Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
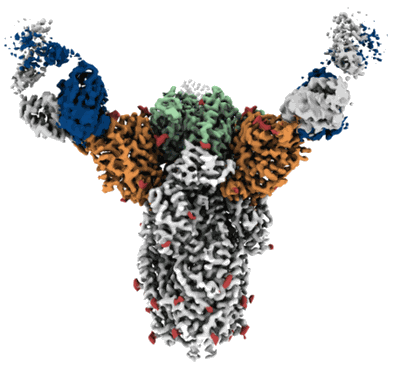
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 7rw2
Q-score: 0.386
Cerutti G, Guo Y, Wang P, Nair MS, Wang M, Huang Y, Yu J, Liu L, Katsamba PS, Bahna F, Reddem ER, Kwong PD, Ho DD, Sheng Z, Shapiro L
Cell Rep (2021) 37 pp. 109928-109928 [ Pubmed: 34706271 DOI: doi:10.1016/j.celrep.2021.109928 ]
- 5-7 heavy chain (27 kDa, Protein from Homo sapiens)
- 5-7 light chain (23 kDa, Protein from Homo sapiens)
- Prefusion sars-cov-2 spike glycoprotein in complex with 5-7 fab (Complex from Severe acute respiratory syndrome coronavirus 2)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
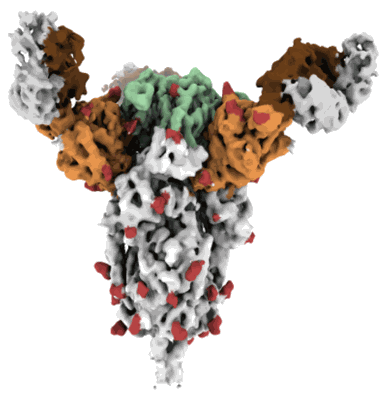
Resolution: 2.97 Å
EM Method: Single-particle
Fitted PDBs: 7l2e
Q-score: 0.505
Cerutti G, Guo Y, Zhou T, Gorman J, Lee M, Rapp M, Reddem ER, Yu J, Bahna F, Bimela J, Huang Y, Katsamba PS, Liu L, Nair MS, Rawi R, Olia AS, Wang P, Zhang B, Chuang GY, Ho DD, Sheng Z, Kwong PD, Shapiro L
Cell Host Microbe (2021) 29 pp. 819- [ Pubmed: 33789084 DOI: doi:10.1016/j.chom.2021.03.005 ]
- Prefusion sars-cov-2 spike glycoprotein in complex with 4-18 fab (Complex from Homo sapiens)
- 4-18 light chain (23 kDa, Protein from Homo sapiens)
- 4-18 heavy chain (24 kDa, Protein from Homo sapiens)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
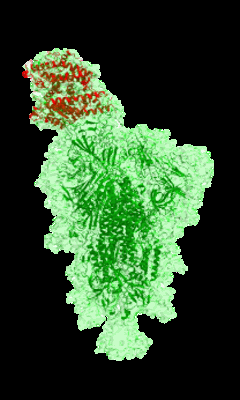
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 7l2f
Q-score: 0.332
Cerutti G, Guo Y, Zhou T, Gorman J, Lee M, Rapp M, Reddem ER, Yu J, Bahna F, Bimela J, Huang Y, Katsamba PS, Liu L, Nair MS, Rawi R, Olia AS, Wang P, Zhang B, Chuang GY, Ho DD, Sheng Z, Kwong PD, Shapiro L
Cell Host Microbe (2021) 29 pp. 819- [ Pubmed: 33789084 DOI: doi:10.1016/j.chom.2021.03.005 ]
- 5-24 light chain (23 kDa, Protein from Homo sapiens)
- 5-24 heavy chain (25 kDa, Protein from Homo sapiens)
- Prefusion sars-cov-2 spike glycoprotein in complex with 5-24 fab (Complex from Homo sapiens)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
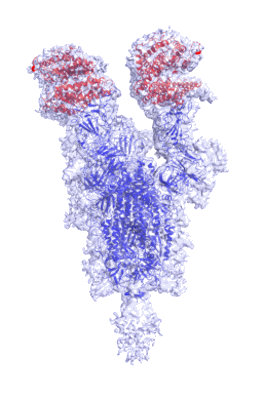
Cryo-EM structure of NTD-directed neutralizing antibody 4-8 Fab in complex with SARS-CoV-2 S2P spike
Resolution: 3.25 Å
EM Method: Single-particle
Fitted PDBs: 7lqv
Q-score: 0.318
Cerutti G, Guo Y, ZHou T, Gorman J, Lee M, Rapp M, Reddem ER, Yu J, Bahna F, Bimela J, Huang Y, Katsamba PS, Liu L, Nair MS, Rawi R, Olia AS, Wang P, Zhang B, Chuang GY, Ho DD, Sheng Z, Kwong PD, Shapiro L
Cell Host Microbe (2021) [ DOI: doi:10.1016/j.chom.2021.03.005 ]
- 4-8 fab in complex with sars-cov-2 s2p spike (Complex from Severe acute respiratory syndrome coronavirus 2)
- Spike glycoprotein (144 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 4-8 heavy chain (13 kDa, Protein from Homo sapiens)
- 4-8 light chain (11 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
Cryo-EM structure of single ACE2-bound SARS-CoV-2 trimer spike at pH 5.5
Resolution: 3.85 Å
EM Method: Single-particle
Fitted PDBs: 7kne
Q-score: 0.253
Zhou T, Tsybovsky Y, Gorman J, Rapp M, Cerutti G, Chuang GY, Katsamba PS, Sampson JM, Schon A, Bimela J, Boyington JC, Nazzari A, Olia AS, Shi W, Sastry M, Stephens T, Stuckey J, Teng IT, Wang P, Wang S, Zhang B, Friesner RA, Ho DD, Mascola JR, Shapiro L, Kwong PD
Cell Host Microbe (2020) 28 pp. 867-879.e5 [ DOI: doi:10.1016/j.chom.2020.11.004 Pubmed: 33271067 ]
- Angiotensin-converting enzyme 2 (69 kDa, Protein from Homo sapiens)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Single ace2-bound sars-cov-2 trimer spike at ph 5.5 (Complex from Severe acute respiratory syndrome coronavirus 2)
Cryo-EM structure of double ACE2-bound SARS-CoV-2 trimer Spike at pH 7.4
Resolution: 3.62 Å
EM Method: Single-particle
Fitted PDBs: 7kmz
Q-score: 0.285
Zhou T, Tsybovsky Y, Gorman J, Rapp M, Cerutti G, Chuang GY, Katsamba PS, Sampson JM, Schon A, Bimela J, Boyington JC, Nazzari A, Olia AS, Shi W, Sastry M, Stephens T, Stuckey J, Teng IT, Wang P, Wang S, Zhang B, Friesner RA, Ho DD, Mascola JR, Shapiro L, Kwong PD
Cell Host Microbe (2020) 28 pp. 867-879.e5 [ DOI: doi:10.1016/j.chom.2020.11.004 Pubmed: 33271067 ]
- Angiotensin-converting enzyme 2 (69 kDa, Protein from Homo sapiens)
- Cryo-em structure of double ace2-bound sars-cov-2 trimer spike at ph 7.4 (Complex from Severe acute respiratory syndrome coronavirus 2)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)