Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
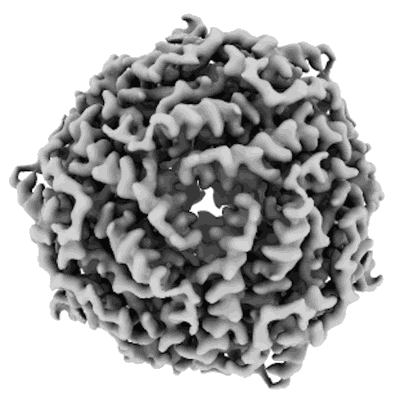
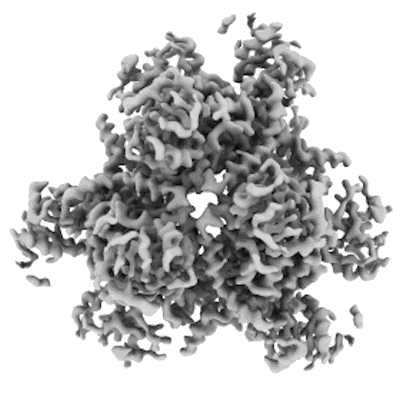
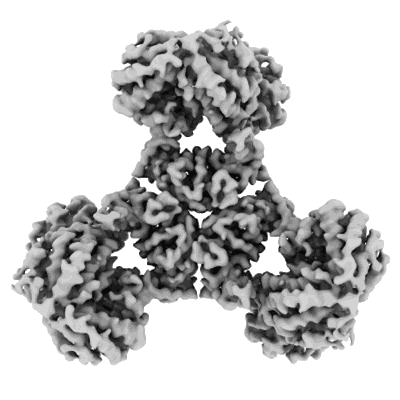
Structure of DPS determined by cryoEM at 100 keV
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 8pv9
Q-score: 0.615
McMullan G, Naydenova K, Mihaylov D, Yamashita K, Peet MJ, Wilson H, Dickerson JL, Chen S, Cannone G, Lee Y, Hutchings KA, Gittins O, Sobhy MA, Wells T, El-Gomati MM, Dalby J, Meffert M, Schulze-Briese C, Henderson R, Russo CJ
PNAS (2023) 120 pp. e2312905120-e2312905120 [ DOI: doi:10.1073/pnas.2312905120 Pubmed: 38011573 ]
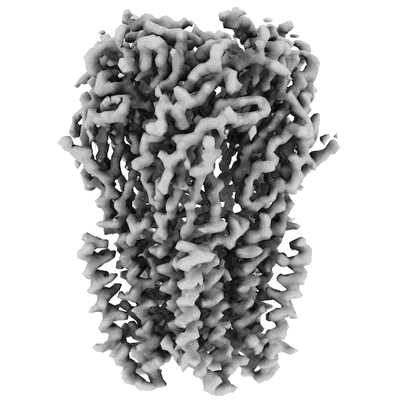
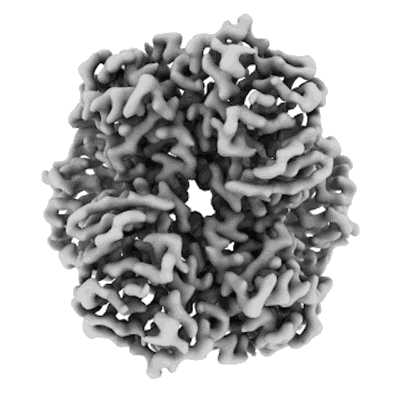
Structure of bacterial ribosome determined by cryoEM at 100 keV
Resolution: 4.5 Å
EM Method: Single-particle
Fitted PDBs: 8pva
Q-score: 0.287
McMullan G, Naydenova K, Mihaylov D, Yamashita K, Peet MJ, Wilson H, Dickerson JL, Chen S, Cannone G, Lee Y, Hutchings KA, Gittins O, Sobhy MA, Wells T, El-Gomati MM, Dalby J, Meffert M, Schulze-Briese C, Henderson R, Russo CJ
PNAS (2023) 120 pp. e2312905120-e2312905120 [ DOI: doi:10.1073/pnas.2312905120 Pubmed: 38011573 ]
- Small ribosomal subunit protein us3 (26 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l25 (10 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l33 (6 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l35 (7 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein bs16 (9 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l19 (13 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l28 (9 kDa, Protein from Escherichia coli)
- Zinc ion (65 Da, Ligand)
- Small ribosomal subunit protein bs6, fully modified isoform (15 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us5 (17 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein bs21 (8 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us4 (23 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us19 (10 kDa, Protein from Escherichia coli)
- Large ribosomal subunit protein ul13 (16 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l29 (7 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein bs18 (9 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l20 (13 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us7 (20 kDa, Protein from Escherichia coli)
- Large ribosomal subunit protein ul3 (22 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us8 (14 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein bs20 (9 kDa, Protein from Escherichia coli)
- Large ribosomal subunit protein bl31a (7 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us17 (9 kDa, Protein from Escherichia coli)
- E. coli 70s ribosome (Complex from Escherichia coli)
- P-site trna-fmet (24 kDa, RNA from Escherichia coli)
- 50s ribosomal protein l15 (15 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l34 (5 kDa, Protein from Escherichia coli)
- E-site trna (589 Da, RNA from Escherichia coli)
- 50s ribosomal protein l36 (4 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l18 (12 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l27 (9 kDa, Protein from Escherichia coli)
- 23s rrna (941 kDa, RNA from Escherichia coli)
- Large ribosomal subunit protein ul6 (18 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l30 (6 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us15 (10 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l23 (11 kDa, Protein from Escherichia coli)
- Large ribosomal subunit protein bl9 (15 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us13 (13 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us2 (26 kDa, Protein from Escherichia coli)
- Large ribosomal subunit protein ul2 (29 kDa, Protein from Escherichia coli)
- 5s rrna (38 kDa, RNA from Escherichia coli)
- 50s ribosomal protein l14 (13 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l32 (6 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l24 (11 kDa, Protein from Escherichia coli)
- Large ribosomal subunit protein ul16 (15 kDa, Protein from Escherichia coli)
- Large ribosomal subunit protein ul5 (20 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us12 (13 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us9 (14 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us10 (11 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us11 (13 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l22 (12 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l17 (14 kDa, Protein from Escherichia coli)
- Mrna (9 kDa, RNA from Escherichia coli)
- A-site trna-val (24 kDa, RNA from Escherichia coli)
- 50s ribosomal protein l21 (11 kDa, Protein from Escherichia coli)
- 16s rrna (499 kDa, RNA from Escherichia coli)
- Large ribosomal subunit protein ul4 (22 kDa, Protein from Escherichia coli)
- Small ribosomal subunit protein us14 (11 kDa, Protein from Escherichia coli)
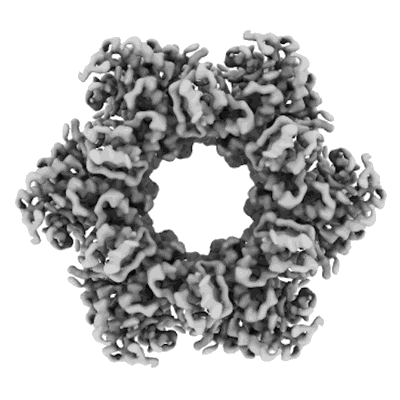
Structure of GABAAR determined by cryoEM at 100 keV
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 8pvb
Q-score: 0.486
McMullan G, Naydenova K, Mihaylov D, Yamashita K, Peet MJ, Wilson H, Dickerson JL, Chen S, Cannone G, Lee Y, Hutchings KA, Gittins O, Sobhy MA, Wells T, El-Gomati MM, Dalby J, Meffert M, Schulze-Briese C, Henderson R, Russo CJ
PNAS (2023) 120 pp. e2312905120-e2312905120 [ DOI: doi:10.1073/pnas.2312905120 Pubmed: 38011573 ]
- Hexadecane (226 Da, Ligand)
- Chloride ion (35 Da, Ligand)
- Decane (142 Da, Ligand)
- Human gaba(a)r-beta3 homopentamer in complex with megabody-25 in lipid nanodisc (Complex from Homo sapiens)
- Megabody mb25 (56 kDa, Protein from Lama glama)
- Etomidate (244 Da, Ligand)
- Gamma-aminobutyric acid receptor subunit beta-3 (45 kDa, Protein from Homo sapiens)
- Histamine (111 Da, Ligand)
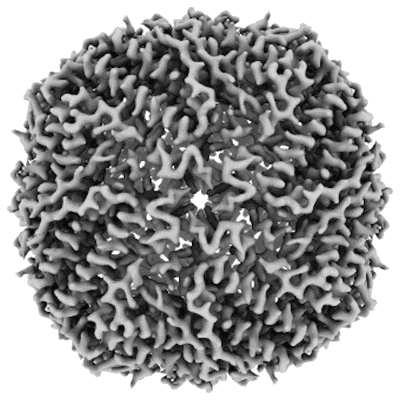
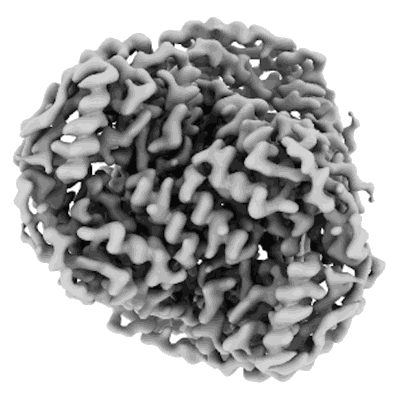
Structure of mouse heavy-chain apoferritin determined by cryoEM at 100 keV
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8pvc
Q-score: 0.675
McMullan G, Naydenova K, Mihaylov D, Yamashita K, Peet MJ, Wilson H, Dickerson JL, Chen S, Cannone G, Lee Y, Hutchings KA, Gittins O, Sobhy MA, Wells T, El-Gomati MM, Dalby J, Meffert M, Schulze-Briese C, Henderson R, Russo CJ
PNAS (2023) 120 pp. e2312905120-e2312905120 [ DOI: doi:10.1073/pnas.2312905120 Pubmed: 38011573 ]
- Zinc ion (65 Da, Ligand)
- Fe (iii) ion (55 Da, Ligand)
- Ferritin heavy chain (21 kDa, Protein from Mus musculus)
- Mouse heavy-chain apoferritin (Complex from Mus musculus)
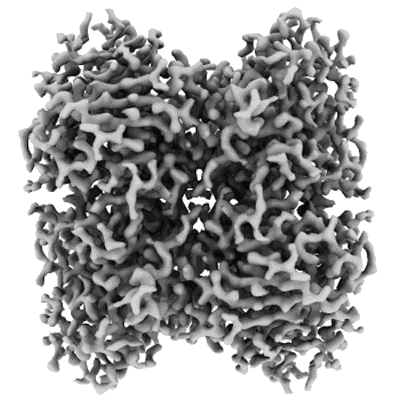
Structure of catalase determined by cryoEM at 100 keV
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8pvd
Q-score: 0.523
McMullan G, Naydenova K, Mihaylov D, Yamashita K, Peet MJ, Wilson H, Dickerson JL, Chen S, Cannone G, Lee Y, Hutchings KA, Gittins O, Sobhy MA, Wells T, El-Gomati MM, Dalby J, Meffert M, Schulze-Briese C, Henderson R, Russo CJ
PNAS (2023) 120 pp. e2312905120-e2312905120 [ DOI: doi:10.1073/pnas.2312905120 Pubmed: 38011573 ]
- Catalase from human erythrocytes (Complex from Homo sapiens)
- Nadph dihydro-nicotinamide-adenine-dinucleotide phosphate (745 Da, Ligand)
- Protoporphyrin ix containing fe (616 Da, Ligand)
- Catalase (59 kDa, Protein from Homo sapiens)
Structure of AHIR determined by cryoEM at 100 keV
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8pve
Q-score: 0.526
McMullan G, Naydenova K, Mihaylov D, Yamashita K, Peet MJ, Wilson H, Dickerson JL, Chen S, Cannone G, Lee Y, Hutchings KA, Gittins O, Sobhy MA, Wells T, El-Gomati MM, Dalby J, Meffert M, Schulze-Briese C, Henderson R, Russo CJ
PNAS (2023) 120 pp. e2312905120-e2312905120 [ DOI: doi:10.1073/pnas.2312905120 Pubmed: 38011573 ]
Structure of GAPDH determined by cryoEM at 100 keV
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 8pvf
Q-score: 0.572
McMullan G, Naydenova K, Mihaylov D, Yamashita K, Peet MJ, Wilson H, Dickerson JL, Chen S, Cannone G, Lee Y, Hutchings KA, Gittins O, Sobhy MA, Wells T, El-Gomati MM, Dalby J, Meffert M, Schulze-Briese C, Henderson R, Russo CJ
PNAS (2023) 120 pp. e2312905120-e2312905120 [ DOI: doi:10.1073/pnas.2312905120 Pubmed: 38011573 ]
Structure of E. coli glutamine synthetase determined by cryoEM at 100 keV
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8pvg
Q-score: 0.463
McMullan G, Naydenova K, Mihaylov D, Yamashita K, Peet MJ, Wilson H, Dickerson JL, Chen S, Cannone G, Lee Y, Hutchings KA, Gittins O, Sobhy MA, Wells T, El-Gomati MM, Dalby J, Meffert M, Schulze-Briese C, Henderson R, Russo CJ
PNAS (2023) 120 pp. e2312905120-e2312905120 [ DOI: doi:10.1073/pnas.2312905120 Pubmed: 38011573 ]
Structure of human apo ALDH1A1 determined by cryoEM at 100 keV
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 8pvh
Q-score: 0.55
McMullan G, Naydenova K, Mihaylov D, Yamashita K, Peet MJ, Wilson H, Dickerson JL, Chen S, Cannone G, Lee Y, Hutchings KA, Gittins O, Sobhy MA, Wells T, El-Gomati MM, Dalby J, Meffert M, Schulze-Briese C, Henderson R, Russo CJ
PNAS (2023) 120 pp. e2312905120-e2312905120 [ DOI: doi:10.1073/pnas.2312905120 Pubmed: 38011573 ]
- Aldehyde dehydrogenase 1a1 (57 kDa, Protein from Homo sapiens)
- Chloride ion (35 Da, Ligand)
- Human apo aldh1a1 (Complex from Homo sapiens)
Structure of PaaZ determined by cryoEM at 100 keV
Resolution: 3.7 Å
EM Method: Single-particle
Fitted PDBs: 8pvi
Q-score: 0.432
McMullan G, Naydenova K, Mihaylov D, Yamashita K, Peet MJ, Wilson H, Dickerson JL, Chen S, Cannone G, Lee Y, Hutchings KA, Gittins O, Sobhy MA, Wells T, El-Gomati MM, Dalby J, Meffert M, Schulze-Briese C, Henderson R, Russo CJ
PNAS (2023) 120 pp. e2312905120-e2312905120 [ DOI: doi:10.1073/pnas.2312905120 Pubmed: 38011573 ]