Resolution: 3.97 Å
EM Method: Single-particle
Fitted PDBs: 7sgd
Q-score: 0.366
Brouwer PJM, Antanasijevic A, Ronk AJ, Muller-Krauter H, Watanabe Y, Claireaux M, Perrett HR, Bijl TPL, Grobben M, Umotoy JC, Schriek AI, Burger JA, Tejjani K, Lloyd NM, Steijaert TH, van Haaren MM, Sliepen K, de Taeye SW, van Gils MJ, Crispin M, Strecker T, Bukreyev A, Ward AB, Sanders RW
Cell Host Microbe (2022) 30 pp. 1759- [ DOI: doi:10.1016/j.chom.2022.10.018 Pubmed: 36400021 ]
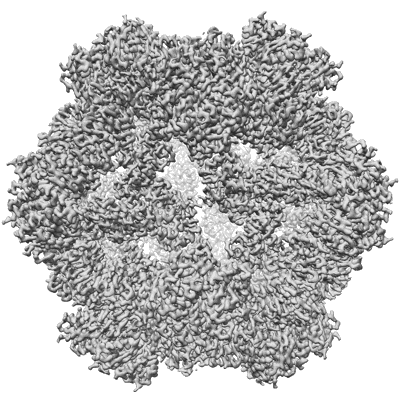
I53-50 nanoparticle core reconstructed from GPC-I53-50NP by focused refinement
Resolution: 3.67 Å
EM Method: Single-particle
Fitted PDBs: 7sge
Q-score: 0.479
Brouwer PJM, Antanasijevic A, Ronk AJ, Muller-Krauter H, Watanabe Y, Claireaux M, Perrett HR, Bijl TPL, Grobben M, Umotoy JC, Schriek AI, Burger JA, Tejjani K, Lloyd NM, Steijaert TH, van Haaren MM, Sliepen K, de Taeye SW, van Gils MJ, Crispin M, Strecker T, Bukreyev A, Ward AB, Sanders RW
Cell Host Microbe (2022) 30 pp. 1759- [ DOI: doi:10.1016/j.chom.2022.10.018 Pubmed: 36400021 ]
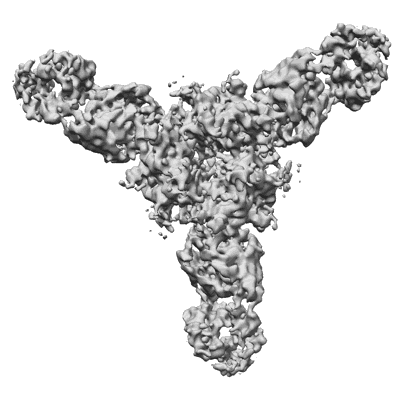
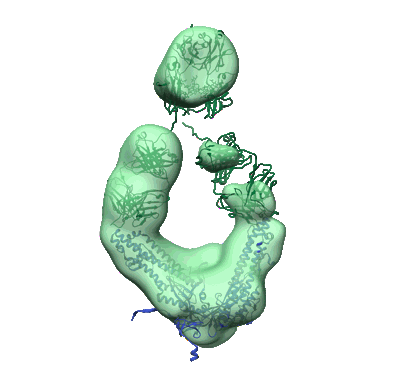
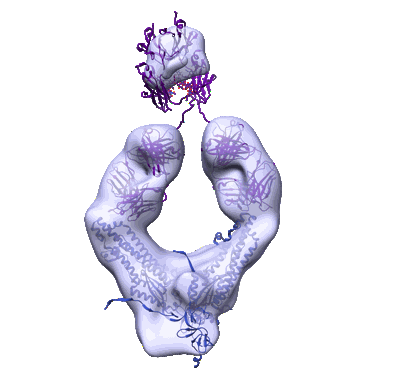
Lassa virus glycoprotein construct (Josiah GPC-I53-50A) in complex with LAVA01 antibody
Resolution: 4.41 Å
EM Method: Single-particle
Fitted PDBs: 7sgf
Q-score: 0.292
Brouwer PJM, Antanasijevic A, Ronk AJ, Muller-Krauter H, Watanabe Y, Claireaux M, Perrett HR, Bijl TPL, Grobben M, Umotoy JC, Schriek AI, Burger JA, Tejjani K, Lloyd NM, Steijaert TH, van Haaren MM, Sliepen K, de Taeye SW, van Gils MJ, Crispin M, Strecker T, Bukreyev A, Ward AB, Sanders RW
Cell Host Microbe (2022) 30 pp. 1759- [ DOI: doi:10.1016/j.chom.2022.10.018 Pubmed: 36400021 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Lava01 mab - heavy chain (15 kDa, Protein from Oryctolagus cuniculus)
- Gpc-i53-50a (73 kDa, Protein from Lassa mammarenavirus)
- Lava01 mab - light chain (12 kDa, Protein from Oryctolagus cuniculus)
- Lassa virus glycoprotein construct(josiah gpc-i53-50a) in complex with lava01 antibody (as fab fragment) (Complex from Lassa mammarenavirus)
Negative stain EM map of COVA1-07 mAb bound to the S2 domain of SARS-CoV-2 S
Claireaux M, Caniels TG, de Gast M, Han J, Guerra D, Kerster G, van Schaik BDC, Jongejan A, Schriek AI, Grobben M, Brouwer PJM, van der Straten K, Aldon Y, Capella-Pujol J, Snitselaar JL, Olijhoek W, Aartse A, Brinkkemper M, Bontjer I, Burger JA, Poniman M, Bijl TPL, Torres JL, Copps J, Martin IC, de Taeye SW, de Bree GJ, Ward AB, Sliepen K, van Kampen AHC, Moerland PD, Sanders RW, van Gils MJ
Nat Commun (2022) 13 pp. 4539-4539 [ Pubmed: 35927266 DOI: doi:10.1038/s41467-022-32232-0 ]
Negative stain EM map of COVA2-14 mAb bound to the S2 domain of SARS-CoV-2 S
Claireaux M, Caniels TG, de Gast M, Han J, Guerra D, Kerster G, van Schaik BDC, Jongejan A, Schriek AI, Grobben M, Brouwer PJM, van der Straten K, Aldon Y, Capella-Pujol J, Snitselaar JL, Olijhoek W, Aartse A, Brinkkemper M, Bontjer I, Burger JA, Poniman M, Bijl TPL, Torres JL, Copps J, Martin IC, de Taeye SW, de Bree GJ, Ward AB, Sliepen K, van Kampen AHC, Moerland PD, Sanders RW, van Gils MJ
Nat Commun (2022) 13 pp. 4539-4539 [ Pubmed: 35927266 DOI: doi:10.1038/s41467-022-32232-0 ]
Negative stain EM map of COVA2-18 mAb bound to the S2 domain of SARS-CoV-2 S
Claireaux M, Caniels TG, de Gast M, Han J, Guerra D, Kerster G, van Schaik BDC, Jongejan A, Schriek AI, Grobben M, Brouwer PJM, van der Straten K, Aldon Y, Capella-Pujol J, Snitselaar JL, Olijhoek W, Aartse A, Brinkkemper M, Bontjer I, Burger JA, Poniman M, Bijl TPL, Torres JL, Copps J, Martin IC, de Taeye SW, de Bree GJ, Ward AB, Sliepen K, van Kampen AHC, Moerland PD, Sanders RW, van Gils MJ
Nat Commun (2022) 13 pp. 4539-4539 [ Pubmed: 35927266 DOI: doi:10.1038/s41467-022-32232-0 ]
Negative stain EM map of the S2 domain of SARS-CoV-2 S
Claireaux M, Caniels TG, de Gast M, Han J, Guerra D, Kerster G, van Schaik BDC, Jongejan A, Schriek AI, Grobben M, Brouwer PJM, van der Straten K, Aldon Y, Capella-Pujol J, Snitselaar JL, Olijhoek W, Aartse A, Brinkkemper M, Bontjer I, Burger JA, Poniman M, Bijl TPL, Torres JL, Copps J, Martin IC, de Taeye SW, de Bree GJ, Ward AB, Sliepen K, van Kampen AHC, Moerland PD, Sanders RW, van Gils MJ
Nat Commun (2022) 13 pp. 4539-4539 [ Pubmed: 35927266 DOI: doi:10.1038/s41467-022-32232-0 ]
BG505 SOSIPv5.2 in complex with PGT122 and two RM20E1 Fabs
Derking R, Allen JD, Cottrell CA, Sliepen K, Seabright GE, Lee WH, Aldon Y, Rantalainen K, Antanasijevic A, Copps J, Yasmeen A, Cupo A, Cruz Portillo VM, Poniman M, Bol N, van der Woude P, de Taeye SW, van den Kerkhof TLGM, Klasse PJ, Ozorowski G, van Gils MJ, Moore JP, Ward AB, Crispin M, Sanders RW
Cell Rep (2021) 35 pp. 108933-108933 [ DOI: doi:10.1016/j.celrep.2021.108933 Pubmed: 33826885 ]
BG505 SOSIPv5.2 in complex with PGT122 and three RM20E1 Fabs
Derking R, Allen JD, Cottrell CA, Sliepen K, Seabright GE, Lee WH, Aldon Y, Rantalainen K, Antanasijevic A, Copps J, Yasmeen A, Cupo A, Cruz Portillo VM, Poniman M, Bol N, van der Woude P, de Taeye SW, van den Kerkhof TLGM, Klasse PJ, Ozorowski G, van Gils MJ, Moore JP, Ward AB, Crispin M, Sanders RW
Cell Rep (2021) 35 pp. 108933-108933 [ DOI: doi:10.1016/j.celrep.2021.108933 Pubmed: 33826885 ]
C3 symmetric BG505 SOSIPv5.2 in complex with PGT122 and three RM20E1 Fabs
Derking R, Allen JD, Cottrell CA, Sliepen K, Seabright GE, Lee WH, Aldon Y, Rantalainen K, Antanasijevic A, Copps J, Yasmeen A, Cupo A, Cruz Portillo VM, Poniman M, Bol N, van der Woude P, de Taeye SW, van den Kerkhof TLGM, Klasse PJ, Ozorowski G, van Gils MJ, Moore JP, Ward AB, Crispin M, Sanders RW
Cell Rep (2021) 35 pp. 108933-108933 [ DOI: doi:10.1016/j.celrep.2021.108933 Pubmed: 33826885 ]