 |
PDBsum entry 2fms

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Transferase/DNA
|
PDB id
|
|

|

|
2fms
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
 |
Contents |
 |
|
|

|

|
|
|
|
|
|
|
|
|
|
|
|
* Residue conservation analysis
|
|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 1:
|
 |
E.C.2.7.7.7
- DNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
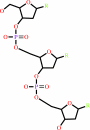 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 2:
|
 |
E.C.4.2.99.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
E.C.4.2.99.18
- DNA-(apurinic or apyrimidinic site) lyase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
2'-deoxyribonucleotide-(2'-deoxyribose 5'-phosphate)- 2'-deoxyribonucleotide-DNA = a 3'-end 2'-deoxyribonucleotide-(2,3- dehydro-2,3-deoxyribose 5'-phosphate)-DNA + a 5'-end 5'-phospho- 2'-deoxyribonucleoside-DNA + H+
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
DOI no:
|
Structure
14:757-766
(2006)
|
|
PubMed id:
|
|
|
|

|
| |
|
Magnesium-induced assembly of a complete DNA polymerase catalytic complex.
|
|
V.K.Batra,
W.A.Beard,
D.D.Shock,
J.M.Krahn,
L.C.Pedersen,
S.H.Wilson.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
The molecular details of the nucleotidyl transferase reaction have remained
speculative, as strategies to trap catalytic intermediates for structure
determination utilize substrates lacking the primer terminus 3'-OH and catalytic
Mg2+, resulting in an incomplete and distorted active site geometry. Since the
geometric arrangement of these essential atoms will impact chemistry, structural
insight into fidelity strategies has been hampered. Here, we present a crystal
structure of a precatalytic complex of a DNA polymerase with bound substrates
that include the primer 3'-OH and catalytic Mg2+. This catalytic intermediate
was trapped with a nonhydrolyzable deoxynucleotide analog. Comparison with two
new structures of DNA polymerase beta lacking the 3'-OH or catalytic Mg2+ is
described. These structures provide direct evidence that both atoms are required
to achieve a proper geometry necessary for an in-line nucleophilic attack of O3'
on the alphaP of the incoming nucleotide.

|

|
|
|

|
| |
Selected figure(s)
|
|

|
| |
 |
 |

|
 |
Figure 7.
Figure 7. Stereoview of the Pol b Active Site
Superimposed structures of pol b with either Na^+ (light blue)
or Mg2+ (yellow) in the catalytic metal site A. Residues that
hydrogen bond (green) to the triphosphate moiety of the incoming
nucleotide and the active site aspartates are shown. When Na^+
occupies the catalytic metal site, O3' of the primer terminus is
3.5 and 4.7 � from the catalytic metal and aP of the incoming
nucleotide, respectively. In contrast, with Mg2+ in the
catalytic metal site, an altered sugar pucker positions O3' of
the primer terminus 2.2 and 3.4 � from the catalytic metal and
Pa of the incoming nucleotide, respectively.
|
 |
|
|

|
| |
The above figure is
reprinted
by permission from Cell Press:
Structure
(2006,
14,
757-766)
copyright 2006.
|
|
| |
Figure was
selected
by the author.
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Literature references that cite this PDB file's key reference
|
|
 |
| |
PubMed id
|
 |
Reference
|
 |
|
|
|
 |
T.Nakamura,
Y.Zhao,
Y.Yamagata,
Y.J.Hua,
and
W.Yang
(2012).
Watching DNA polymerase η make a phosphodiester bond.
|
| |
Nature,
487,
196-201.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
D.L.Ma,
D.S.Chan,
B.Y.Man,
and
C.H.Leung
(2011).
Oligonucleotide-based luminescent detection of metal ions.
|
| |
Chem Asian J,
6,
986.
|
 |
|
|
|
|
 |
G.K.Surya Prakash,
M.Zibinsky,
T.G.Upton,
B.A.Kashemirov,
C.E.McKenna,
K.Oertell,
M.F.Goodman,
V.K.Batra,
L.C.Pedersen,
W.A.Beard,
D.D.Shock,
S.H.Wilson,
and
G.A.Olah
(2010).
Synthesis and biological evaluation of fluorinated deoxynucleotide analogs based on bis-(difluoromethylene)triphosphoric acid.
|
| |
Proc Natl Acad Sci U S A,
107,
15693-15698.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
G.L.Butterfoss,
E.F.DeRose,
S.A.Gabel,
L.Perera,
J.M.Krahn,
G.A.Mueller,
X.Zheng,
and
R.E.London
(2010).
Conformational dependence of 13C shielding and coupling constants for methionine methyl groups.
|
| |
J Biomol NMR,
48,
31-47.
|
 |
|
|
|
|
 |
R.Rucker,
P.Oelschlaeger,
and
A.Warshel
(2010).
A binding free energy decomposition approach for accurate calculations of the fidelity of DNA polymerases.
|
| |
Proteins,
78,
671-680.
|
 |
|
|
|
|
 |
S.H.Wilson,
W.A.Beard,
D.D.Shock,
V.K.Batra,
N.A.Cavanaugh,
R.Prasad,
E.W.Hou,
Y.Liu,
K.Asagoshi,
J.K.Horton,
D.F.Stefanick,
P.S.Kedar,
M.J.Carrozza,
A.Masaoka,
and
M.L.Heacock
(2010).
Base excision repair and design of small molecule inhibitors of human DNA polymerase β.
|
| |
Cell Mol Life Sci,
67,
3633-3647.
|
 |
|
|
|
|
 |
V.K.Batra,
W.A.Beard,
E.W.Hou,
L.C.Pedersen,
R.Prasad,
and
S.H.Wilson
(2010).
Mutagenic conformation of 8-oxo-7,8-dihydro-2'-dGTP in the confines of a DNA polymerase active site.
|
| |
Nat Struct Mol Biol,
17,
889-890.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
A.Irimia,
R.L.Eoff,
F.P.Guengerich,
and
M.Egli
(2009).
Structural and functional elucidation of the mechanism promoting error-prone synthesis by human DNA polymerase kappa opposite the 7,8-dihydro-8-oxo-2'-deoxyguanosine adduct.
|
| |
J Biol Chem,
284,
22467-22480.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
K.Donny-Clark,
R.Shapiro,
and
S.Broyde
(2009).
Accommodation of an N-(deoxyguanosin-8-yl)-2-acetylaminofluorene adduct in the active site of human DNA polymerase iota: Hoogsteen or Watson-Crick base pairing?
|
| |
Biochemistry,
48,
7.
|
 |
|
|
|
|
 |
K.Donny-Clark,
and
S.Broyde
(2009).
Influence of local sequence context on damaged base conformation in human DNA polymerase iota: molecular dynamics studies of nucleotide incorporation opposite a benzo[a]pyrene-derived adenine lesion.
|
| |
Nucleic Acids Res,
37,
7095-7109.
|
 |
|
|
|
|
 |
L.Wang,
S.Broyde,
and
Y.Zhang
(2009).
Polymerase-tailored variations in the water-mediated and substrate-assisted mechanism for nucleotidyl transfer: insights from a study of T7 DNA polymerase.
|
| |
J Mol Biol,
389,
787-796.
|
 |
|
|
|
|
 |
M.Morar,
K.Bhullar,
D.W.Hughes,
M.Junop,
and
G.D.Wright
(2009).
Structure and mechanism of the lincosamide antibiotic adenylyltransferase LinB.
|
| |
Structure,
17,
1649-1659.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
P.Xu,
L.Oum,
Y.C.Lee,
N.E.Geacintov,
and
S.Broyde
(2009).
Visualizing sequence-governed nucleotide selectivities and mutagenic consequences through a replicative cycle: processing of a bulky carcinogen N2-dG lesion in a Y-family DNA polymerase.
|
| |
Biochemistry,
48,
4677-4690.
|
 |
|
|
|
|
 |
S.C.Kamerlin,
C.E.McKenna,
M.F.Goodman,
M.F.Goondman,
and
A.Warshel
(2009).
A computational study of the hydrolysis of dGTP analogues with halomethylene-modified leaving groups in solution: implications for the mechanism of DNA polymerases.
|
| |
Biochemistry,
48,
5963-5971.
|
 |
|
|
|
|
 |
S.M.Sherrer,
J.A.Brown,
L.R.Pack,
V.P.Jasti,
J.D.Fowler,
A.K.Basu,
and
Z.Suo
(2009).
Mechanistic Studies of the Bypass of a Bulky Single-base Lesion Catalyzed by a Y-family DNA Polymerase.
|
| |
J Biol Chem,
284,
6379-6388.
|
 |
|
|
|
|
 |
S.Nakane,
N.Nakagawa,
S.Kuramitsu,
and
R.Masui
(2009).
Characterization of DNA polymerase X from Thermus thermophilus HB8 reveals the POLXc and PHP domains are both required for 3'-5' exonuclease activity.
|
| |
Nucleic Acids Res,
37,
2037-2052.
|
 |
|
|
|
|
 |
T.G.Upton,
B.A.Kashemirov,
C.E.McKenna,
M.F.Goodman,
G.K.Prakash,
R.Kultyshev,
V.K.Batra,
D.D.Shock,
L.C.Pedersen,
W.A.Beard,
and
S.H.Wilson
(2009).
Alpha,beta-difluoromethylene deoxynucleoside 5'-triphosphates: a convenient synthesis of useful probes for DNA polymerase beta structure and function.
|
| |
Org Lett,
11,
1883-1886.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
W.A.Beard,
D.D.Shock,
V.K.Batra,
L.C.Pedersen,
and
S.H.Wilson
(2009).
DNA polymerase beta substrate specificity: side chain modulation of the "A-rule".
|
| |
J Biol Chem,
284,
31680-31689.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
A.Abyzov,
A.Uzun,
P.R.Strauss,
and
V.A.Ilyin
(2008).
An AP endonuclease 1-DNA polymerase beta complex: theoretical prediction of interacting surfaces.
|
| |
PLoS Comput Biol,
4,
e1000066.
|
 |
|
|
|
|
 |
D.F.Zamyatkin,
F.Parra,
J.M.Alonso,
D.A.Harki,
B.R.Peterson,
P.Grochulski,
and
K.K.Ng
(2008).
Structural insights into mechanisms of catalysis and inhibition in Norwalk virus polymerase.
|
| |
J Biol Chem,
283,
7705-7712.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
D.L.Murphy,
J.Kosa,
J.Jaeger,
and
J.B.Sweasy
(2008).
The Asp285 variant of DNA polymerase beta extends mispaired primer termini via increased nucleotide binding.
|
| |
Biochemistry,
47,
8048-8057.
|
 |
|
|
|
|
 |
G.Martin,
S.Doublié,
and
W.Keller
(2008).
Determinants of substrate specificity in RNA-dependent nucleotidyl transferases.
|
| |
Biochim Biophys Acta,
1779,
206-216.
|
 |
|
|
|
|
 |
J.Mendieta,
C.E.Cases-González,
T.Matamoros,
G.Ramírez,
and
L.Menéndez-Arias
(2008).
A Mg2+-induced conformational switch rendering a competent DNA polymerase catalytic complex.
|
| |
Proteins,
71,
565-574.
|
 |
|
|
|
|
 |
L.Jia,
N.E.Geacintov,
and
S.Broyde
(2008).
The N-clasp of human DNA polymerase kappa promotes blockage or error-free bypass of adenine- or guanine-benzo[a]pyrenyl lesions.
|
| |
Nucleic Acids Res,
36,
6571-6584.
|
 |
|
|
|
|
 |
M.Pandey,
S.S.Patel,
and
A.Gabriel
(2008).
Kinetic pathway of pyrophosphorolysis by a retrotransposon reverse transcriptase.
|
| |
PLoS ONE,
3,
e1389.
|
 |
|
|
|
|
 |
P.Lin,
V.K.Batra,
L.C.Pedersen,
W.A.Beard,
S.H.Wilson,
and
L.G.Pedersen
(2008).
Incorrect nucleotide insertion at the active site of a G:A mismatch catalyzed by DNA polymerase beta.
|
| |
Proc Natl Acad Sci U S A,
105,
5670-5674.
|
 |
|
|
|
|
 |
V.K.Batra,
W.A.Beard,
D.D.Shock,
L.C.Pedersen,
and
S.H.Wilson
(2008).
Structures of DNA polymerase beta with active-site mismatches suggest a transient abasic site intermediate during misincorporation.
|
| |
Mol Cell,
30,
315-324.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
W.Yang
(2008).
An equivalent metal ion in one- and two-metal-ion catalysis.
|
| |
Nat Struct Mol Biol,
15,
1228-1231.
|
 |
|
|
|
|
 |
A.F.Moon,
M.Garcia-Diaz,
K.Bebenek,
B.J.Davis,
X.Zhong,
D.A.Ramsden,
T.A.Kunkel,
and
L.C.Pedersen
(2007).
Structural insight into the substrate specificity of DNA Polymerase mu.
|
| |
Nat Struct Mol Biol,
14,
45-53.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
A.F.Moon,
M.Garcia-Diaz,
V.K.Batra,
W.A.Beard,
K.Bebenek,
T.A.Kunkel,
S.H.Wilson,
and
L.C.Pedersen
(2007).
The X family portrait: structural insights into biological functions of X family polymerases.
|
| |
DNA Repair (Amst),
6,
1709-1725.
|
 |
|
|
|
|
 |
C.E.McKenna,
B.A.Kashemirov,
T.G.Upton,
V.K.Batra,
M.F.Goodman,
L.C.Pedersen,
W.A.Beard,
and
S.H.Wilson
(2007).
(R)-beta,gamma-fluoromethylene-dGTP-DNA ternary complex with DNA polymerase beta.
|
| |
J Am Chem Soc,
129,
15412-15413.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
G.C.Lin,
J.Jaeger,
and
J.B.Sweasy
(2007).
Loop II of DNA polymerase beta is important for polymerization activity and fidelity.
|
| |
Nucleic Acids Res,
35,
2924-2935.
|
 |
|
|
|
|
 |
K.A.Fiala,
C.D.Hypes,
and
Z.Suo
(2007).
Mechanism of abasic lesion bypass catalyzed by a Y-family DNA polymerase.
|
| |
J Biol Chem,
282,
8188-8198.
|
 |
|
|
|
|
 |
K.A.Fiala,
J.A.Brown,
H.Ling,
A.K.Kshetry,
J.Zhang,
J.S.Taylor,
W.Yang,
and
Z.Suo
(2007).
Mechanism of template-independent nucleotide incorporation catalyzed by a template-dependent DNA polymerase.
|
| |
J Mol Biol,
365,
590-602.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
L.Wang,
X.Yu,
P.Hu,
S.Broyde,
and
Y.Zhang
(2007).
A water-mediated and substrate-assisted catalytic mechanism for Sulfolobus solfataricus DNA polymerase IV.
|
| |
J Am Chem Soc,
129,
4731-4737.
|
 |
|
|
|
|
 |
M.Garcia-Diaz,
K.Bebenek,
J.M.Krahn,
L.C.Pedersen,
and
T.A.Kunkel
(2007).
Role of the catalytic metal during polymerization by DNA polymerase lambda.
|
| |
DNA Repair (Amst),
6,
1333-1340.
|
 |
|
PDB codes:
|
 |
|
|
|
|
|
 |
M.Garcia-Diaz,
and
K.Bebenek
(2007).
Multiple functions of DNA polymerases.
|
| |
CRC Crit Rev Plant Sci,
26,
105-122.
|
 |
|
|
|
|
 |
P.B.Balbo,
and
A.Bohm
(2007).
Mechanism of poly(A) polymerase: structure of the enzyme-MgATP-RNA ternary complex and kinetic analysis.
|
| |
Structure,
15,
1117-1131.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
P.Oelschlaeger,
M.Klahn,
W.A.Beard,
S.H.Wilson,
and
A.Warshel
(2007).
Magnesium-cationic dummy atom molecules enhance representation of DNA polymerase beta in molecular dynamics simulations: improved accuracy in studies of structural features and mutational effects.
|
| |
J Mol Biol,
366,
687-701.
|
 |
|
|
|
|
 |
P.Xu,
L.Oum,
L.S.Beese,
N.E.Geacintov,
and
S.Broyde
(2007).
Following an environmental carcinogen N2-dG adduct through replication: elucidating blockage and bypass in a high-fidelity DNA polymerase.
|
| |
Nucleic Acids Res,
35,
4275-4288.
|
 |
|
|
|
|
 |
Y.Wang,
S.Reddy,
W.A.Beard,
S.H.Wilson,
and
T.Schlick
(2007).
Differing conformational pathways before and after chemistry for insertion of dATP versus dCTP opposite 8-oxoG in DNA polymerase beta.
|
| |
Biophys J,
92,
3063-3070.
|
 |
|
|
|
|
 |
L.Zhang,
O.Rechkoblit,
L.Wang,
D.J.Patel,
R.Shapiro,
and
S.Broyde
(2006).
Mutagenic nucleotide incorporation and hindered translocation by a food carcinogen C8-dG adduct in Sulfolobus solfataricus P2 DNA polymerase IV (Dpo4): modeling and dynamics studies.
|
| |
Nucleic Acids Res,
34,
3326-3337.
|
 |
|
|
|
|
 |
M.C.Foley,
K.Arora,
and
T.Schlick
(2006).
Sequential side-chain residue motions transform the binary into the ternary state of DNA polymerase lambda.
|
| |
Biophys J,
91,
3182-3195.
|
 |
|
|
|
|
 |
R.Radhakrishnan,
K.Arora,
Y.Wang,
W.A.Beard,
S.H.Wilson,
and
T.Schlick
(2006).
Regulation of DNA repair fidelity by molecular checkpoints: "gates" in DNA polymerase beta's substrate selection.
|
| |
Biochemistry,
45,
15142-15156.
|
 |
|
|
|
|
 |
W.W.Duym,
K.A.Fiala,
N.Bhatt,
and
Z.Suo
(2006).
Kinetic effect of a downstream strand and its 5'-terminal moieties on single nucleotide gap-filling synthesis catalyzed by human DNA polymerase lambda.
|
| |
J Biol Chem,
281,
35649-35655.
|
 |
|
 |
 |
|
The most recent references are shown first.
Citation data come partly from CiteXplore and partly
from an automated harvesting procedure. Note that this is likely to be
only a partial list as not all journals are covered by
either method. However, we are continually building up the citation data
so more and more references will be included with time.
Where a reference describes a PDB structure, the PDB
codes are
shown on the right.
|
 |

