purines, but not pyrimidines, alters catalytic efficiency and fidelity.
|
| |
J Biol Chem,
277,
8235-8242.
|
 |
|
|
|
|
 |
W.H.Zhang,
E.S.Svarovskaia,
R.Barr,
and
V.K.Pathak
(2002).
Y586F mutation in murine leukemia virus reverse transcriptase decreases fidelity of DNA synthesis in regions associated with adenine-thymine tracts.
|
| |
Proc Natl Acad Sci U S A,
99,
10090-10095.
|
 |
|
|
|
|
 |
Y.W.Yin,
and
T.A.Steitz
(2002).
Structural basis for the transition from initiation to elongation transcription in T7 RNA polymerase.
|
| |
Science,
298,
1387-1395.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
H.Ling,
F.Boudsocq,
R.Woodgate,
and
W.Yang
(2001).
Crystal structure of a Y-family DNA polymerase in action: a mechanism for error-prone and lesion-bypass replication.
|
| |
Cell,
107,
91.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
W.A.Beard,
and
S.H.Wilson
(2001).
DNA lesion bypass polymerases open up.
|
| |
Structure,
9,
759-764.
|
 |
|
 |
 |
|
The most recent references are shown first.
Citation data come partly from CiteXplore and partly
from an automated harvesting procedure. Note that this is likely to be
only a partial list as not all journals are covered by
either method. However, we are continually building up the citation data
so more and more references will be included with time.
Where a reference describes a PDB structure, the PDB
codes are
shown on the right.
|
 |

