Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
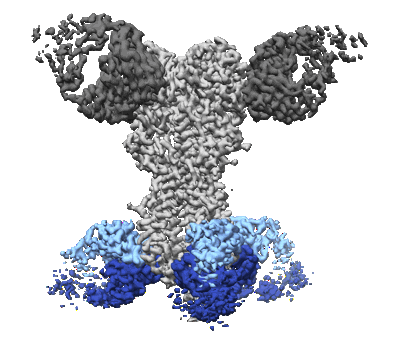
CryoEM map of anchor 222-1C06 Fab and lateral patch 2B05 Fab binding H1 HA
Resolution: 3.38 Å
EM Method: Single-particle
Fitted PDBs: 7t3d
Q-score: 0.464
Guthmiller JJ, Han J, Utset HA, Li L, Lan LY, Henry C, Stamper CT, McMahon M, O'Dell G, Fernandez-Quintero ML, Freyn AW, Amanat F, Stovicek O, Gentles L, Richey ST, de la Pena AT, Rosado V, Dugan HL, Zheng NY, Tepora ME, Bitar DJ, Changrob S, Strohmeier S, Huang M, Garcia-Sastre A, Liedl KR, Bloom JD, Nachbagauer R, Palese P, Krammer F, Coughlan L, Ward AB, Wilson PC
Nature (2022) 602 pp. 314-320 [ Pubmed: 34942633 DOI: doi:10.1038/s41586-021-04356-8 ]
- Cryoem map of anchor 222-1c06 fab and lateral patch 2b05 fab binding h1 ha (Complex from Homo sapiens)
- 2b05 mab light chain (11 kDa, Protein from Homo sapiens)
- 222-1c06 mab heavy chain (13 kDa, Protein from Homo sapiens)
- Hemagglutinin (Complex from Influenza A virus (A/California/04/2009(H1N1)))
- 2b05 mab heavy chain (12 kDa, Protein from Homo sapiens)
- 222-1c06 mab light chain (11 kDa, Protein from Homo sapiens)
- Hemagglutinin ha2 chain (19 kDa, Protein from Influenza A virus (A/California/04/2009(H1N1)))
- Hemagglutinin ha1 chain (36 kDa, Protein from Influenza A virus (A/California/04/2009(H1N1)))
- Monoclonal antibodies 222-1c06 and 2b05 (Complex from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
Proteinase K Determined by MicroED Phased by ARCIMBOLDO_SHREDDER
Richards LR, Millan C, Miao J, Martynowycz MW, Sawaya MR, Gonen T, Borges RJ, Uson I, Rodriguez JA
Acta Crystallogr D Struct Biol (2020) 76 pp. 703-712 [ DOI: doi:10.1107/S2059798320008049 ]
- Water (18 Da, Ligand)
- Proteinase k (Complex from Parengyodontium album)
- Proteinase k (28 kDa, Protein from Parengyodontium album)
- Calcium ion (40 Da, Ligand)
1.4A low-dose structure of GSNQNNF determined from initial phases generated using radiation damage
Resolution: 1.4 Å
EM Method: Electron Crystallography
Fitted PDBs: 6vhb
Q-score: 0.832
Martynowycz MW, Hattne J, Gonen T
Structure (2020) 28 pp. 458- [ DOI: doi:10.1016/j.str.2020.01.008 Pubmed: 32023481 ]
- Gsnqnnf (779 Da, Protein from synthetic construct)
- Zinc ion (65 Da, Ligand)
- Synthetic proto-filament (899 Da, Complex from synthetic construct)
- Water (18 Da, Ligand)
- Acetate ion (59 Da, Ligand)
1.4A damaged structure of GSNQNNF used to determine initial phases from radiation damage
Martynowycz MW, Hattne J, Gonen T
Structure (2020) 28 pp. 458- [ DOI: doi:10.1016/j.str.2020.01.008 Pubmed: 32023481 ]
- Gsnqnnf (779 Da, Protein from synthetic construct)
- Zinc ion (65 Da, Ligand)
- Synthetic proto-filament (899 Da, Complex from synthetic construct)
- Water (18 Da, Ligand)
- Acetate ion (59 Da, Ligand)
MicroED structure of a FIB-milled CypA Crystal
Resolution: 2.5 Å
EM Method: Electron Crystallography
Fitted PDBs: 6u5g
Q-score: 0.702
Wolff AM, Young ID, Sierra RG, Brewster AS, Martynowycz MW, Nango E, Sugahara M, Nakane T, Ito K, Aquila A, Bhowmick A, Biel JT, Carbajo S, Cohen AE, Cortez S, Gonzalez A, Hino T, Im D, Koralek JD, Kubo M, Lazarou TS, Nomura T, Owada S, Samelson A, Tanaka T, Tanaka R, Thompson EM, van den Bedem H, Woldeyes RA, Yumoto F, Zhao W, Tono K, Boutet S, Iwata S, Gonen T, Sauter NK, Fraser JS, Thompson MC
IUCrJ (2020) 7 pp. 306-323 [ DOI: doi:10.1107/S205225252000072X ]
- Peptidyl-prolyl cis-trans isomerase a (18 kDa, Protein from Homo sapiens)
- Water (18 Da, Ligand)
- Peptidyl-prolyl cis-trans isomerase a (Complex from Homo sapiens)
MicroED structure of NaK ion channel reveals a process of Na+ partition into the selectivity filter
Resolution: 2.5002 Å
EM Method: Electron Crystallography
Fitted PDBs: 6cpv
Q-score: 0.666
Commun Biol (2018) 1 pp. 38-38 [ DOI: doi:10.1038/s42003-018-0040-8 Pubmed: 30167468 ]
- Potassium channel protein (10 kDa, Protein from Bacillus cereus)
- (4s)-2-methyl-2,4-pentanediol (118 Da, Ligand)
- Water (18 Da, Ligand)
- Nak (Complex from Bacillus cereus)
- Sodium ion (22 Da, Ligand)
Guenther EL, Cao Q, Trinh H, Lu J, Sawaya MR, Cascio D, Boyer DR, Rodriguez JA, Hughes MP, Eisenberg DS
Nat Struct Mol Biol (2018) 25 pp. 463-471 [ DOI: doi:10.1038/s41594-018-0064-2 Pubmed: 29786080 ]
- Water (18 Da, Ligand)
- Crystal of nfgtfs phosphorylated on threonine. (Complex from Homo sapiens)
- Nfgtfs (751 Da, Protein from Homo sapiens)
Resolution: 1.0 Å
EM Method: Electron Crystallography
Fitted PDBs: 5wkb
Q-score: 0.383
Guenther EL, Cao Q, Trinh H, Lu J, Sawaya MR, Cascio D, Boyer DR, Rodriguez JA, Hughes MP, Eisenberg DS
Nat Struct Mol Biol (2018) 25 pp. 463-471 [ DOI: doi:10.1038/s41594-018-0064-2 Pubmed: 29786080 ]
- Water (18 Da, Ligand)
- Segment from tdp-43 (Cellular component from Homo sapiens)
- Tar dna-binding protein 43 (699 Da, Protein from Homo sapiens)
1.01 A MicroED structure of GSNQNNF at 0.27 e- / A^2
Resolution: 1.01 Å
EM Method: Electron Crystallography
Fitted PDBs: 6clc
Q-score: 0.958
Hattne J, Shi D, Glynn C, Zee CT, Gallagher-Jones M, Martynowycz MW, Rodriguez JA, Gonen T
Structure (2018) 26 pp. 759-766.e4 [ DOI: doi:10.1016/j.str.2018.03.021 Pubmed: 29706530 ]
- Gsnqnnf (779 Da, Protein from synthetic construct)
- Zinc ion (65 Da, Ligand)
- Water (18 Da, Ligand)
- Acetate ion (59 Da, Ligand)
- Synthetic proto-filament (899 Da, Complex)
1.01 A MicroED structure of GSNQNNF at 0.81 e- / A^2
Resolution: 1.01 Å
EM Method: Electron Crystallography
Fitted PDBs: 6cld
Q-score: 0.96
Hattne J, Shi D, Glynn C, Zee CT, Gallagher-Jones M, Martynowycz MW, Rodriguez JA, Gonen T
Structure (2018) 26 pp. 759-766.e4 [ DOI: doi:10.1016/j.str.2018.03.021 Pubmed: 29706530 ]
- Gsnqnnf (779 Da, Protein from synthetic construct)
- Zinc ion (65 Da, Ligand)
- Water (18 Da, Ligand)
- Acetate ion (59 Da, Ligand)
- Synthetic proto-filament (899 Da, Complex)