 |
PDBsum entry 1lv5

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Transferase/DNA
|
PDB id
|
|

|

|
1lv5
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
 |
Contents |
 |
|
|

|

|
|
|
|
|
|
|
|
|
|
|
|
* Residue conservation analysis
|
|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class:
|
 |
E.C.2.7.7.7
- DNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
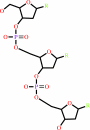 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
DOI no:
|
Proc Natl Acad Sci U S A
100:3895-3900
(2003)
|
|
PubMed id:
|
|
|
|

|
| |
|
Processive DNA synthesis observed in a polymerase crystal suggests a mechanism for the prevention of frameshift mutations.
|
|
S.J.Johnson,
J.S.Taylor,
L.S.Beese.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
DNA polymerases replicate DNA by adding nucleotides to a growing primer strand
while avoiding frameshift and point mutations. Here we present a series of up to
six successive replication events that were obtained by extension of a primed
template directly in a crystal of the thermostable Bacillus DNA polymerase I.
The 6-bp extension involves a 20-A translocation of the DNA duplex, representing
the largest molecular movement observed in a protein crystal. In addition, we
obtained the structure of a "closed" conformation of the enzyme with a
bound triphosphate juxtaposed to a template and a dideoxy-terminated primer by
constructing a point mutant that destroys a crystal lattice contact stabilizing
the wild-type polymerase in an "open" conformation. Together, these
observations allow many of the steps involved in DNA replication to be observed
in the same enzyme at near atomic detail. The successive replication events
observed directly by catalysis in the crystal confirm the general reaction
sequence deduced from observations obtained by using several other polymerases
and further refine critical aspects of the known reaction mechanism, and also
allow us to propose new features that concern the regulated transfer of the
template strand between a preinsertion site and an insertion site. We propose
that such regulated transfer is an important element in the prevention of
frameshift mutations in high-fidelity DNA polymerases. The ability to observe
processive, high-fidelity replication directly in a crystal establishes this
polymerase as a powerful model system for mechanistic studies in which the
structural consequences of mismatches and DNA adducts are observed.

|

|
|
|

|
| |
Selected figure(s)
|
|

|
| |
 |
 |

|
 |

|
 |
Figure 2.
Fig. 2. Active site superposition of open and closed BF
structures. (A) The 11-bp open binary complex (yellow) and the
closed ternary complex (blue) are shown in stereo view. The
largest conformational differences occur in the fingers domain,
including the O helix, O1 helix, and preinsertion site. The
acceptor template base (n) occupies the preinsertion site in the
open conformation and the insertion site in the closed
conformation. (B) A close-up view of the preinsertion site. The
locations of the conserved Tyr-714 are indicated.
|
 |
Figure 3.
Fig. 3. Conformational interlocks during DNA synthesis. A
schematic overview of the polymerase active site (A) and atomic
coordinates (B) derived from the open and closed BF structures
represent a complete round of DNA synthesis. The conformational
changes described here are presented in animated form in Movie
1, which is published as supporting information on the PNAS web
site. The reaction cycle starts with the acceptor template base
(n, red) bound at the template preinsertion site (between the O
and O1 helices; green shading); Tyr-714 blocks access to the
insertion site (blue shading) and stacks with the n-1 base pair
at the postinsertion site (gray shading). Formation of the
closed conformation involves rearrangement of the O and O1
helices, which simultaneously blocks the template preinsertion
site and unblocks the insertion site. These rearrangements move
the acceptor template base (n) to the insertion site, where it
pairs with an incoming dNTP (green). Nucleotide incorporation
occurs on formation of a cognate base pair and proper assembly
of the catalytic site (orange shading). The cycle is completed
with translocation of the DNA by one base pair position. The
polymerase resets to the open conformation in preparation for
the next round of DNA synthesis.
|
 |
|
|
| |
Figures were
selected
by an automated process.
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Literature references that cite this PDB file's key reference
|
|
 |
| |
PubMed id
|
 |
Reference
|
 |
|
|
|
 |
T.Nakamura,
Y.Zhao,
Y.Yamagata,
Y.J.Hua,
and
W.Yang
(2012).
Watching DNA polymerase η make a phosphodiester bond.
|
| |
Nature,
487,
196-201.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
J.S.Fraser,
and
C.J.Jackson
(2011).
Mining electron density for functionally relevant protein polysterism in crystal structures.
|
| |
Cell Mol Life Sci,
68,
1829-1841.
|
 |
|
|
|
|
 |
M.L.Gleghorn,
E.K.Davydova,
R.Basu,
L.B.Rothman-Denes,
and
K.S.Murakami
(2011).
X-ray crystal structures elucidate the nucleotidyl transfer reaction of transcript initiation using two nucleotides.
|
| |
Proc Natl Acad Sci U S A,
108,
3566-3571.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
A.A.Golosov,
J.J.Warren,
L.S.Beese,
and
M.Karplus
(2010).
The mechanism of the translocation step in DNA replication by DNA polymerase I: a computer simulation analysis.
|
| |
Structure,
18,
83-93.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
C.M.Joyce
(2010).
Techniques used to study the DNA polymerase reaction pathway.
|
| |
Biochim Biophys Acta,
1804,
1032-1040.
|
 |
|
|
|
|
 |
D.Zhi,
M.Shatsky,
and
S.E.Brenner
(2010).
Alignment-free local structural search by writhe decomposition.
|
| |
Bioinformatics,
26,
1176-1184.
|
 |
|
|
|
|
 |
G.Stengel,
M.Urban,
B.W.Purse,
and
R.D.Kuchta
(2010).
Incorporation of the fluorescent ribonucleotide analogue tCTP by T7 RNA polymerase.
|
| |
Anal Chem,
82,
1082-1089.
|
 |
|
|
|
|
 |
J.D.Pata
(2010).
Structural diversity of the Y-family DNA polymerases.
|
| |
Biochim Biophys Acta,
1804,
1124-1135.
|
 |
|
|
|
|
 |
J.D.Pata,
and
J.Jaeger
(2010).
Molecular machines and targeted molecular dynamics: DNA in motion.
|
| |
Structure,
18,
4-6.
|
 |
|
|
|
|
 |
J.Hafner,
and
W.Zheng
(2010).
Optimal modeling of atomic fluctuations in protein crystal structures for weak crystal contact interactions.
|
| |
J Chem Phys,
132,
014111.
|
 |
|
|
|
|
 |
K.Datta,
N.P.Johnson,
and
P.H.von Hippel
(2010).
DNA conformational changes at the primer-template junction regulate the fidelity of replication by DNA polymerase.
|
| |
Proc Natl Acad Sci U S A,
107,
17980-17985.
|
 |
|
|
|
|
 |
M.Yokoyama,
H.Mori,
and
H.Sato
(2010).
Allosteric regulation of HIV-1 reverse transcriptase by ATP for nucleotide selection.
|
| |
PLoS One,
5,
e8867.
|
 |
|
|
|
|
 |
P.Gong,
and
O.B.Peersen
(2010).
Structural basis for active site closure by the poliovirus RNA-dependent RNA polymerase.
|
| |
Proc Natl Acad Sci U S A,
107,
22505-22510.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
R.G.Federley,
and
L.J.Romano
(2010).
DNA polymerase: structural homology, conformational dynamics, and the effects of carcinogenic DNA adducts.
|
| |
J Nucleic Acids,
2010,
0.
|
 |
|
|
|
|
 |
R.Venkatramani,
and
R.Radhakrishnan
(2010).
Computational delineation of the catalytic step of a high-fidelity DNA polymerase.
|
| |
Protein Sci,
19,
815-825.
|
 |
|
|
|
|
 |
S.K.Perumal,
H.Yue,
Z.Hu,
M.M.Spiering,
and
S.J.Benkovic
(2010).
Single-molecule studies of DNA replisome function.
|
| |
Biochim Biophys Acta,
1804,
1094-1112.
|
 |
|
|
|
|
 |
Y.Santoso,
C.M.Joyce,
O.Potapova,
L.Le Reste,
J.Hohlbein,
J.P.Torella,
N.D.Grindley,
and
A.N.Kapanidis
(2010).
Conformational transitions in DNA polymerase I revealed by single-molecule FRET.
|
| |
Proc Natl Acad Sci U S A,
107,
715-720.
|
 |
|
|
|
|
 |
Y.Santoso,
J.P.Torella,
and
A.N.Kapanidis
(2010).
Characterizing single-molecule FRET dynamics with probability distribution analysis.
|
| |
Chemphyschem,
11,
2209-2219.
|
 |
|
|
|
|
 |
C.Castro,
E.D.Smidansky,
J.J.Arnold,
K.R.Maksimchuk,
I.Moustafa,
A.Uchida,
M.Götte,
W.Konigsberg,
and
C.E.Cameron
(2009).
Nucleic acid polymerases use a general acid for nucleotidyl transfer.
|
| |
Nat Struct Mol Biol,
16,
212-218.
|
 |
|
|
|
|
 |
C.Xu,
B.A.Maxwell,
J.A.Brown,
L.Zhang,
and
Z.Suo
(2009).
Global conformational dynamics of a Y-family DNA polymerase during catalysis.
|
| |
PLoS Biol,
7,
e1000225.
|
 |
|
|
|
|
 |
K.Datta,
N.P.Johnson,
V.J.LiCata,
and
P.H.von Hippel
(2009).
Local conformations and competitive binding affinities of single- and double-stranded primer-template DNA at the polymerization and editing active sites of DNA polymerases.
|
| |
J Biol Chem,
284,
17180-17193.
|
 |
|
|
|
|
 |
M.Trostler,
A.Delier,
J.Beckman,
M.Urban,
J.N.Patro,
T.E.Spratt,
L.S.Beese,
and
R.D.Kuchta
(2009).
Discrimination between right and wrong purine dNTPs by DNA polymerase I from Bacillus stearothermophilus.
|
| |
Biochemistry,
48,
4633-4641.
|
 |
|
|
|
|
 |
N.A.Wilson,
R.Abu-Shumays,
B.Gyarfas,
H.Wang,
K.R.Lieberman,
M.Akeson,
and
W.B.Dunbar
(2009).
Electronic control of DNA polymerase binding and unbinding to single DNA molecules.
|
| |
ACS Nano,
3,
995.
|
 |
|
|
|
|
 |
N.Hurt,
H.Wang,
M.Akeson,
and
K.R.Lieberman
(2009).
Specific nucleotide binding and rebinding to individual DNA polymerase complexes captured on a nanopore.
|
| |
J Am Chem Soc,
131,
3772-3778.
|
 |
|
|
|
|
 |
P.Xu,
L.Oum,
Y.C.Lee,
N.E.Geacintov,
and
S.Broyde
(2009).
Visualizing sequence-governed nucleotide selectivities and mutagenic consequences through a replicative cycle: processing of a bulky carcinogen N2-dG lesion in a Y-family DNA polymerase.
|
| |
Biochemistry,
48,
4677-4690.
|
 |
|
|
|
|
 |
J.Cramer,
G.Rangam,
A.Marx,
and
T.Restle
(2008).
Varied active-site constraints in the klenow fragment of E. coli DNA polymerase I and the lesion-bypass Dbh DNA polymerase.
|
| |
Chembiochem,
9,
1243-1250.
|
 |
|
|
|
|
 |
K.Bebenek,
M.Garcia-Diaz,
M.C.Foley,
L.C.Pedersen,
T.Schlick,
and
T.A.Kunkel
(2008).
Substrate-induced DNA strand misalignment during catalytic cycling by DNA polymerase lambda.
|
| |
EMBO Rep,
9,
459-464.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
K.K.Ng,
J.J.Arnold,
and
C.E.Cameron
(2008).
Structure-function relationships among RNA-dependent RNA polymerases.
|
| |
Curr Top Microbiol Immunol,
320,
137-156.
|
 |
|
|
|
|
 |
M.Renders,
R.Lievrouw,
M.Krecmerová,
A.Holý,
and
P.Herdewijn
(2008).
Enzymatic polymerization of phosphonate nucleosides.
|
| |
Chembiochem,
9,
2883-2888.
|
 |
|
|
|
|
 |
R.Venkatramani,
and
R.Radhakrishnan
(2008).
Effect of oxidatively damaged DNA on the active site preorganization during nucleotide incorporation in a high fidelity polymerase from Bacillus stearothermophilus.
|
| |
Proteins,
71,
1360-1372.
|
 |
|
|
|
|
 |
R.Venkatramani,
and
R.Radhakrishnan
(2008).
Computational study of the force dependence of phosphoryl transfer during DNA synthesis by a high fidelity polymerase.
|
| |
Phys Rev Lett,
100,
088102.
|
 |
|
|
|
|
 |
S.Broyde,
L.Wang,
O.Rechkoblit,
N.E.Geacintov,
and
D.J.Patel
(2008).
Lesion processing: high-fidelity versus lesion-bypass DNA polymerases.
|
| |
Trends Biochem Sci,
33,
209-219.
|
 |
|
|
|
|
 |
A.A.Thompson,
R.A.Albertini,
and
O.B.Peersen
(2007).
Stabilization of poliovirus polymerase by NTP binding and fingers-thumb interactions.
|
| |
J Mol Biol,
366,
1459-1474.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
A.J.Berman,
S.Kamtekar,
J.L.Goodman,
J.M.Lázaro,
M.de Vega,
L.Blanco,
M.Salas,
and
T.A.Steitz
(2007).
Structures of phi29 DNA polymerase complexed with substrate: the mechanism of translocation in B-family polymerases.
|
| |
EMBO J,
26,
3494-3505.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
C.Ferrer-Orta,
A.Arias,
R.Pérez-Luque,
C.Escarmís,
E.Domingo,
and
N.Verdaguer
(2007).
Sequential structures provide insights into the fidelity of RNA replication.
|
| |
Proc Natl Acad Sci U S A,
104,
9463-9468.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
C.H.Tsai,
J.Chen,
and
J.W.Szostak
(2007).
Enzymatic synthesis of DNA on glycerol nucleic acid templates without stable duplex formation between product and template.
|
| |
Proc Natl Acad Sci U S A,
104,
14598-14603.
|
 |
|
|
|
|
 |
G.Luo,
M.Wang,
W.H.Konigsberg,
and
X.S.Xie
(2007).
Single-molecule and ensemble fluorescence assays for a functionally important conformational change in T7 DNA polymerase.
|
| |
Proc Natl Acad Sci U S A,
104,
12610-12615.
|
 |
|
|
|
|
 |
N.Z.Rudinger,
R.Kranaster,
and
A.Marx
(2007).
Hydrophobic amino acid and single-atom substitutions increase DNA polymerase selectivity.
|
| |
Chem Biol,
14,
185-194.
|
 |
|
|
|
|
 |
P.R.Meyer,
W.Rutvisuttinunt,
S.E.Matsuura,
A.G.So,
and
W.A.Scott
(2007).
Stable complexes formed by HIV-1 reverse transcriptase at distinct positions on the primer-template controlled by binding deoxynucleoside triphosphates or foscarnet.
|
| |
J Mol Biol,
369,
41-54.
|
 |
|
|
|
|
 |
P.Xu,
L.Oum,
L.S.Beese,
N.E.Geacintov,
and
S.Broyde
(2007).
Following an environmental carcinogen N2-dG adduct through replication: elucidating blockage and bypass in a high-fidelity DNA polymerase.
|
| |
Nucleic Acids Res,
35,
4275-4288.
|
 |
|
|
|
|
 |
S.Benner,
R.J.Chen,
N.A.Wilson,
R.Abu-Shumays,
N.Hurt,
K.R.Lieberman,
D.W.Deamer,
W.B.Dunbar,
and
M.Akeson
(2007).
Sequence-specific detection of individual DNA polymerase complexes in real time using a nanopore.
|
| |
Nat Nanotechnol,
2,
718-724.
|
 |
|
|
|
|
 |
S.Meneni,
F.Liang,
and
B.P.Cho
(2007).
Examination of the long-range effects of aminofluorene-induced conformational heterogeneity and its relevance to the mechanism of translesional DNA synthesis.
|
| |
J Mol Biol,
366,
1387-1400.
|
 |
|
|
|
|
 |
E.Kashkina,
M.Anikin,
F.Brueckner,
R.T.Pomerantz,
W.T.McAllister,
P.Cramer,
and
D.Temiakov
(2006).
Template misalignment in multisubunit RNA polymerases and transcription fidelity.
|
| |
Mol Cell,
24,
257-266.
|
 |
|
|
|
|
 |
J.J.Warren,
L.J.Forsberg,
and
L.S.Beese
(2006).
The structural basis for the mutagenicity of O(6)-methyl-guanine lesions.
|
| |
Proc Natl Acad Sci U S A,
103,
19701-19706.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
L.V.Gening,
S.A.Klincheva,
A.Reshetnjak,
A.P.Grollman,
and
H.Miller
(2006).
RNA aptamers selected against DNA polymerase beta inhibit the polymerase activities of DNA polymerases beta and kappa.
|
| |
Nucleic Acids Res,
34,
2579-2586.
|
 |
|
|
|
|
 |
M.Garcia-Diaz,
K.Bebenek,
J.M.Krahn,
L.C.Pedersen,
and
T.A.Kunkel
(2006).
Structural analysis of strand misalignment during DNA synthesis by a human DNA polymerase.
|
| |
Cell,
124,
331-342.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
M.Garcia-Diaz,
and
T.A.Kunkel
(2006).
Mechanism of a genetic glissando: structural biology of indel mutations.
|
| |
Trends Biochem Sci,
31,
206-214.
|
 |
|
|
|
|
 |
O.Potapova,
C.Chan,
A.M.DeLucia,
S.A.Helquist,
E.T.Kool,
N.D.Grindley,
and
C.M.Joyce
(2006).
DNA polymerase catalysis in the absence of Watson-Crick hydrogen bonds: analysis by single-turnover kinetics.
|
| |
Biochemistry,
45,
890-898.
|
 |
|
|
|
|
 |
O.Rechkoblit,
L.Malinina,
Y.Cheng,
V.Kuryavyi,
S.Broyde,
N.E.Geacintov,
and
D.J.Patel
(2006).
Stepwise translocation of Dpo4 polymerase during error-free bypass of an oxoG lesion.
|
| |
PLoS Biol,
4,
e11.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
R.T.Pomerantz,
D.Temiakov,
M.Anikin,
D.G.Vassylyev,
and
W.T.McAllister
(2006).
A mechanism of nucleotide misincorporation during transcription due to template-strand misalignment.
|
| |
Mol Cell,
24,
245-255.
|
 |
|
|
|
|
 |
S.Vichier-Guerre,
S.Ferris,
N.Auberger,
K.Mahiddine,
and
J.L.Jestin
(2006).
A population of thermostable reverse transcriptases evolved from Thermus aquaticus DNA polymerase I by phage display.
|
| |
Angew Chem Int Ed Engl,
45,
6133-6137.
|
 |
|
|
|
|
 |
T.A.Steitz
(2006).
Visualizing polynucleotide polymerase machines at work.
|
| |
EMBO J,
25,
3458-3468.
|
 |
|
|
|
|
 |
V.K.Batra,
W.A.Beard,
D.D.Shock,
J.M.Krahn,
L.C.Pedersen,
and
S.H.Wilson
(2006).
Magnesium-induced assembly of a complete DNA polymerase catalytic complex.
|
| |
Structure,
14,
757-766.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
D.Bourgeois,
and
A.Royant
(2005).
Advances in kinetic protein crystallography.
|
| |
Curr Opin Struct Biol,
15,
538-547.
|
 |
|
|
|
|
 |
J.Florián,
M.F.Goodman,
and
A.Warshel
(2005).
Computer simulations of protein functions: searching for the molecular origin of the replication fidelity of DNA polymerases.
|
| |
Proc Natl Acad Sci U S A,
102,
6819-6824.
|
 |
|
|
|
|
 |
M.Garcia-Diaz,
K.Bebenek,
J.M.Krahn,
T.A.Kunkel,
and
L.C.Pedersen
(2005).
A closed conformation for the Pol lambda catalytic cycle.
|
| |
Nat Struct Mol Biol,
12,
97-98.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
R.Chen,
M.Yokoyama,
H.Sato,
C.Reilly,
and
L.M.Mansky
(2005).
Human immunodeficiency virus mutagenesis during antiviral therapy: impact of drug-resistant reverse transcriptase and nucleoside and nonnucleoside reverse transcriptase inhibitors on human immunodeficiency virus type 1 mutation frequencies.
|
| |
J Virol,
79,
12045-12057.
|
 |
|
|
|
|
 |
V.K.Batra,
W.A.Beard,
D.D.Shock,
L.C.Pedersen,
and
S.H.Wilson
(2005).
Nucleotide-induced DNA polymerase active site motions accommodating a mutagenic DNA intermediate.
|
| |
Structure,
13,
1225-1233.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
Z.F.Burton,
M.Feig,
X.Q.Gong,
C.Zhang,
Y.A.Nedialkov,
and
Y.Xiong
(2005).
NTP-driven translocation and regulation of downstream template opening by multi-subunit RNA polymerases.
|
| |
Biochem Cell Biol,
83,
486-496.
|
 |
|
|
|
|
 |
D.Das,
and
M.M.Georgiadis
(2004).
The crystal structure of the monomeric reverse transcriptase from Moloney murine leukemia virus.
|
| |
Structure,
12,
819-829.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
D.Temiakov,
V.Patlan,
M.Anikin,
W.T.McAllister,
S.Yokoyama,
and
D.G.Vassylyev
(2004).
Structural basis for substrate selection by t7 RNA polymerase.
|
| |
Cell,
116,
381-391.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
D.W.Gohara,
J.J.Arnold,
and
C.E.Cameron
(2004).
Poliovirus RNA-dependent RNA polymerase (3Dpol): kinetic, thermodynamic, and structural analysis of ribonucleotide selection.
|
| |
Biochemistry,
43,
5149-5158.
|
 |
|
|
|
|
 |
G.W.Hsu,
M.Ober,
T.Carell,
and
L.S.Beese
(2004).
Error-prone replication of oxidatively damaged DNA by a high-fidelity DNA polymerase.
|
| |
Nature,
431,
217-221.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
K.Arora,
and
T.Schlick
(2004).
In silico evidence for DNA polymerase-beta's substrate-induced conformational change.
|
| |
Biophys J,
87,
3088-3099.
|
 |
|
|
|
|
 |
K.S.Gajiwala,
and
C.Pinko
(2004).
Structural rearrangement accompanying NAD+ synthesis within a bacterial DNA ligase crystal.
|
| |
Structure,
12,
1449-1459.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
L.G.Brieba,
B.F.Eichman,
R.J.Kokoska,
S.Doublié,
T.A.Kunkel,
and
T.Ellenberger
(2004).
Structural basis for the dual coding potential of 8-oxoguanosine by a high-fidelity DNA polymerase.
|
| |
EMBO J,
23,
3452-3461.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
M.Hogg,
S.S.Wallace,
and
S.Doublié
(2004).
Crystallographic snapshots of a replicative DNA polymerase encountering an abasic site.
|
| |
EMBO J,
23,
1483-1493.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
M.Karplus,
and
Y.Q.Gao
(2004).
Biomolecular motors: the F1-ATPase paradigm.
|
| |
Curr Opin Struct Biol,
14,
250-259.
|
 |
|
|
|
|
 |
M.Strerath,
J.Gaster,
and
A.Marx
(2004).
Recognition of remote mismatches by DNA polymerases.
|
| |
Chembiochem,
5,
1585-1588.
|
 |
|
|
|
|
 |
R.L.Crowther,
D.P.Remeta,
C.A.Minetti,
D.Das,
S.P.Montano,
and
M.M.Georgiadis
(2004).
Structural and energetic characterization of nucleic acid-binding to the fingers domain of Moloney murine leukemia virus reverse transcriptase.
|
| |
Proteins,
57,
15-26.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
R.Landick
(2004).
Active-site dynamics in RNA polymerases.
|
| |
Cell,
116,
351-353.
|
 |
|
|
|
|
 |
R.Radhakrishnan,
and
T.Schlick
(2004).
Orchestration of cooperative events in DNA synthesis and repair mechanism unraveled by transition path sampling of DNA polymerase beta's closing.
|
| |
Proc Natl Acad Sci U S A,
101,
5970-5975.
|
 |
|
|
|
|
 |
S.Dutta,
Y.Li,
D.Johnson,
L.Dzantiev,
C.C.Richardson,
L.J.Romano,
and
T.Ellenberger
(2004).
Crystal structures of 2-acetylaminofluorene and 2-aminofluorene in complex with T7 DNA polymerase reveal mechanisms of mutagenesis.
|
| |
Proc Natl Acad Sci U S A,
101,
16186-16191.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
S.Fujii,
and
R.P.Fuchs
(2004).
Defining the position of the switches between replicative and bypass DNA polymerases.
|
| |
EMBO J,
23,
4342-4352.
|
 |
|
|
|
|
 |
S.J.Johnson,
and
L.S.Beese
(2004).
Structures of mismatch replication errors observed in a DNA polymerase.
|
| |
Cell,
116,
803-816.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
T.A.Steitz,
and
Y.W.Yin
(2004).
Accuracy, lesion bypass, strand displacement and translocation by DNA polymerases.
|
| |
Philos Trans R Soc Lond B Biol Sci,
359,
17-23.
|
 |
|
|
|
|
 |
T.A.Steitz
(2004).
The structural basis of the transition from initiation to elongation phases of transcription, as well as translocation and strand separation, by T7 RNA polymerase.
|
| |
Curr Opin Struct Biol,
14,
4-9.
|
 |
|
|
|
|
 |
Y.W.Yin,
and
T.A.Steitz
(2004).
The structural mechanism of translocation and helicase activity in T7 RNA polymerase.
|
| |
Cell,
116,
393-404.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
A.M.DeLucia,
N.D.Grindley,
and
C.M.Joyce
(2003).
An error-prone family Y DNA polymerase (DinB homolog from Sulfolobus solfataricus) uses a 'steric gate' residue for discrimination against ribonucleotides.
|
| |
Nucleic Acids Res,
31,
4129-4137.
|
 |
|
|
|
|
 |
L.Tsujikawa,
M.Weinfield,
and
L.J.Reha-Krantz
(2003).
Differences in replication of a DNA template containing an ethyl phosphotriester by T4 DNA polymerase and Escherichia coli DNA polymerase I.
|
| |
Nucleic Acids Res,
31,
4965-4972.
|
 |
|
|
|
|
 |
O.Kornyushyna,
and
C.J.Burrows
(2003).
Effect of the oxidized guanosine lesions spiroiminodihydantoin and guanidinohydantoin on proofreading by Escherichia coli DNA polymerase I (Klenow fragment) in different sequence contexts.
|
| |
Biochemistry,
42,
13008-13018.
|
 |
|
 |
 |
|
The most recent references are shown first.
Citation data come partly from CiteXplore and partly
from an automated harvesting procedure. Note that this is likely to be
only a partial list as not all journals are covered by
either method. However, we are continually building up the citation data
so more and more references will be included with time.
Where a reference describes a PDB structure, the PDB
codes are
shown on the right.
|
 |

