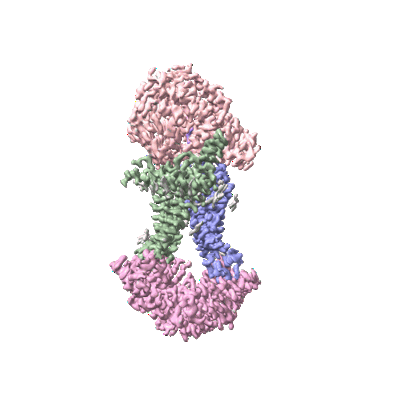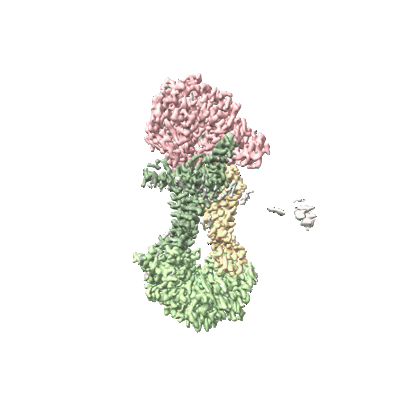Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the pre-translocation state
Resolution: 3.25 Å
EM Method: Single-particle
Fitted PDBs: 8j5q
Q-score: 0.519
Yang X, Hu T, Liang J, Xiong Z, Lin Z, Zhao Y, Zhou X, Gao Y, Sun S, Yang X, Guddat LW, Yang H, Rao Z, Zhang B
Nat Struct Mol Biol (2024) [ Pubmed: 38548954 DOI: doi:10.1038/s41594-024-01256-z ]
- Iron/sulfur cluster (351 Da, Ligand)
- Uncharacterized abc transporter atp-binding protein rv1281c (65 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Cryo-em structure of mycobacterium tuberculosis oppabcd in the pre-translocation state (Complex from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1283c (35 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Uncharacterized protein rv1280c (64 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1282c (31 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Endogenous oligopeptide (783 Da, Protein from Mycolicibacterium smegmatis MC2 155)
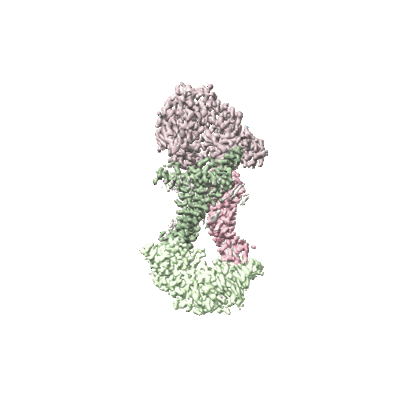
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the resting state
Resolution: 3.28 Å
EM Method: Single-particle
Fitted PDBs: 8j5r
Q-score: 0.534
Yang X, Hu T, Liang J, Xiong Z, Lin Z, Zhao Y, Zhou X, Gao Y, Sun S, Yang X, Guddat LW, Yang H, Rao Z, Zhang B
Nat Struct Mol Biol (2024) [ Pubmed: 38548954 DOI: doi:10.1038/s41594-024-01256-z ]
- Iron/sulfur cluster (351 Da, Ligand)
- Uncharacterized abc transporter atp-binding protein rv1281c (65 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1283c (35 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Uncharacterized protein rv1280c (64 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1282c (31 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Cryo-em structure of mycobacterium tuberculosis oppabcd in the resting state (Complex from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the pre-catalytic intermediate state
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 8j5s
Q-score: 0.566
Yang X, Hu T, Liang J, Xiong Z, Lin Z, Zhao Y, Zhou X, Gao Y, Sun S, Yang X, Guddat LW, Yang H, Rao Z, Zhang B
Nat Struct Mol Biol (2024) [ Pubmed: 38548954 DOI: doi:10.1038/s41594-024-01256-z ]
- Iron/sulfur cluster (351 Da, Ligand)
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)
- Endogenous oligopeptide (698 Da, Protein from Mycolicibacterium smegmatis MC2 155)
- Putative peptide transport permease protein rv1283c (35 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Uncharacterized protein rv1280c (64 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Cryo-em structure of mycobacterium tuberculosis oppabcd in the pre-catalytic intermediate state (Complex from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1282c (31 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Magnesium ion (24 Da, Ligand)
- Uncharacterized abc transporter atp-binding protein rv1281c (65 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
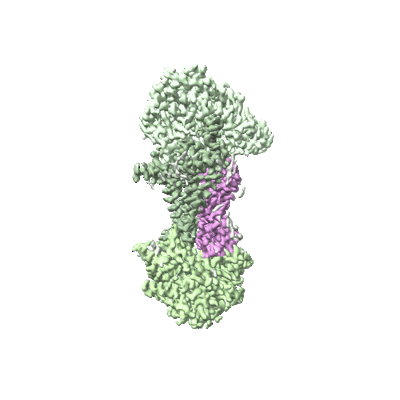
Cryo-EM structure of Mycobacterium tuberculosis OppABCD in the catalytic intermediate state
Resolution: 2.98 Å
EM Method: Single-particle
Fitted PDBs: 8j5t
Q-score: 0.572
Yang X, Hu T, Liang J, Xiong Z, Lin Z, Zhao Y, Zhou X, Gao Y, Sun S, Yang X, Guddat LW, Yang H, Rao Z, Zhang B
Nat Struct Mol Biol (2024) [ Pubmed: 38548954 DOI: doi:10.1038/s41594-024-01256-z ]
- Iron/sulfur cluster (351 Da, Ligand)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Cryo-em structure of mycobacterium tuberculosis oppabcd in the catalytic intermediate state (Complex from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1283c (35 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Uncharacterized protein rv1280c (64 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Putative peptide transport permease protein rv1282c (31 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
- Magnesium ion (24 Da, Ligand)
- Uncharacterized abc transporter atp-binding protein rv1281c (65 kDa, Protein from Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv))
cryo-EM structure of the human respiratory complex II
Resolution: 2.86 Å
EM Method: Single-particle
Fitted PDBs: 8gs8
Q-score: 0.577
Du Z, Zhou X, Lai Y, Xu J, Zhang Y, Zhou S, Feng Z, Yu L, Tang Y, Wang W, Yu L, Tian C, Ran T, Chen H, Guddat LW, Liu F, Gao Y, Rao Z, Gong H
PNAS (2023) 120 pp. e2216713120-e2216713120 [ DOI: doi:10.1073/pnas.2216713120 Pubmed: 37098072 ]
- Iron/sulfur cluster (351 Da, Ligand)
- Succinate dehydrogenase [ubiquinone] iron-sulfur subunit, mitochondrial (31 kDa, Protein from Homo sapiens)
- Fe3-s4 cluster (295 Da, Ligand)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Ubiquinone-1 (250 Da, Ligand)
- Protoporphyrin ix containing fe (616 Da, Ligand)
- Succinate dehydrogenase [ubiquinone] cytochrome b small subunit, mitochondrial (17 kDa, Protein from Homo sapiens)
- (1s)-2-{[(2-aminoethoxy)(hydroxy)phosphoryl]oxy}-1-[(palmitoyloxy)methyl]ethyl stearate (720 Da, Ligand)
- Succinate dehydrogenase cytochrome b560 subunit, mitochondrial (18 kDa, Protein from Homo sapiens)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- The human mitochondrial complex ii, composed of sdha, sdhb, sdhc and sdhd (Complex from Homo sapiens)
- Succinate dehydrogenase [ubiquinone] flavoprotein subunit, mitochondrial (72 kDa, Protein from Homo sapiens)
Cryo-EM structure of Mycobacterium tuberculosis irtAB in inward-facing state
Resolution: 3.48 Å
EM Method: Single-particle
Fitted PDBs: 7wiu
Q-score: 0.476
Cryo-EM structure of Mycobacterium tuberculosis irtAB in complex with an AMP-PNP
Resolution: 2.88 Å
EM Method: Single-particle
Fitted PDBs: 7wiv
Q-score: 0.577
Sun S, Gao Y, Yang X, Yang X, Hu T, Liang J, Xiong Z, Ran Y, Ren P, Bai F, Guddat LW, Yang H, Rao Z, Zhang B
Protein Cell (2023) 14 pp. 448-458 [ Pubmed: 36882106 DOI: doi:10.1093/procel/pwac060 ]
- Irtab (Complex from Mycobacterium tuberculosis H37Rv)
- Mycobactin import atp-binding/permease protein irta (93 kDa, Protein from Mycobacterium tuberculosis H37Rv)
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)
- Mycobactin import atp-binding/permease protein irtb (61 kDa, Protein from Mycobacterium tuberculosis H37Rv)
Cryo-EM structure of Mycobacterium tuberculosis irtAB complexed with ATP in an occluded conformation
Resolution: 3.12 Å
EM Method: Single-particle
Fitted PDBs: 7wiw
Q-score: 0.56
Sun S, Gao Y, Yang X, Yang X, Hu T, Liang J, Xiong Z, Ran Y, Ren P, Bai F, Guddat LW, Yang H, Rao Z, Zhang B
Protein Cell (2023) 14 pp. 448-458 [ Pubmed: 36882106 DOI: doi:10.1093/procel/pwac060 ]
- Irtab (Complex from Mycobacterium tuberculosis H37Rv)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Mycobactin import atp-binding/permease protein irtb (61 kDa, Protein from Mycobacterium tuberculosis H37Rv)
- Magnesium ion (24 Da, Ligand)
- Mycobactin import atp-binding/permease protein irta (93 kDa, Protein from Mycobacterium tuberculosis H37Rv)
Cryo-EM structure of Mycobacterium tuberculosis irtAB in complex with ADP
Resolution: 3.53 Å
EM Method: Single-particle
Fitted PDBs: 7wix
Q-score: 0.48
Sun S, Gao Y, Yang X, Yang X, Hu T, Liang J, Xiong Z, Ran Y, Ren P, Bai F, Guddat LW, Yang H, Rao Z, Zhang B
Protein Cell (2023) 14 pp. 448-458 [ Pubmed: 36882106 DOI: doi:10.1093/procel/pwac060 ]
- Irtab (Complex from Mycobacterium tuberculosis H37Rv)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Mycobactin import atp-binding/permease protein irta (93 kDa, Protein from Mycobacterium tuberculosis H37Rv)
- Mycobactin import atp-binding/permease protein irtb (61 kDa, Protein from Mycobacterium tuberculosis H37Rv)
Mycobacterium smegmatis Sdh1 in complex with UQ1
Resolution: 2.53 Å
EM Method: Single-particle
Fitted PDBs: 7d6v
Q-score: 0.636
Zhou X, Gao Y, Wang W, Yang X, Yang X, Liu F, Tang Y, Lam SM, Shui G, Yu L, Tian C, Guddat LW, Wang Q, Rao Z, Gong H
PNAS (2021) 118 [ Pubmed: 33876763 DOI: doi:10.1073/pnas.2022308118 ]
- Iron/sulfur cluster (351 Da, Ligand)
- Fe3-s4 cluster (295 Da, Ligand)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Fumarate reductase iron-sulfur subunit (28 kDa, Protein from Mycolicibacterium smegmatis)
- (1s)-2-{[(2-aminoethoxy)(hydroxy)phosphoryl]oxy}-1-[(palmitoyloxy)methyl]ethyl stearate (720 Da, Ligand)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- Succinate dehydrogenase subunit a (70 kDa, Protein from Mycolicibacterium smegmatis)
- Mycobacterium smegmatis sdh1 in complex with uq1 (Complex from Mycolicibacterium smegmatis MC2 51)
- Ubiquinone-1 (250 Da, Ligand)
- Succinate dehydrogenase (membrane anchor subunit) (32 kDa, Protein from Mycolicibacterium smegmatis)