 |
PDBsum entry 2bcs

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Transferase, lyase/DNA
|
PDB id
|
|

|

|
2bcs
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
 |
Contents |
 |
|
|

|

|
|
|
|
|
|
|
|
|
|
|
|
* Residue conservation analysis
|
|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 2:
|
 |
E.C.2.7.7.7
- DNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
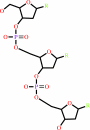 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
Bound ligand (Het Group name = )
corresponds exactly
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
E.C.4.2.99.-
- ?????
|
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
DOI no:
|
Cell
124:331-342
(2006)
|
|
PubMed id:
|
|
|
|

|
| |
|
Structural analysis of strand misalignment during DNA synthesis by a human DNA polymerase.
|
|
M.Garcia-Diaz,
K.Bebenek,
J.M.Krahn,
L.C.Pedersen,
T.A.Kunkel.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
Insertions and deletions in coding sequences can alter the reading frame of
genes and have profound biological consequences. In 1966, Streisinger proposed
that these mutations result from strand slippage, which in repetitive sequences
generates misaligned intermediates stabilized by correct base pairing that
support polymerization. We report here crystal structures of human DNA
polymerase lambda, which frequently generates deletion mutations, bound to such
intermediates. Each contains an extrahelical template nucleotide upstream of the
active site. Surprisingly, the extra nucleotide, even when combined with an
adjacent mismatch, does not perturb polymerase active site geometry, which is
indistinguishable from that for correctly aligned strands. These structures
reveal how pol lambda can polymerize on substrates with minimal homology during
repair of double-strand breaks and represent strand-slippage intermediates
consistent with Streisinger's classical hypothesis. They are thus relevant to
the origin of single-base deletions, a class of mutations that can confer strong
biological phenotypes.

|

|
|
|

|
| |
Selected figure(s)
|
|

|
| |
 |
 |

|
 |

|
 |
Figure 3.
Figure 3. The Extrahelical Adenine Is in Close Proximity to
β strand 8
|
 |
Figure 4.
Figure 4. Pol λ Can Tolerate Distortion upstream of the
Primer Terminus
|
 |
|
|

|
| |
The above figures are
reprinted
by permission from Cell Press:
Cell
(2006,
124,
331-342)
copyright 2006.
|
|
| |
Figures were
selected
by an automated process.
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Literature references that cite this PDB file's key reference
|
|
 |
| |
PubMed id
|
 |
Reference
|
 |
|
|
|
 |
A.L.Abdulovic,
S.E.Hile,
T.A.Kunkel,
and
K.A.Eckert
(2011).
The in vitro fidelity of yeast DNA polymerase δ and polymerase ɛ holoenzymes during dinucleotide microsatellite DNA synthesis.
|
| |
DNA Repair (Amst),
10,
497-505.
|
 |
|
|
|
|
 |
K.Bebenek,
L.C.Pedersen,
and
T.A.Kunkel
(2011).
Replication infidelity via a mismatch with Watson-Crick geometry.
|
| |
Proc Natl Acad Sci U S A,
108,
1862-1867.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
P.Xie
(2011).
A model for the dynamics of mammalian family X DNA polymerases.
|
| |
J Theor Biol,
277,
111-122.
|
 |
|
|
|
|
 |
A.Demogines,
A.M.East,
J.H.Lee,
S.R.Grossman,
P.C.Sabeti,
T.T.Paull,
and
S.L.Sawyer
(2010).
Ancient and recent adaptive evolution of primate non-homologous end joining genes.
|
| |
PLoS Genet,
6,
e1001169.
|
 |
|
|
|
|
 |
J.Yamtich,
and
J.B.Sweasy
(2010).
DNA polymerase family X: function, structure, and cellular roles.
|
| |
Biochim Biophys Acta,
1804,
1136-1150.
|
 |
|
|
|
|
 |
K.Bebenek,
M.Garcia-Diaz,
R.Z.Zhou,
L.F.Povirk,
and
T.A.Kunkel
(2010).
Loop 1 modulates the fidelity of DNA polymerase lambda.
|
| |
Nucleic Acids Res,
38,
5419-5431.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
K.Takakusagi,
Y.Takakusagi,
K.Ohta,
S.Aoki,
F.Sugawara,
and
K.Sakaguchi
(2010).
A sulfoglycolipid beta-sulfoquinovosyldiacylglycerol (betaSQDG) binds to Met1-Arg95 region of murine DNA polymerase lambda (Mmpol lambda) and inhibits its nuclear transit.
|
| |
Protein Eng Des Sel,
23,
51-60.
|
 |
|
|
|
|
 |
O.Rechkoblit,
A.Kolbanovskiy,
L.Malinina,
N.E.Geacintov,
S.Broyde,
and
D.J.Patel
(2010).
Mechanism of error-free and semitargeted mutagenic bypass of an aromatic amine lesion by Y-family polymerase Dpo4.
|
| |
Nat Struct Mol Biol,
17,
379-388.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
F.Romain,
I.Barbosa,
J.Gouge,
F.Rougeon,
and
M.Delarue
(2009).
Conferring a template-dependent polymerase activity to terminal deoxynucleotidyltransferase by mutations in the Loop1 region.
|
| |
Nucleic Acids Res,
37,
4642-4656.
|
 |
|
|
|
|
 |
F.Wang,
and
W.Yang
(2009).
Structural insight into translesion synthesis by DNA Pol II.
|
| |
Cell,
139,
1279-1289.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
M.C.Foley,
and
T.Schlick
(2009).
Relationship between conformational changes in pol lambda's active site upon binding incorrect nucleotides and mismatch incorporation rates.
|
| |
J Phys Chem B,
113,
13035-13047.
|
 |
|
|
|
|
 |
M.Garcia-Diaz,
K.Bebenek,
A.A.Larrea,
J.M.Havener,
L.Perera,
J.M.Krahn,
L.C.Pedersen,
D.A.Ramsden,
and
T.A.Kunkel
(2009).
Template strand scrunching during DNA gap repair synthesis by human polymerase lambda.
|
| |
Nat Struct Mol Biol,
16,
967-972.
|
 |
|
|
|
|
 |
M.Urban,
N.Joubert,
M.Hocek,
R.E.Alexander,
and
R.D.Kuchta
(2009).
Herpes simplex virus-1 DNA primase: a remarkably inaccurate yet selective polymerase.
|
| |
Biochemistry,
48,
10866-10881.
|
 |
|
|
|
|
 |
W.A.Baase,
D.Jose,
B.C.Ponedel,
P.H.von Hippel,
and
N.P.Johnson
(2009).
DNA models of trinucleotide frameshift deletions: the formation of loops and bulges at the primer-template junction.
|
| |
Nucleic Acids Res,
37,
1682-1689.
|
 |
|
|
|
|
 |
J.M.Daley,
and
T.E.Wilson
(2008).
Evidence that base stacking potential in annealed 3' overhangs determines polymerase utilization in yeast nonhomologous end joining.
|
| |
DNA Repair (Amst),
7,
67-76.
|
 |
|
|
|
|
 |
M.E.Arana,
M.Seki,
R.D.Wood,
I.B.Rogozin,
and
T.A.Kunkel
(2008).
Low-fidelity DNA synthesis by human DNA polymerase theta.
|
| |
Nucleic Acids Res,
36,
3847-3856.
|
 |
|
|
|
|
 |
A.F.Moon,
M.Garcia-Diaz,
K.Bebenek,
B.J.Davis,
X.Zhong,
D.A.Ramsden,
T.A.Kunkel,
and
L.C.Pedersen
(2007).
Structural insight into the substrate specificity of DNA Polymerase mu.
|
| |
Nat Struct Mol Biol,
14,
45-53.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
A.F.Moon,
M.Garcia-Diaz,
V.K.Batra,
W.A.Beard,
K.Bebenek,
T.A.Kunkel,
S.H.Wilson,
and
L.C.Pedersen
(2007).
The X family portrait: structural insights into biological functions of X family polymerases.
|
| |
DNA Repair (Amst),
6,
1709-1725.
|
 |
|
|
|
|
 |
K.A.Fiala,
J.A.Brown,
H.Ling,
A.K.Kshetry,
J.Zhang,
J.S.Taylor,
W.Yang,
and
Z.Suo
(2007).
Mechanism of template-independent nucleotide incorporation catalyzed by a template-dependent DNA polymerase.
|
| |
J Mol Biol,
365,
590-602.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
M.E.Arana,
K.Takata,
M.Garcia-Diaz,
R.D.Wood,
and
T.A.Kunkel
(2007).
A unique error signature for human DNA polymerase nu.
|
| |
DNA Repair (Amst),
6,
213-223.
|
 |
|
|
|
|
 |
M.Garcia-Diaz,
K.Bebenek,
J.M.Krahn,
L.C.Pedersen,
and
T.A.Kunkel
(2007).
Role of the catalytic metal during polymerization by DNA polymerase lambda.
|
| |
DNA Repair (Amst),
6,
1333-1340.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
M.Garcia-Diaz,
and
K.Bebenek
(2007).
Multiple functions of DNA polymerases.
|
| |
CRC Crit Rev Plant Sci,
26,
105-122.
|
 |
|
|
|
|
 |
N.C.Brissett,
R.S.Pitcher,
R.Juarez,
A.J.Picher,
A.J.Green,
T.R.Dafforn,
G.C.Fox,
L.Blanco,
and
A.J.Doherty
(2007).
Structure of a NHEJ polymerase-mediated DNA synaptic complex.
|
| |
Science,
318,
456-459.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
P.R.Meyer,
W.Rutvisuttinunt,
S.E.Matsuura,
A.G.So,
and
W.A.Scott
(2007).
Stable complexes formed by HIV-1 reverse transcriptase at distinct positions on the primer-template controlled by binding deoxynucleoside triphosphates or foscarnet.
|
| |
J Mol Biol,
369,
41-54.
|
 |
|
|
|
|
 |
V.J.Cannistraro,
and
J.S.Taylor
(2007).
Ability of polymerase eta and T7 DNA polymerase to bypass bulge structures.
|
| |
J Biol Chem,
282,
11188-11196.
|
 |
|
|
|
|
 |
A.J.Picher,
M.García-Díaz,
K.Bebenek,
L.C.Pedersen,
T.A.Kunkel,
and
L.Blanco
(2006).
Promiscuous mismatch extension by human DNA polymerase lambda.
|
| |
Nucleic Acids Res,
34,
3259-3266.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
B.Bertocci,
A.De Smet,
J.C.Weill,
and
C.A.Reynaud
(2006).
Nonoverlapping functions of DNA polymerases mu, lambda, and terminal deoxynucleotidyltransferase during immunoglobulin V(D)J recombination in vivo.
|
| |
Immunity,
25,
31-41.
|
 |
|
|
|
|
 |
E.Kashkina,
M.Anikin,
F.Brueckner,
R.T.Pomerantz,
W.T.McAllister,
P.Cramer,
and
D.Temiakov
(2006).
Template misalignment in multisubunit RNA polymerases and transcription fidelity.
|
| |
Mol Cell,
24,
257-266.
|
 |
|
|
|
|
 |
M.Garcia-Diaz,
and
T.A.Kunkel
(2006).
Mechanism of a genetic glissando: structural biology of indel mutations.
|
| |
Trends Biochem Sci,
31,
206-214.
|
 |
|
|
|
|
 |
R.T.Pomerantz,
D.Temiakov,
M.Anikin,
D.G.Vassylyev,
and
W.T.McAllister
(2006).
A mechanism of nucleotide misincorporation during transcription due to template-strand misalignment.
|
| |
Mol Cell,
24,
245-255.
|
 |
|
|
|
|
 |
W.W.Duym,
K.A.Fiala,
N.Bhatt,
and
Z.Suo
(2006).
Kinetic effect of a downstream strand and its 5'-terminal moieties on single nucleotide gap-filling synthesis catalyzed by human DNA polymerase lambda.
|
| |
J Biol Chem,
281,
35649-35655.
|
 |
|
 |
 |
|
The most recent references are shown first.
Citation data come partly from CiteXplore and partly
from an automated harvesting procedure. Note that this is likely to be
only a partial list as not all journals are covered by
either method. However, we are continually building up the citation data
so more and more references will be included with time.
Where a reference describes a PDB structure, the PDB
codes are
shown on the right.
|
 |

