Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
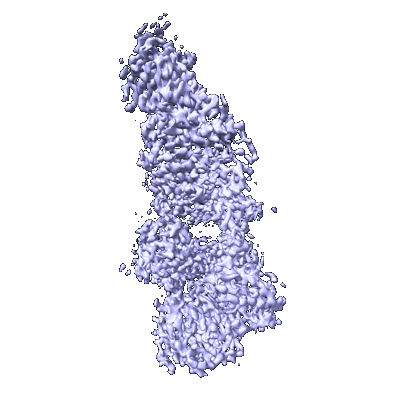
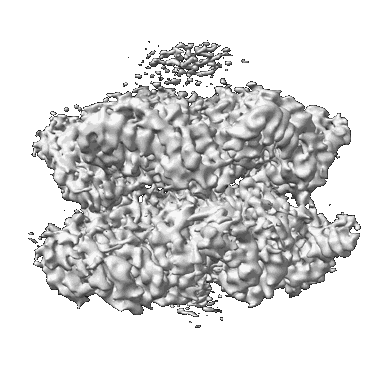
Chaetomium thermophilum SETX - NPPC internal deletion
Resolution: 3.17 Å
EM Method: Single-particle
Fitted PDBs: 8fth
Q-score: 0.479
Appel CD, Bermek O, Dandey VP, Wood M, Viverette E, Williams JG, Bouvette J, Riccio AA, Krahn JM, Borgnia MJ, Williams RS
Mol Cell (2023) 83 pp. 3692- [ DOI: doi:10.1016/j.molcel.2023.09.024 Pubmed: 37832548 ]
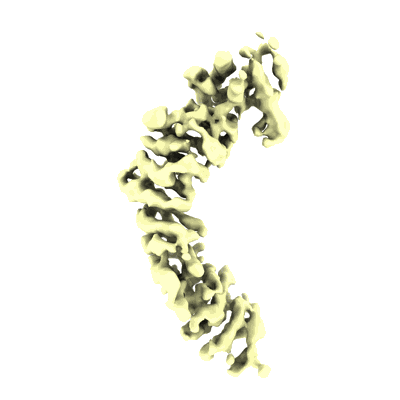
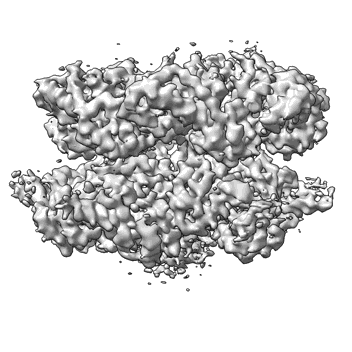
Chaetomium thermophilum SETX (Full-length)
Resolution: 4.56 Å
EM Method: Single-particle
Fitted PDBs: 8ftk
Q-score: 0.288
Appel CD, Bermek O, Dandey VP, Wood M, Viverette E, Williams JG, Bouvette J, Riccio AA, Krahn JM, Borgnia MJ, Williams RS
Mol Cell (2023) 83 pp. 3692- [ DOI: doi:10.1016/j.molcel.2023.09.024 Pubmed: 37832548 ]
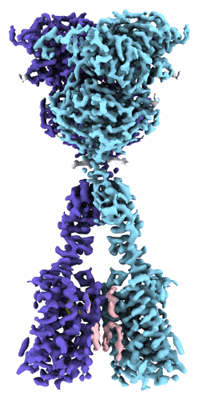
Appel CD, Bermek O, Dandey VP, Wood M, Viverette E, Williams JG, Bouvette J, Riccio AA, Krahn JM, Borgnia MJ, Williams RS
Mol Cell (2023) 83 pp. 3692-3706.e5 [ DOI: doi:10.1016/j.molcel.2023.09.024 Pubmed: 37832548 ]
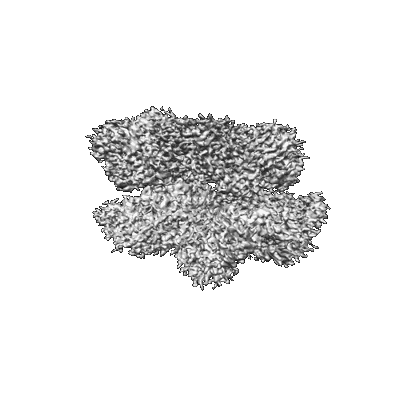
Structures of positive allosteric modulator-bound and unbound active human calcium-sensing receptor
Park J, Zuo H, Frangaj A, Fu Z, Yen LY, Zhang Z, Mosyak L, Slavkovich VN, Liu J, Ray KM, Cao B, Vallese F, Geng Y, Chen S, Grassucci R, Dandey VP, Tan YZ, Eng E, Lee Y, Kloss B, Liu Z, Hendrickson WA, Potter CS, Carragher B, Graziano J, Conigrave AD, Frank J, Clarke OB, Fan QR
Proc Natl Acad Sci U S A (2021) 118 [ DOI: 10.1073/pnas.2115849118 Pubmed: 34916296 ]
Image sets:
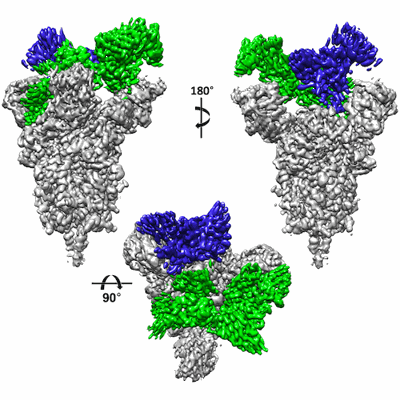
Camel nanobodies 7A3 and 8A2 broadly neutralize SARS-CoV-2 variants
Resolution: 2.39 Å
EM Method: Single-particle
Fitted PDBs: 7tpr
Q-score: 0.407
Hong J, Kwon HJ, Cachau R, Chen CZ, Dandey VP, Martin N, Esposito D, Ortega-Rodriguez U, Xu M, Borgnia MJ, Xie H, Ho M
bioRxiv (2021) [ DOI: doi:10.1101/2021.10.27.465996 Pubmed: 34751270 ]
- Camel nanobody 7a3 (Complex from Camelus dromedarius)
- Nanobody 7a3 (13 kDa, Protein from Camelus dromedarius)
- Spike glycoprotein (126 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Nanobody 8a2 (13 kDa, Protein from Camelus dromedarius)
- Sars-cov-2 spike glycoprotein (Complex from Severe acute respiratory syndrome coronavirus 2)
- Camel nanobody 8a2 (Complex from Camelus dromedarius)
- Ternary complex of sars-cov-2 spike protein ectodomain (wuhan strain) and camel nanobodies 7a3 and 8a2. (17 kDa, Complex)
Structure of negative allosteric modulator-bound inactive human calcium-sensing receptor
Park J, Zuo H, Frangaj A, Fu Z, Yen LY, Zhang Z, Mosyak L, Slavkovich VN, Liu J, Ray KM, Cao B, Vallese F, Geng Y, Chen S, Grassucci R, Dandey VP, Tan YZ, Eng E, Lee Y, Kloss B, Liu Z, Hendrickson WA, Potter CS, Carragher B, Graziano J, Conigrave AD, Frank J, Clarke OB, Fan QR
Proc Natl Acad Sci U S A (2021) 118 [ DOI: 10.1073/pnas.2115849118 Pubmed: 34916296 ]
Image sets:
CryoEM structure of the N-terminal-deleted Rix7 AAA-ATPase
Resolution: 2.88 Å
EM Method: Single-particle
Fitted PDBs: 7swl
Q-score: 0.546
Kocaman S, Lo YH, Krahn JM, Sobhany M, Dandey VP, Petrovich ML, Etigunta SK, Williams JG, Deterding LJ, Borgnia MJ, Stanley RE
Pnas Nexus (2022) 1 pp. pgac118-pgac118 [ DOI: doi:10.1093/pnasnexus/pgac118 Pubmed: 36090660 ]
- Polyleucine (2 kDa, Protein from synthetic construct)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Rix7 (69 kDa, Protein from Chaetomium thermophilum)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Cryoem structure of the n-terminal-deleted rix7 aaa-atpase (100 kDa, Complex from Chaetomium thermophilum)
CryoEM structure of the crosslinked Rix7 AAA-ATPase
Resolution: 3.67 Å
EM Method: Single-particle
Fitted PDBs: 7t0v
Q-score: 0.381
Kocaman S, Lo YH, Krahn JM, Sobhany M, Dandey VP, Petrovich ML, Etigunta SK, Williams JG, Deterding LJ, Borgnia MJ, Stanley RE
Pnas Nexus (2022) 1 pp. pgac118-pgac118 [ DOI: doi:10.1093/pnasnexus/pgac118 Pubmed: 36090660 ]
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Rix7 (89 kDa, Protein from Chaetomium thermophilum)
- Magnesium ion (24 Da, Ligand)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Polyvaline (2 kDa, Protein from synthetic construct)
- Cryoem structure of the crosslinked rix7 aaa-atpase (100 kDa, Complex from Chaetomium thermophilum)
CryoEM structure of the Rix7 D2 Walker B mutant
Resolution: 4.3 Å
EM Method: Single-particle
Fitted PDBs: 7t3i
Q-score: 0.38
Kocaman S, Lo YH, Krahn JM, Sobhany M, Dandey VP, Petrovich ML, Etigunta SK, Williams JG, Deterding LJ, Borgnia MJ, Stanley RE
Pnas Nexus (2022) 1 pp. pgac118-pgac118 [ DOI: doi:10.1093/pnasnexus/pgac118 Pubmed: 36090660 ]
- Cryoem structure of rix7 d2 walker b mutant (Complex from Chaetomium thermophilum)
- Phosphate ion (94 Da, Ligand)
- Substrate peptide (2 kDa, Protein from Escherichia coli)
- Rix7 (89 kDa, Protein from Chaetomium thermophilum)
- Adenosine-5'-triphosphate (507 Da, Ligand)
Structure of positive allosteric modulator-bound active human calcium-sensing receptor
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 7sil
Q-score: 0.562
Park J, Zuo H, Frangaj A, Fu Z, Yen LY, Zhang Z, Mosyak L, Slavkovich VN, Liu J, Ray KM, Cao B, Vallese F, Geng Y, Chen S, Grassucci R, Dandey VP, Tan YZ, Eng E, Lee Y, Kloss B, Liu Z, Hendrickson WA, Potter CS, Carragher B, Graziano J, Conigrave AD, Frank J, Clarke OB, Fan QR
PNAS (2021) 118 [ DOI: doi:10.1073/pnas.2115849118 Pubmed: 34916296 ]
- Cyclomethyltryptophan (216 Da, Ligand)
- Calcium ion (40 Da, Ligand)
- Phosphate ion (94 Da, Ligand)
- Cholesterol (386 Da, Ligand)
- 3-(2-chlorophenyl)-n-[(1r)-1-(3-methoxyphenyl)ethyl]propan-1-amine (303 Da, Ligand)
- Homodimer human calcium sensing receptor (192 kDa, Complex from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Isoform 1 of extracellular calcium-sensing receptor (99 kDa, Protein from Homo sapiens)