Resolution: 2.4 Å
EM Method: Single-particle
Fitted PDBs: 8ef9
Q-score: 0.605
Li C, Zhu H, Jin S, Maksoud LM, Jain N, Sun J, Gao Y
Nucleic Acids Res (2023) 51 pp. 463-474 [ Pubmed: 36583344 DOI: doi:10.1093/nar/gkac1201 ]
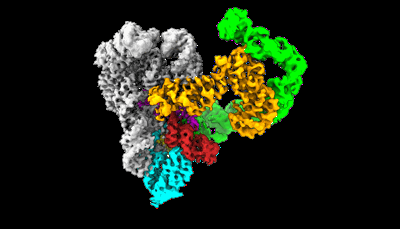
- Dna (5'-d(*ap*gp*cp*ap*tp*cp*cp*gp*tp*ap*gp*(2da))-3') (5 kDa, DNA from Lates calcarifer)
- Ternary complex of lates calcarifer dna polymerase theta with duplex dna and incoming nucleotide (Complex from Lates calcarifer)
- 2'-deoxyguanosine-5'-triphosphate (507 Da, Ligand)
- Dna polymerase theta (95 kDa, Protein from Lates calcarifer)
- Magnesium ion (24 Da, Ligand)
- Dna (5'-d(*ap*gp*cp*tp*cp*tp*ap*cp*gp*gp*ap*tp*gp*c)-3') (8 kDa, DNA from Lates calcarifer)
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 8efc
Q-score: 0.492
Li C, Zhu H, Jin S, Maksoud LM, Jain N, Sun J, Gao Y
Nucleic Acids Res (2023) 51 pp. 463-474 [ Pubmed: 36583344 DOI: doi:10.1093/nar/gkac1201 ]
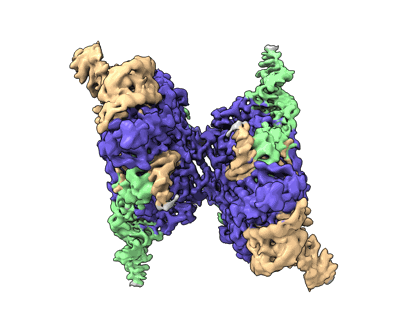
- Ternary complex of lates calcarifer dna polymerase theta with duplex dna and incoming nucleotide (Complex from Lates calcarifer)
- Dna (5'-d(*ap*cp*tp*gp*tp*gp*ap*gp*gp*cp*ap*tp*cp*cp*gp*tp*ap*gp*(2da))-3') (6 kDa, DNA from Lates calcarifer)
- 2'-deoxyguanosine-5'-triphosphate (507 Da, Ligand)
- Dna polymerase theta (95 kDa, Protein from Lates calcarifer)
- Magnesium ion (24 Da, Ligand)
- Dna (5'-d(*ap*gp*cp*tp*cp*tp*ap*cp*gp*gp*ap*tp*gp*cp*cp*tp*cp*ap*cp*ap*g)-3') (8 kDa, DNA from Lates calcarifer)
Structure of Lates calcarifer DNA polymerase theta polymerase domain with hairpin DNA
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 8efk
Q-score: 0.525
Li C, Zhu H, Jin S, Maksoud LM, Jain N, Sun J, Gao Y
Nucleic Acids Res (2023) 51 pp. 463-474 [ Pubmed: 36583344 DOI: doi:10.1093/nar/gkac1201 ]
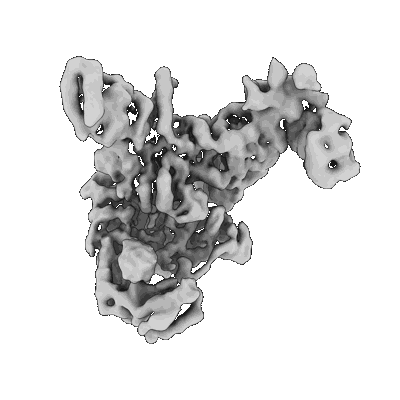
- Ternary complex of lates calcarifer dna polymerase theta with duplex dna and incoming nucleotide (Complex from Lates calcarifer)
- Dna (5'-d(p*tp*tp*tp*tp*gp*gp*cp*tp*tp*tp*tp*gp*cp*cp*(2da))-3') (7 kDa, DNA from Lates calcarifer)
- Magnesium ion (24 Da, Ligand)
- Lates calcarifer dna polymerase theta (95 kDa, Protein from Lates calcarifer)
- 2',3'-dideoxyadenosine triphosphate (475 Da, Ligand)
Resolution: 3.38 Å
EM Method: Single-particle
Fitted PDBs: 7uo4
Q-score: 0.457
Malone BF, Perry JK, Olinares PDB, Lee HW, Chen J, Appleby TC, Feng JY, Bilello JP, Ng H, Sotiris J, Ebrahim M, Chua EYD, Mendez JH, Eng ET, Landick R, Gotte M, Chait BT, Campbell EA, Darst SA
Nature (2023) 614 pp. 781-787 [ Pubmed: 36725929 DOI: doi:10.1038/s41586-022-05664-3 ]
- Sars-cov-2 replication-transcription complex (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Water (18 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Template rna (55-mer) (17 kDa, RNA from synthetic construct)
- [[(2~{r},3~{s},4~{r},5~{r})-5-(4-azanylpyrrolo[2,1-f][1,2,4]triazin-7-yl)-5-cyano-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl] phosphono hydrogen phosphate (531 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Product rna (35-mer) (11 kDa, RNA from synthetic construct)
- Non-structural protein 7 (10 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
CryoEM Structure of an Group II Intron Retroelement (apo-complex)
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7uim
Q-score: 0.262
Chung K, Xu L, Chai P, Peng J, Devarkar SC, Pyle AM
Science (2022) 378 pp. 627-634 [ Pubmed: 36356138 DOI: doi:10.1126/science.abq2844 ]
- Ammonium ion (18 Da, Ligand)
- E.r iic intron (206 kDa, RNA from [Eubacterium] rectale)
- Complex of rna and protein (250 kDa, Complex from [Eubacterium] rectale)
- Magnesium ion (24 Da, Ligand)
- Group ii intron reverse transcriptase/maturase (49 kDa, Protein from [Eubacterium] rectale)
CryoEM Structure of an Group II Intron Retroelement
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 7uin
Q-score: 0.467
Chung K, Xu L, Chai P, Peng J, Devarkar SC, Pyle AM
Science (2022) 378 pp. 627-634 [ Pubmed: 36356138 DOI: doi:10.1126/science.abq2844 ]
- Ammonium ion (18 Da, Ligand)
- E.r iic intron (206 kDa, RNA from [Eubacterium] rectale)
- Ternary complex of rna, protein and dna (260 kDa, Complex from [Eubacterium] rectale)
- Water (18 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Dna (37-mer) (11 kDa, DNA from [Eubacterium] rectale)
- Group ii intron reverse transcriptase/maturase (49 kDa, Protein from [Eubacterium] rectale)
Cryo-EM structure of Escherichia coli release complex of transcription termination (TTC-release)
Resolution: 3.48 Å
EM Method: Single-particle
Fitted PDBs: 7ypb
Q-score: 0.399
You L, Omollo EO, Yu C, Mooney RA, Shi J, Shen L, Wu X, Wen A, He D, Zeng Y, Feng Y, Landick R, Zhang Y
Nature (2023) 613 pp. 783-789 [ Pubmed: 36631609 DOI: doi:10.1038/s41586-022-05604-1 ]
- Rna (5'-r(*gp*cp*gp*up*cp*gp*cp*ap*gp*gp*cp*cp*up*up*up*up*up*ap*up*u)-3') (6 kDa, RNA from synthetic construct)
- Dna-directed rna polymerase (Complex)
- Dna-directed rna polymerase subunit alpha (36 kDa, Protein from Escherichia coli K-12)
- Dna-directed rna polymerase subunit beta (150 kDa, Protein from Escherichia coli K-12)
- Dna (31-mer) (9 kDa, DNA from synthetic construct)
- Dna (31-mer) (9 kDa, DNA from synthetic construct)
- Dna-directed rna polymerase subunit omega (10 kDa, Protein from Escherichia coli K-12)
- Dna/rna (Complex)
- E. coli release complex of transcription termination (400 kDa, Complex from Escherichia coli K-12)
- Dna-directed rna polymerase subunit beta' (156 kDa, Protein from Escherichia coli K-12)
- Magnesium ion (24 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Rna (5'-r(*gp*gp*cp*cp*up*gp*cp*gp*ap*cp*gp*a)-3') (3 kDa, RNA from synthetic construct)
Cryo-EM structure of E.coli retron-Ec86 (RT-msDNA-RNA) at 3.2 angstrom
Resolution: 3.12 Å
EM Method: Single-particle
Fitted PDBs: 7v9u
Q-score: 0.471
Wang Y, Guan Z, Wang C, Nie Y, Chen Y, Qian Z, Cui Y, Xu H, Wang Q, Zhao F, Zhang D, Tao P, Sun M, Yin P, Jin S, Wu S, Zou T
Nat Microbiol (2022) 7 pp. 1480-1489 [ Pubmed: 35982312 DOI: doi:10.1038/s41564-022-01197-7 ]
- Rna (81-mer) (25 kDa, RNA from Escherichia coli)
- Rna-directed dna polymerase from retron ec86 (36 kDa, Protein from Escherichia coli)
- Retron-ec86 (rt-msdna-rna) (Complex from Escherichia coli)
- Rna (5'-r(p*cp*gp*up*ap*ap*gp*gp*g)-3') (4 kDa, RNA from Escherichia coli)
- Dna (105-mer) (32 kDa, DNA from Escherichia coli)
Cryo-EM structure of E.coli retron-Ec86 in complex with its effector at 2.8 angstrom
Resolution: 2.82 Å
EM Method: Single-particle
Fitted PDBs: 7v9x
Q-score: 0.549
To Be Published
- Rna (81-mer) (25 kDa, RNA from Escherichia coli)
- Retron-ec86 (rt-msdna-rna) (Complex from Escherichia coli)
- Rna-directed dna polymerase from retron ec86 (37 kDa, Protein from Escherichia coli)
- Dna (105-mer) (32 kDa, DNA from Escherichia coli)
- Rna (14-mer) (4 kDa, RNA from Escherichia coli)
- Retron st85 family effector protein (35 kDa, Protein from Escherichia coli)
Tetrahymena CST with Polymerase alpha-Primase
Resolution: 4.2 Å
EM Method: Single-particle
Fitted PDBs: 7uy7
Q-score: 0.293
He Y, Song H, Chan H, Liu B, Wang Y, Susac L, Zhou ZH, Feigon J
Nature (2022) 608 pp. 813-818 [ Pubmed: 35831498 DOI: doi:10.1038/s41586-022-04931-7 ]
- Telomere dna (19 kDa, DNA from Tetrahymena thermophila)
- Telomerase-associated protein of 19 kda (19 kDa, Protein from Tetrahymena thermophila)
- Telomerase-associated protein of 75 kda (73 kDa, Protein from Tetrahymena thermophila)
- Dna polymerase (161 kDa, Protein from Tetrahymena thermophila)
- Telomerase associated protein p50 (50 kDa, Protein from Tetrahymena thermophila)
- Tetrahymena cst (Complex from Tetrahymena thermophila)
- Tetrahymena cst-polaprim (Complex)
- Zinc ion (65 Da, Ligand)
- Tetrahymena dna polymerase alpha-primase (Complex from Tetrahymena thermophila)
- Telomerase-associated protein of 45 kda (43 kDa, Protein from Tetrahymena thermophila)