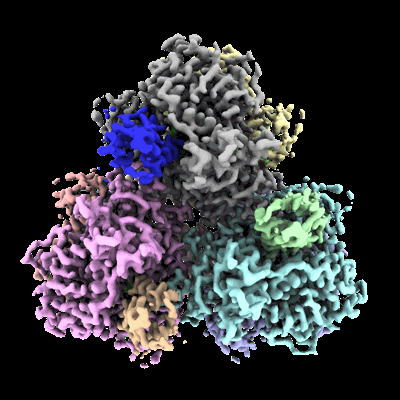
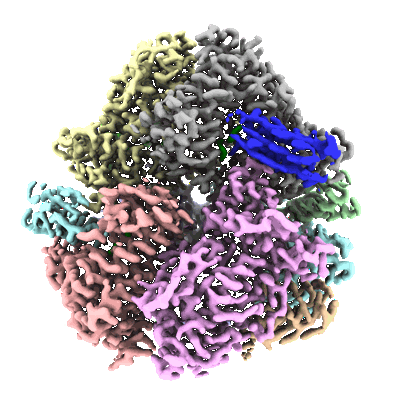
Structure of E. coli dGTPase bound to T7 bacteriophage protein Gp1.2
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 7u65
Q-score: 0.551
Klemm BP, Singh D, Smith CE, Hsu AL, Dillard LB, Krahn JM, London RE, Mueller GA, Borgnia MJ, Schaaper RM
PNAS (2022) 119 pp. e2123092119-e2123092119 [ DOI: doi:10.1073/pnas.2123092119 Pubmed: 36067314 ]
- Dgtpase hexamer bound to six copies of gp1.2 (Complex from Escherichia coli str. K-12 substr. MG1655)
- Inhibitor of dgtpase (10 kDa, Protein from Escherichia phage T7)
- Deoxyguanosinetriphosphate triphosphohydrolase (59 kDa, Protein from Escherichia coli str. K-12 substr. MG1655)
- Gene 1.2 protein (Complex from Escherichia phage T7)
- Dgtp triphosphohydrolase (Complex from Escherichia coli str. K-12 substr. MG1655)
Structure of E. coli dGTPase bound to T7 bacteriophage protein Gp1.2 and dGTP
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7u66
Q-score: 0.539
Klemm BP, Singh D, Smith CE, Hsu AL, Dillard LB, Krahn JM, London RE, Mueller GA, Borgnia MJ, Schaaper RM
PNAS (2022) 119 pp. e2123092119-e2123092119 [ DOI: doi:10.1073/pnas.2123092119 Pubmed: 36067314 ]
- Dgtpase hexamer bound to six copies of gp1.2 (Complex from Escherichia coli str. K-12 substr. MG1655)
- Inhibitor of dgtpase (10 kDa, Protein from Escherichia phage T7)
- Deoxyguanosinetriphosphate triphosphohydrolase (59 kDa, Protein from Escherichia coli str. K-12 substr. MG1655)
- Gene 1.2 protein (Complex from Escherichia phage T7)
- 2'-deoxyguanosine-5'-triphosphate (507 Da, Ligand)
- Dgtp triphosphohydrolase (Complex from Escherichia coli str. K-12 substr. MG1655)
- Magnesium ion (24 Da, Ligand)
Structure of E. coli dGTPase bound to T7 bacteriophage protein Gp1.2 and GTP
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 7u67
Q-score: 0.588
Klemm BP, Singh D, Smith CE, Hsu AL, Dillard LB, Krahn JM, London RE, Mueller GA, Borgnia MJ, Schaaper RM
PNAS (2022) 119 pp. e2123092119-e2123092119 [ DOI: doi:10.1073/pnas.2123092119 Pubmed: 36067314 ]
- Dgtpase hexamer bound to six copies of gp1.2 (Complex from Escherichia coli str. K-12 substr. MG1655)
- Inhibitor of dgtpase (10 kDa, Protein from Escherichia phage T7)
- Deoxyguanosinetriphosphate triphosphohydrolase (59 kDa, Protein from Escherichia coli str. K-12 substr. MG1655)
- Gene 1.2 protein (Complex from Escherichia phage T7)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Dgtp triphosphohydrolase (Complex from Escherichia coli str. K-12 substr. MG1655)
- Magnesium ion (24 Da, Ligand)
Structure of the L. blandensis dGTPase in the apo form
Resolution: 2.1 Å
EM Method: Single-particle
Fitted PDBs: 7tu5
Q-score: 0.651
Klemm BP, Sikkema AP, Hsu AL, Horng JC, Hall TMT, Borgnia MJ, Schaaper RM
J Biol Chem (2022) 298 pp. 102073-102073 [ DOI: doi:10.1016/j.jbc.2022.102073 Pubmed: 35643313 ]
- Magnesium ion (24 Da, Ligand)
- Dgtp triphosphohydrolase (52 kDa, Protein from Leeuwenhoekiella blandensis MED217)
- Dgtp triphosphohydrolase (Complex from Leeuwenhoekiella blandensis MED217)
- Water (18 Da, Ligand)
Structure of the L. blandensis dGTPase bound to dATP
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 7tu6
Q-score: 0.556
Klemm BP, Sikkema AP, Hsu AL, Horng JC, Hall TMT, Borgnia MJ, Schaaper RM
J Biol Chem (2022) 298 pp. 102073-102073 [ DOI: doi:10.1016/j.jbc.2022.102073 Pubmed: 35643313 ]
- Magnesium ion (24 Da, Ligand)
- 2'-deoxyadenosine 5'-triphosphate (491 Da, Ligand)
- Dgtp triphosphohydrolase (52 kDa, Protein from Leeuwenhoekiella blandensis MED217)
- Dgtp triphosphohydrolase (Complex from Leeuwenhoekiella blandensis MED217)
Structure of the L. blandensis dGTPase H125A mutant bound to dGTP
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 7tu7
Q-score: 0.637
Klemm BP, Sikkema AP, Hsu AL, Horng JC, Hall TMT, Borgnia MJ, Schaaper RM
J Biol Chem (2022) 298 pp. 102073-102073 [ DOI: doi:10.1016/j.jbc.2022.102073 Pubmed: 35643313 ]
- Magnesium ion (24 Da, Ligand)
- Dgtp triphosphohydrolase (52 kDa, Protein from Leeuwenhoekiella blandensis MED217)
- 2'-deoxyguanosine-5'-triphosphate (507 Da, Ligand)
- Dgtp triphosphohydrolase (Complex from Leeuwenhoekiella blandensis MED217)
Structure of the L. blandensis dGTPase H125A mutant bound to dGTP and dATP
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 7tu8
Q-score: 0.623
Klemm BP, Sikkema AP, Hsu AL, Horng JC, Hall TMT, Borgnia MJ, Schaaper RM
J Biol Chem (2022) 298 pp. 102073-102073 [ DOI: doi:10.1016/j.jbc.2022.102073 Pubmed: 35643313 ]
- 2'-deoxyadenosine 5'-triphosphate (491 Da, Ligand)
- Dgtp triphosphohydrolase (52 kDa, Protein from Leeuwenhoekiella blandensis MED217)
- 2'-deoxyguanosine-5'-triphosphate (507 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Dgtp triphosphohydrolase (Complex from Leeuwenhoekiella blandensis MED217)
Structure of dNTPase at 3.3 Angstrom Resolution
Bouvette J, Liu HF, Du X, Zhou Y, Sikkema AP, da Fonseca Rezende E Mello J, Klemm BP, Huang R, Schaaper RM, Borgnia MJ, Bartesaghi A
Nat Commun (2021) 12 pp. 1957-1957 [ DOI: doi:10.1038/s41467-021-22251-8 Pubmed: 33785757 ]
- Dntpase (Complex from Vibrio cholerae)
Structure of dNTPase at 3.6 Angstrom Resolution
Bouvette J, Liu HF, Du X, Zhou Y, Sikkema AP, da Fonseca Rezende E Mello J, Klemm BP, Huang R, Schaaper RM, Borgnia MJ, Bartesaghi A
Nat Commun (2021) 12 pp. 1957-1957 [ DOI: doi:10.1038/s41467-021-22251-8 Pubmed: 33785757 ]
- Dntpase (Complex from Vibrio cholerae)
Structure of 80S ribosome from EMPIAR-10064 at 5.6 Angstrom Resolution
Bouvette J, Liu HF, Du X, Zhou Y, Sikkema AP, da Fonseca Rezende E Mello J, Klemm BP, Huang R, Schaaper RM, Borgnia MJ, Bartesaghi A
Nat Commun (2021) 12 pp. 1957-1957 [ DOI: doi:10.1038/s41467-021-22251-8 Pubmed: 33785757 ]