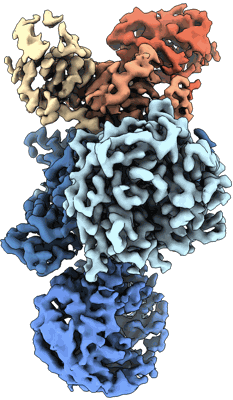
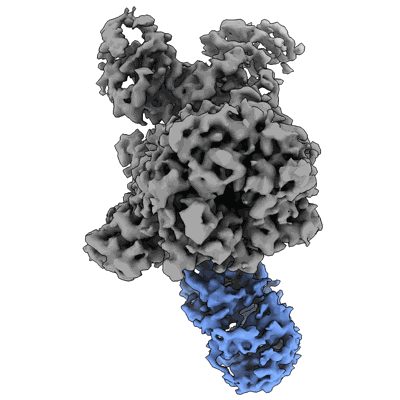
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8cvp
Q-score: 0.465
Watson ER, Novick S, Matyskiela ME, Chamberlain PP, H de la Pena A, Zhu J, Tran E, Griffin PR, Wertz IE, Lander GC
Science (2022) 378 pp. 549-553 [ DOI: doi:10.1126/science.add7574 Pubmed: 36378961 ]
- Cereblon~ddb1 in the apo form (177 kDa, Complex from Homo sapiens)
- Zinc ion (65 Da, Ligand)
- Dna damage-binding protein 1 (127 kDa, Protein from Homo sapiens)
- Protein cereblon (50 kDa, Protein from Homo sapiens)
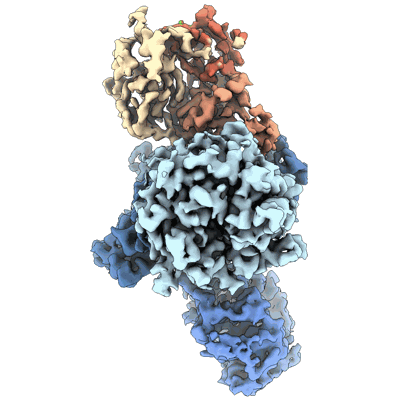
Cereblon~DDB1 bound to CC-92480 with DDB1 in the linear conformation
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 8d7u
Q-score: 0.515
Watson ER, Novick S, Matyskiela ME, Chamberlain PP, H de la Pena A, Zhu J, Tran E, Griffin PR, Wertz IE, Lander GC
Science (2022) 378 pp. 549-553 [ DOI: doi:10.1126/science.add7574 Pubmed: 36378961 ]
- Cereblon~ddb1 bound to cc-92480 with ddb1 in the linear conformation (177 kDa, Complex from Homo sapiens)
- Protein cereblon (50 kDa, Protein from Homo sapiens)
- Zinc ion (65 Da, Ligand)
- Mezigdomide (567 Da, Ligand)
- Dna damage-binding protein 1 (127 kDa, Protein from Homo sapiens)
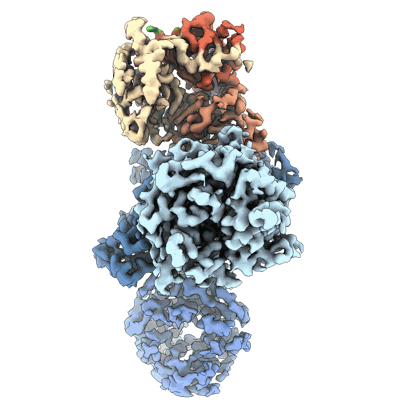
Cereblon~DDB1 bound to CC-92480 with DDB1 in the twisted conformation
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 8d7v
Q-score: 0.45
Watson ER, Novick S, Matyskiela ME, Chamberlain PP, H de la Pena A, Zhu J, Tran E, Griffin PR, Wertz IE, Lander GC
Science (2022) 378 pp. 549-553 [ DOI: doi:10.1126/science.add7574 Pubmed: 36378961 ]
- Cereblon~ddb1 bound to cc-92480 with ddb1 in the twisted conformation (177 kDa, Complex from Homo sapiens)
- Protein cereblon (50 kDa, Protein from Homo sapiens)
- Zinc ion (65 Da, Ligand)
- Mezigdomide (567 Da, Ligand)
- Dna damage-binding protein 1 (127 kDa, Protein from Homo sapiens)
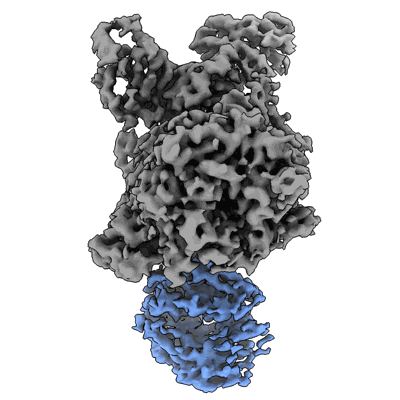
Cereblon~DDB1 bound to CC-92480 with DDB1 in the hinged conformation
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 8d7w
Q-score: 0.476
Watson ER, Novick S, Matyskiela ME, Chamberlain PP, H de la Pena A, Zhu J, Tran E, Griffin PR, Wertz IE, Lander GC
Science (2022) 378 pp. 549-553 [ DOI: doi:10.1126/science.add7574 Pubmed: 36378961 ]
- Protein cereblon (50 kDa, Protein from Homo sapiens)
- Cereblon~ddb1 bound to cc-92480 with ddb1 in the hinged conformation (177 kDa, Complex from Homo sapiens)
- Zinc ion (65 Da, Ligand)
- Mezigdomide (567 Da, Ligand)
- Dna damage-binding protein 1 (127 kDa, Protein from Homo sapiens)
Cereblon~DDB1 in the Apo form with DDB1 in the hinged conformation
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8d7x
Q-score: 0.488
Watson ER, Novick S, Matyskiela ME, Chamberlain PP, H de la Pena A, Zhu J, Tran E, Griffin PR, Wertz IE, Lander GC
Science (2022) 378 pp. 549-553 [ DOI: doi:10.1126/science.add7574 Pubmed: 36378961 ]
- Cereblon~ddb1 in the apo form with ddb1 in the hinged conformation (177 kDa, Complex from Homo sapiens)
- Zinc ion (65 Da, Ligand)
- Dna damage-binding protein 1 (127 kDa, Protein from Homo sapiens)
- Protein cereblon (50 kDa, Protein from Homo sapiens)
Cereblon-DDB1 in the Apo form with DDB1 in the twisted conformation
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8d7y
Q-score: 0.453
Watson ER, Novick S, Matyskiela ME, Chamberlain PP, H de la Pena A, Zhu J, Tran E, Griffin PR, Wertz IE, Lander GC
Science (2022) 378 pp. 549-553 [ DOI: doi:10.1126/science.add7574 Pubmed: 36378961 ]
- Protein cereblon (50 kDa, Protein from Homo sapiens)
- Zinc ion (65 Da, Ligand)
- Cereblon~ddb1 in the apo form with ddb1 in twisted conformation (177 kDa, Complex from Homo sapiens)
- Dna damage-binding protein 1 (127 kDa, Protein from Homo sapiens)
Cereblon~DDB1 bound to Pomalidomide
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 8d81
Q-score: 0.436
Watson ER, Novick S, Matyskiela ME, Chamberlain PP, H de la Pena A, Zhu J, Tran E, Griffin PR, Wertz IE, Lander GC
Science (2022) 378 pp. 549-553 [ DOI: doi:10.1126/science.add7574 Pubmed: 36378961 ]
- Cereblon~ddb1 bound to pomalidomide (143 kDa, Complex from Homo sapiens)
- S-pomalidomide (273 Da, Ligand)
- Protein cereblon (50 kDa, Protein from Homo sapiens)
- Dna damage-binding protein 1 (93 kDa, Protein from Homo sapiens)
- Zinc ion (65 Da, Ligand)
ATP-bound AMP-activated protein kinase
Resolution: 3.93 Å
EM Method: Single-particle
Fitted PDBs: 7m74
Q-score: 0.27
Yan Y, Mukherjee S, Harikumar KG, Strutzenberg TS, Zhou XE, Suino-Powell K, Xu TH, Sheldon RD, Lamp J, Brunzelle JS, Radziwon K, Ellis A, Novick SJ, Vega IE, Jones RG, Miller LJ, Xu HE, Griffin PR, Kossiakoff AA, Melcher K
Science (2021) 373 pp. 413-419 [ DOI: doi:10.1126/science.abe7565 Pubmed: 34437114 ]
- 5'-amp-activated protein kinase subunit beta-2 (22 kDa, Protein from Homo sapiens)
- Adenosine monophosphate (347 Da, Ligand)
- Fab light chain (22 kDa, Protein from Homo sapiens)
- Nanobody (13 kDa, Protein from Lama glama)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Fab heavy chain (24 kDa, Protein from Homo sapiens)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- 5'-amp-activated protein kinase subunit gamma-1 (34 kDa, Protein from Homo sapiens)
- 6-[4-(2-piperidin-1-ylethoxy)phenyl]-3-pyridin-4-ylpyrazolo[1,5-a]pyrimidine (399 Da, Ligand)
- Mbp-fused atp bound ampk in complex with c-compound stabilized by fab and a nanobody (Complex from Homo sapiens)
- Maltose/maltodextrin abc transporter substrate-binding protein male (40 kDa, Protein from Escherichia coli)
- 5'-amp-activated protein kinase catalytic subunit alpha-1 (56 kDa, Protein from Homo sapiens)
Cryo-EM structure of human GPR158
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7she
Q-score: 0.352
Patil DN, Singh S, Laboute T, Strutzenberg TS, Qiu X, Wu D, Novick SJ, Robinson CV, Griffin PR, Hunt JF, Izard T, Singh AK, Martemyanov KA
Science (2022) 375 pp. 86-91 [ DOI: doi:10.1126/science.abl4732 Pubmed: 34793198 ]
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- G-protein coupled receptor 158 (88 kDa, Protein from Homo sapiens)
- Cholesterol (386 Da, Ligand)
- (2s)-1-{[(s)-hydroxy{[(1s,2r,3r,4r,5s,6s)-2,3,4,5,6-pentahydroxycyclohexyl]oxy}phosphoryl]oxy}-3-(octadecanoyloxy)propan-2-yl (5e,8e,11e,14e)-icosa-5,8,11,14-tetraenoate (887 Da, Ligand)
- Receptor (170 kDa, Complex from Homo sapiens)
Cryo-EM structure of GPR158 coupled to the RGS7-Gbeta5 complex
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7shf
Q-score: 0.478
Patil DN, Singh S, Laboute T, Strutzenberg TS, Qiu X, Wu D, Novick SJ, Robinson CV, Griffin PR, Hunt JF, Izard T, Singh AK, Martemyanov KA
Science (2022) 375 pp. 86-91 [ DOI: doi:10.1126/science.abl4732 Pubmed: 34793198 ]
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Isoform 2 of regulator of g-protein signaling 7 (54 kDa, Protein from Homo sapiens)
- Guanine nucleotide-binding protein subunit beta-5 (38 kDa, Protein from Mus musculus)
- G-protein coupled receptor 158 (88 kDa, Protein from Homo sapiens)
- Cholesterol (386 Da, Ligand)
- (2s)-1-{[(s)-hydroxy{[(1s,2r,3r,4r,5s,6s)-2,3,4,5,6-pentahydroxycyclohexyl]oxy}phosphoryl]oxy}-3-(octadecanoyloxy)propan-2-yl (5e,8e,11e,14e)-icosa-5,8,11,14-tetraenoate (887 Da, Ligand)
- Gpcr complex (270 kDa, Complex from Homo sapiens)