Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
RNA-triggered protein cleavage and cell growth arrest by the type III-E CRISPR nuclease-protease
Kato K, Okazaki S, Schmitt-Ulms C, Jiang K, Zhou W, Ishikawa J, Isayama Y, Adachi S, Nishizawa T, Makarova KS, Koonin EV, Abudayyeh OO, Gootenberg JS, Nishimasu H
Science (2022) 378 pp. 882-889 [ Pubmed: 36423304 DOI: 10.1126/science.add7347 ]
Image sets:
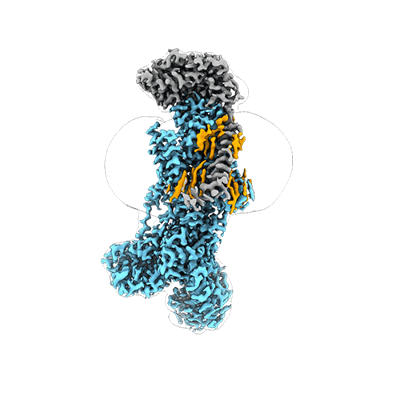
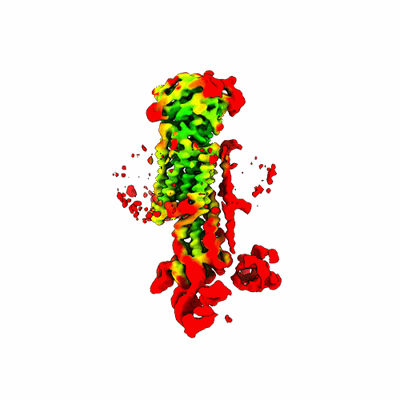
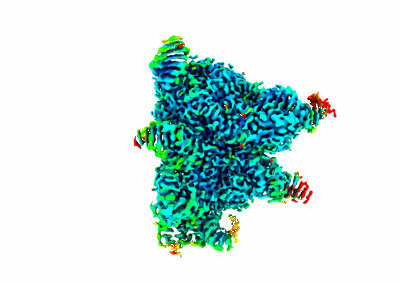
Structure and engineering of the type III-E CRISPR-Cas7-11 effector complex
Kato K, Zhou W, Okazaki S, Isayama Y, Nishizawa T, Gootenberg JS, Abudayyeh OO, Nishimasu H
Cell (2022) 185 pp. 2324-2337.e16 [ Pubmed: 35643083 DOI: 10.1016/j.cell.2022.05.003 ]
Image sets:
Cyo-EM structure of wildtype non-gastric proton pump in the presence of Na+, AlF and ADP
Resolution: 2.62 Å
EM Method: Single-particle
Fitted PDBs: 8ijl
Q-score: 0.612
Biochim Biophys Acta Mol Cell Res (2023) 1870 pp. 119543-119543 [ DOI: doi:10.1016/j.bbamcr.2023.119543 Pubmed: 37482134 ]
- Cholesterol (386 Da, Ligand)
- Tetrafluoroaluminate ion (102 Da, Ligand)
- Sodium/potassium-transporting atpase subunit beta-1 (37 kDa, Protein from Rattus)
- 1,2-dioleoyl-sn-glycero-3-phosphocholine (787 Da, Ligand)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Water (18 Da, Ligand)
- Hetero dimer of alpha and beta subunit (135 MDa, Complex from Rattus)
- Sodium ion (22 Da, Ligand)
- Sodium/potassium-transporting atpase subunit alpha (109 kDa, Protein from Rattus)
- Magnesium ion (24 Da, Ligand)
- 2-{[(4-o-alpha-d-glucopyranosyl-alpha-d-glucopyranosyl)oxy]methyl}-4-{[(3beta,9beta,14beta,17beta,25r)-spirost-5-en-3-yl]oxy}butyl 4-o-alpha-d-glucopyranosyl-alpha-d-glucopyranoside (1 kDa, Ligand)
Resolution: 2.81 Å
EM Method: Single-particle
Fitted PDBs: 8gui
Q-score: 0.532
Onishi S, Uchiyama K, Sato K, Okada C, Kobayashi S, Hamada K, Nishizawa T, Nureki O, Ogata K, Sengoku T
Nat Commun (2024) 15 pp. 2580-2580 [ Pubmed: 38519511 DOI: doi:10.1038/s41467-024-46910-8 ]
- Histone h4 (11 kDa, Protein from Homo sapiens)
- Dna (Complex)
- Zinc ion (65 Da, Ligand)
- Histone h2a type 1 (13 kDa, Protein from Homo sapiens)
- E3 ubiquitin-protein ligase bre1b (113 kDa, Protein from Homo sapiens)
- Histone (Complex from Homo sapiens)
- E3 ubiquitin-protein ligase bre1a (113 kDa, Protein from Homo sapiens)
- Bre1 (Complex from Homo sapiens)
- Histone h3.1 (15 kDa, Protein from Homo sapiens)
- Histone h2b type 1-j (14 kDa, Protein from Homo sapiens)
- Dna (147-mer) (45 kDa, DNA from synthetic construct)
- Dna (147-mer) (45 kDa, DNA from synthetic construct)
- Bre1-nucleosome complex (Complex)
Human nucleosome core particle (free form)
Resolution: 2.51 Å
EM Method: Single-particle
Fitted PDBs: 8guk
Q-score: 0.588
Onishi S, Uchiyama K, Sato K, Okada C, Kobayashi S, Hamada K, Nishizawa T, Nureki O, Ogata K, Sengoku T
Nat Commun (2024) 15 pp. 2580-2580 [ Pubmed: 38519511 DOI: doi:10.1038/s41467-024-46910-8 ]
- Histone h4 (11 kDa, Protein from Homo sapiens)
- Dna (Complex)
- Histone h2a type 1 (13 kDa, Protein from Homo sapiens)
- Bre1-nucleosome complex (Complex from Homo sapiens)
- Dna (147-mer) (45 kDa, DNA from synthetic construct)
- Histone h3.1 (15 kDa, Protein from Homo sapiens)
- Histone h2b type 1-j (14 kDa, Protein from Homo sapiens)
- Dna (147-mer) (45 kDa, DNA from synthetic construct)
- Hiistone (Complex)
Cyo-EM structure of K794A non-gastric proton pump in Na+ bound E1AMPPCP state
Resolution: 3.13 Å
EM Method: Single-particle
Fitted PDBs: 8ijm
Q-score: 0.412
Biochim Biophys Acta Mol Cell Res (2023) 1870 pp. 119543-119543 [ DOI: doi:10.1016/j.bbamcr.2023.119543 Pubmed: 37482134 ]
- Phosphomethylphosphonic acid adenylate ester (505 Da, Ligand)
- Cholesterol (386 Da, Ligand)
- Sodium/potassium-transporting atpase subunit beta-1 (37 kDa, Protein from Rattus)
- 1,2-dioleoyl-sn-glycero-3-phosphocholine (787 Da, Ligand)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Hetero dimer of alpha and beta subunit (135 MDa, Complex from Rattus)
- Sodium ion (22 Da, Ligand)
- Sodium/potassium-transporting atpase subunit alpha (109 kDa, Protein from Rattus)
Cryo-EM structure of non-gastric proton pump K794S mutant in Na+ bound E1AMPPCP state
Biochim Biophys Acta Mol Cell Res (2023) 1870 pp. 119543-119543 [ DOI: doi:10.1016/j.bbamcr.2023.119543 Pubmed: 37482134 ]
- Hetero dimer of alpha and beta subunit (135 MDa, Complex from Rattus)
Thermostabilized human prestin in complex with chloride, sulfate or salicylate
Futamata H, Fukuda M, Umeda R, Yamashita K, Tomita A, Takahashi S, Shikakura T, Hayashi S, Kusakizako T, Nishizawa T, Homma K, Nureki O
Nat Commun (2022) 13 [ DOI: 10.1038/s41467-022-34017-x Pubmed: 36266333 ]
Image sets:
Dog beta3 adrenergic receptor bound to mirabegron in complex with a miniGs heterotrimer
Nagiri C, Kobayashi K, Tomita A, Kato M, Kobayashi K, Yamashita K, Nishizawa T, Inoue A, Shihoya W, Nureki O
Mol Cell (2021) 81 pp. 3205-321500000.0 [ Pubmed: 34314699 DOI: 10.1016/j.molcel.2021.06.024 ]
Image sets:
Structure of the OgeuIscB-omega RNA-target DNA complex
Resolution: 2.55 Å
EM Method: Single-particle
Fitted PDBs: 7xht
Q-score: 0.619
Kato K, Okazaki S, Kannan S, Altae-Tran H, Esra Demircioglu F, Isayama Y, Ishikawa J, Fukuda M, Macrae RK, Nishizawa T, Makarova KS, Koonin EV, Zhang F, Nishimasu H
Nat Commun (2022) 13 pp. 6719-6719 [ Pubmed: 36344504 DOI: doi:10.1038/s41467-022-34378-3 ]
- Dna (Complex)
- Dna (5'-d(p*gp*ap*ap*gp*ap*ap*ap*ap*cp*cp*ap*t)-3') (4 kDa, DNA from synthetic construct)
- Rna (228-mer) (73 kDa, RNA from metagenome)
- Iscb-omegarna (Complex from metagenome)
- Dna (49-mer) (15 kDa, DNA from synthetic construct)
- Lauryl dimethylamine-n-oxide (229 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Structure of the iscb-omegarna ribonucleoprotein complex (Complex)
- Ogeuiscb (56 kDa, Protein from metagenome)