Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
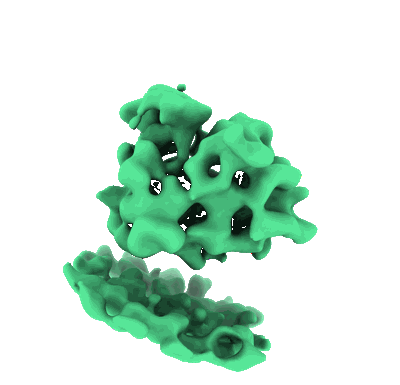
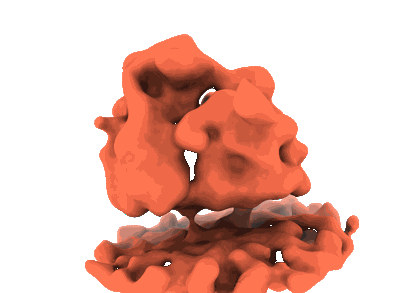
Cryo-electron tomogram acquired on a cryo-FIB lamella of a retinal pigment epithelial (RPE1) cell
de Teresa-Trueba I, Goetz SK, Mattausch A, Stojanovska F, Zimmerli CE, Toro-Nahuelpan M, Cheng DWC, Tollervey F, Pape C, Beck M, Diz-Munoz A, Kreshuk A, Mahamid J, Zaugg JB
Nat Methods (2023) 20 pp. 284-294 [ DOI: doi:10.1038/s41592-022-01746-2 Pubmed: 36690741 ]
Teresa I, Goetz SK, Mattausch A, Stojanovska F, Zimmerli CE, Toro-Nahuelpan M, Cheng DW, Tollervey F, Pape C, Beck M, Kreshuk A, Mahamid J, Zaugg J
bioRxiv (2022) [ DOI: 10.1101/2022.04.12.488077 ]
Image sets:
- Reconstructed cryo-electron tomograms acquired with a Volta potential phase plate (VPP) and defocus on S. pombe cryo-FIB lamellae
- Tilt series stacks and metadata of cryo-electron tomograms acquired with a Volta potential phase plate (VPP) and defocus on S. pombe cryo-FIB lamellae
- Tilt series stacks and metadata of cryo-electron tomograms acquired with defocus-only (DEF) on S. pombe cryo-FIB lamellae
- Raw unaligned multi-frame micrographs for each tilt in10 tilt-series acquired with a Volta potential phase plate (VPP) and defocus on S. pombe cryo-FIB lamellae
- Reconstructed cryo-electron tomograms acquired with defocus-only (DEF) on S. pombe cryo-FIB lamellae
- Raw unaligned multi-frame micrographs for each tilt in 10 tilt-series acquired with defocus-only (DEF) on S. pombe cryo-FIB lamellae
de Teresa-Trueba I, Goetz SK, Mattausch A, Stojanovska F, Zimmerli CE, Toro-Nahuelpan M, Cheng DWC, Tollervey F, Pape C, Beck M, Diz-Munoz A, Kreshuk A, Mahamid J, Zaugg JB
Nat Methods (2023) 20 pp. 284-294 [ DOI: doi:10.1038/s41592-022-01746-2 Pubmed: 36690741 ]
de Teresa-Trueba I, Goetz SK, Mattausch A, Stojanovska F, Zimmerli CE, Toro-Nahuelpan M, Cheng DWC, Tollervey F, Pape C, Beck M, Diz-Munoz A, Kreshuk A, Mahamid J, Zaugg JB
Nat Methods (2023) 20 pp. 284-294 [ DOI: doi:10.1038/s41592-022-01746-2 Pubmed: 36690741 ]
de Teresa-Trueba I, Goetz SK, Mattausch A, Stojanovska F, Zimmerli CE, Toro-Nahuelpan M, Cheng DWC, Tollervey F, Pape C, Beck M, Diz-Munoz A, Kreshuk A, Mahamid J, Zaugg JB
Nat Methods (2023) 20 pp. 284-294 [ DOI: doi:10.1038/s41592-022-01746-2 Pubmed: 36690741 ]
de Teresa-Trueba I, Goetz SK, Mattausch A, Stojanovska F, Zimmerli CE, Toro-Nahuelpan M, Cheng DWC, Tollervey F, Pape C, Beck M, Diz-Munoz A, Kreshuk A, Mahamid J, Zaugg JB
Nat Methods (2023) 20 pp. 284-294 [ DOI: doi:10.1038/s41592-022-01746-2 Pubmed: 36690741 ]
de Teresa-Trueba I, Goetz SK, Mattausch A, Stojanovska F, Zimmerli CE, Toro-Nahuelpan M, Cheng DWC, Tollervey F, Pape C, Beck M, Diz-Munoz A, Kreshuk A, Mahamid J, Zaugg JB
Nat Methods (2023) 20 pp. 284-294 [ DOI: doi:10.1038/s41592-022-01746-2 Pubmed: 36690741 ]
de Teresa-Trueba I, Goetz SK, Mattausch A, Stojanovska F, Zimmerli CE, Toro-Nahuelpan M, Cheng DWC, Tollervey F, Pape C, Beck M, Diz-Munoz A, Kreshuk A, Mahamid J, Zaugg JB
Nat Methods (2023) 20 pp. 284-294 [ DOI: doi:10.1038/s41592-022-01746-2 Pubmed: 36690741 ]
de Teresa-Trueba I, Goetz SK, Mattausch A, Stojanovska F, Zimmerli CE, Toro-Nahuelpan M, Cheng DWC, Tollervey F, Pape C, Beck M, Diz-Munoz A, Kreshuk A, Mahamid J, Zaugg JB
Nat Methods (2023) 20 pp. 284-294 [ DOI: doi:10.1038/s41592-022-01746-2 Pubmed: 36690741 ]