Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
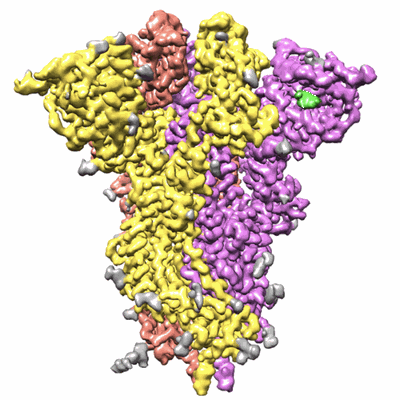
Trimeric SARS-CoV-2 spike ectodomain in complex with biliverdin (closed conformation)
Resolution: 3.36 Å
EM Method: Single-particle
Fitted PDBs: 7nt9
Q-score: 0.458
Rosa A, Pye VE, Graham C, Muir L, Seow J, Ng KW, Cook NJ, Rees-Spear C, Parker E, Dos Santos MS, Rosadas C, Susana A, Rhys H, Nans A, Masino L, Roustan C, Christodoulou E, Ulferts R, Wrobel AG, Short CE, Fertleman M, Sanders RW, Heaney J, Spyer M, Kjaer S, Riddell A, Malim MH, Beale R, MacRae JI, Taylor GP, Nastouli E, van Gils MJ, Rosenthal PB, Pizzato M, McClure MO, Tedder RS, Kassiotis G, McCoy LE, Doores KJ, Cherepanov P
Sci Adv (2021) 7 [ Pubmed: 33888467 DOI: doi:10.1126/sciadv.abg7607 ]
- Ectodomain of spike glycoprotein from sars-cov-2 expressed in hek cells and purified. data were collected in the presence of excess biliverdin. (426 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Biliverdine ix alpha (582 Da, Ligand)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
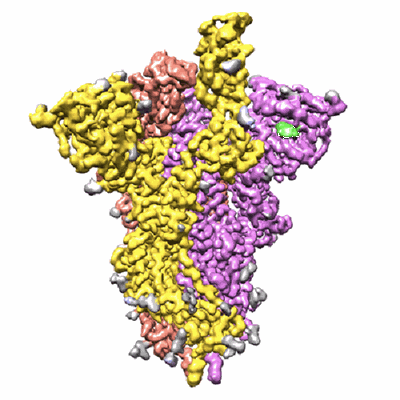
Trimeric SARS-CoV-2 spike ectodomain in complex with biliverdin (one RBD erect)
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 7nta
Q-score: 0.436
Rosa A, Pye VE, Graham C, Muir L, Seow J, Ng KW, Cook NJ, Rees-Spear C, Parker E, Dos Santos MS, Rosadas C, Susana A, Rhys H, Nans A, Masino L, Roustan C, Christodoulou E, Ulferts R, Wrobel AG, Short CE, Fertleman M, Sanders RW, Heaney J, Spyer M, Kjaer S, Riddell A, Malim MH, Beale R, MacRae JI, Taylor GP, Nastouli E, van Gils MJ, Rosenthal PB, Pizzato M, McClure MO, Tedder RS, Kassiotis G, McCoy LE, Doores KJ, Cherepanov P
Sci Adv (2021) 7 [ Pubmed: 33888467 DOI: doi:10.1126/sciadv.abg7607 ]
- Biliverdine ix alpha (582 Da, Ligand)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Ectodomain of s1 protein from sars-cov-2 expressed in hek cells and purified. sample in the presence of excess biliverdin. (426 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
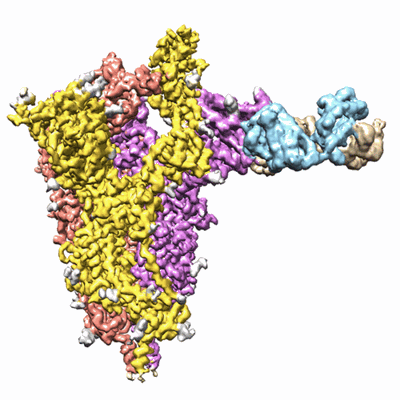
Trimeric SARS-CoV-2 spike ectodomain bound to P008_056 Fab
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 7ntc
Q-score: 0.391
Rosa A, Pye VE, Graham C, Muir L, Seow J, Ng KW, Cook NJ, Rees-Spear C, Parker E, Dos Santos MS, Rosadas C, Susana A, Rhys H, Nans A, Masino L, Roustan C, Christodoulou E, Ulferts R, Wrobel AG, Short CE, Fertleman M, Sanders RW, Heaney J, Spyer M, Kjaer S, Riddell A, Malim MH, Beale R, MacRae JI, Taylor GP, Nastouli E, van Gils MJ, Rosenthal PB, Pizzato M, McClure MO, Tedder RS, Kassiotis G, McCoy LE, Doores KJ, Cherepanov P
Sci Adv (2021) 7 [ Pubmed: 33888467 DOI: doi:10.1126/sciadv.abg7607 ]
- P008_056 fab heavy chain (27 kDa, Protein from Homo sapiens)
- P008_056 fab light chain (25 kDa, Protein from Homo sapiens)
- P008_056 fab (Complex from Homo sapiens)
- Biliverdine ix alpha (582 Da, Ligand)
- Spike glycoprotein (142 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Stabilised sars-cov-2 spike ectodomain trimer bound to one p008_056 fab (478 kDa, Complex)
- Stabilised sars-cov-2 spike ectodomain trimer (Complex from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)