Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 8fhw
Q-score: 0.579
Washington EJ, Zhou Y, Hsu AL, Petrovich M, Borgnia MJ, Bartesaghi A, Brennan RG
bioRxiv (2023) [ DOI: doi:10.1101/2023.03.14.530545 Pubmed: 36993618 ]
- Uridine-5'-diphosphate (404 Da, Ligand)
- 6-o-phosphono-alpha-d-glucopyranose (260 Da, Ligand)
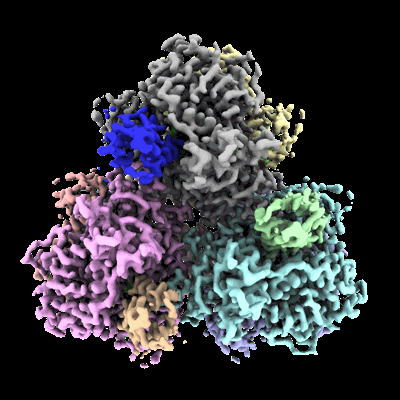
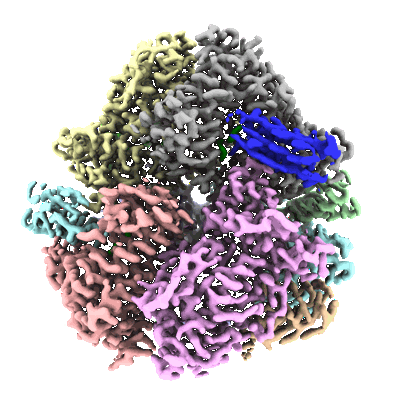
- Alpha,alpha-trehalose-phosphate synthase (udp-forming) (76 kDa, Protein from Cryptococcus neoformans var. grubii H99)
- Cryo-em structure of cryptococcus neoformans trehalose-6-phosphate synthase homotetramer in complex with uridine diphosphate and glucose-6-phosphate (307 kDa, Complex from Cryptococcus neoformans var. grubii H99)
Structure of E. coli dGTPase bound to T7 bacteriophage protein Gp1.2
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 7u65
Q-score: 0.551
Klemm BP, Singh D, Smith CE, Hsu AL, Dillard LB, Krahn JM, London RE, Mueller GA, Borgnia MJ, Schaaper RM
PNAS (2022) 119 pp. e2123092119-e2123092119 [ DOI: doi:10.1073/pnas.2123092119 Pubmed: 36067314 ]
- Dgtpase hexamer bound to six copies of gp1.2 (Complex from Escherichia coli str. K-12 substr. MG1655)
- Deoxyguanosinetriphosphate triphosphohydrolase (59 kDa, Protein from Escherichia coli str. K-12 substr. MG1655)
- Inhibitor of dgtpase (10 kDa, Protein from Escherichia phage T7)
- Dgtp triphosphohydrolase (Complex from Escherichia coli str. K-12 substr. MG1655)
- Gene 1.2 protein (Complex from Escherichia phage T7)
Structure of E. coli dGTPase bound to T7 bacteriophage protein Gp1.2 and dGTP
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7u66
Q-score: 0.539
Klemm BP, Singh D, Smith CE, Hsu AL, Dillard LB, Krahn JM, London RE, Mueller GA, Borgnia MJ, Schaaper RM
PNAS (2022) 119 pp. e2123092119-e2123092119 [ DOI: doi:10.1073/pnas.2123092119 Pubmed: 36067314 ]
- Dgtpase hexamer bound to six copies of gp1.2 (Complex from Escherichia coli str. K-12 substr. MG1655)
- Magnesium ion (24 Da, Ligand)
- 2'-deoxyguanosine-5'-triphosphate (507 Da, Ligand)
- Deoxyguanosinetriphosphate triphosphohydrolase (59 kDa, Protein from Escherichia coli str. K-12 substr. MG1655)
- Inhibitor of dgtpase (10 kDa, Protein from Escherichia phage T7)
- Dgtp triphosphohydrolase (Complex from Escherichia coli str. K-12 substr. MG1655)
- Gene 1.2 protein (Complex from Escherichia phage T7)
Structure of E. coli dGTPase bound to T7 bacteriophage protein Gp1.2 and GTP
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 7u67
Q-score: 0.588
Klemm BP, Singh D, Smith CE, Hsu AL, Dillard LB, Krahn JM, London RE, Mueller GA, Borgnia MJ, Schaaper RM
PNAS (2022) 119 pp. e2123092119-e2123092119 [ DOI: doi:10.1073/pnas.2123092119 Pubmed: 36067314 ]
- Dgtpase hexamer bound to six copies of gp1.2 (Complex from Escherichia coli str. K-12 substr. MG1655)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Deoxyguanosinetriphosphate triphosphohydrolase (59 kDa, Protein from Escherichia coli str. K-12 substr. MG1655)
- Inhibitor of dgtpase (10 kDa, Protein from Escherichia phage T7)
- Dgtp triphosphohydrolase (Complex from Escherichia coli str. K-12 substr. MG1655)
- Gene 1.2 protein (Complex from Escherichia phage T7)
Structure of the L. blandensis dGTPase in the apo form
Resolution: 2.1 Å
EM Method: Single-particle
Fitted PDBs: 7tu5
Q-score: 0.651
Klemm BP, Sikkema AP, Hsu AL, Horng JC, Hall TMT, Borgnia MJ, Schaaper RM
J Biol Chem (2022) 298 pp. 102073-102073 [ Pubmed: 35643313 DOI: doi:10.1016/j.jbc.2022.102073 ]
- Magnesium ion (24 Da, Ligand)
- Water (18 Da, Ligand)
- Dgtp triphosphohydrolase (52 kDa, Protein from Leeuwenhoekiella blandensis MED217)
- Dgtp triphosphohydrolase (Complex from Leeuwenhoekiella blandensis MED217)
Structure of the L. blandensis dGTPase bound to dATP
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 7tu6
Q-score: 0.556
Klemm BP, Sikkema AP, Hsu AL, Horng JC, Hall TMT, Borgnia MJ, Schaaper RM
J Biol Chem (2022) 298 pp. 102073-102073 [ Pubmed: 35643313 DOI: doi:10.1016/j.jbc.2022.102073 ]
- 2'-deoxyadenosine 5'-triphosphate (491 Da, Ligand)
- Dgtp triphosphohydrolase (52 kDa, Protein from Leeuwenhoekiella blandensis MED217)
- Magnesium ion (24 Da, Ligand)
- Dgtp triphosphohydrolase (Complex from Leeuwenhoekiella blandensis MED217)
Structure of the L. blandensis dGTPase H125A mutant bound to dGTP
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 7tu7
Q-score: 0.637
Klemm BP, Sikkema AP, Hsu AL, Horng JC, Hall TMT, Borgnia MJ, Schaaper RM
J Biol Chem (2022) 298 pp. 102073-102073 [ Pubmed: 35643313 DOI: doi:10.1016/j.jbc.2022.102073 ]
- Dgtp triphosphohydrolase (52 kDa, Protein from Leeuwenhoekiella blandensis MED217)
- Magnesium ion (24 Da, Ligand)
- 2'-deoxyguanosine-5'-triphosphate (507 Da, Ligand)
- Dgtp triphosphohydrolase (Complex from Leeuwenhoekiella blandensis MED217)
Structure of the L. blandensis dGTPase H125A mutant bound to dGTP and dATP
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 7tu8
Q-score: 0.623
Klemm BP, Sikkema AP, Hsu AL, Horng JC, Hall TMT, Borgnia MJ, Schaaper RM
J Biol Chem (2022) 298 pp. 102073-102073 [ Pubmed: 35643313 DOI: doi:10.1016/j.jbc.2022.102073 ]
- 2'-deoxyadenosine 5'-triphosphate (491 Da, Ligand)
- Dgtp triphosphohydrolase (Complex from Leeuwenhoekiella blandensis MED217)
- Dgtp triphosphohydrolase (52 kDa, Protein from Leeuwenhoekiella blandensis MED217)
- 2'-deoxyguanosine-5'-triphosphate (507 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
Cryo-EM map of DH851.3 bound to HIV-1 CH505 Env
Resolution: 5.6 Å
EM Method: Single-particle
Fitted PDBs: 7lu9
Q-score: 0.22
Williams WB, Meyerhoff RR, Edwards RJ, Li H, Manne K, Nicely NI, Henderson R, Zhou Y, Janowska K, Mansouri K, Gobeil S, Evangelous T, Hora B, Berry M, Abuahmad AY, Sprenz J, Deyton M, Stalls V, Kopp M, Hsu AL, Borgnia MJ, Stewart-Jones GBE, Lee MS, Bronkema N, Moody MA, Wiehe K, Bradley T, Alam SM, Parks RJ, Foulger A, Oguin T, Sempowski GD, Bonsignori M, LaBranche CC, Montefiori DC, Seaman M, Santra S, Perfect J, Francica JR, Lynn GM, Aussedat B, Walkowicz WE, Laga R, Kelsoe G, Saunders KO, Fera D, Kwong PD, Seder RA, Bartesaghi A, Shaw GM, Acharya P, Haynes BF
Cell (2021) 184 pp. 2955- [ DOI: doi:10.1016/j.cell.2021.04.042 Pubmed: 34019795 ]
- Envelope glycoprotein gp120 (51 kDa, Protein from Human immunodeficiency virus 1)
- Dh851.3 bound to hiv-1 ch505 env (Complex from Human immunodeficiency virus 1)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Envelope glycoprotein gp41 (17 kDa, Protein from Human immunodeficiency virus 1)
- Dh851.3 heavy chain (24 kDa, Protein from Macaca mulatta)
- Dh851.3 light chain (21 kDa, Protein from Macaca mulatta)
- Dh851.3 light chain (22 kDa, Protein from Macaca mulatta)
- Dh851.3 heavy chain (23 kDa, Protein from Macaca mulatta)
Cryo-EM map of DH898.1 Fab-dimer bound near the CD4 binding site of HIV-1 Env CH848 SOSIP trimer
Resolution: 4.7 Å
EM Method: Single-particle
Fitted PDBs: 7lua
Q-score: 0.267
Williams WB, Meyerhoff RR, Edwards RJ, Li H, Manne K, Nicely NI, Henderson R, Zhou Y, Janowska K, Mansouri K, Gobeil S, Evangelous T, Hora B, Berry M, Abuahmad AY, Sprenz J, Deyton M, Stalls V, Kopp M, Hsu AL, Borgnia MJ, Stewart-Jones GBE, Lee MS, Bronkema N, Moody MA, Wiehe K, Bradley T, Alam SM, Parks RJ, Foulger A, Oguin T, Sempowski GD, Bonsignori M, LaBranche CC, Montefiori DC, Seaman M, Santra S, Perfect J, Francica JR, Lynn GM, Aussedat B, Walkowicz WE, Laga R, Kelsoe G, Saunders KO, Fera D, Kwong PD, Seder RA, Bartesaghi A, Shaw GM, Acharya P, Haynes BF
Cell (2021) 184 pp. 2955- [ DOI: doi:10.1016/j.cell.2021.04.042 Pubmed: 34019795 ]
- Ch848 sosip gp120 (52 kDa, Protein from Human immunodeficiency virus 1)
- Dh898.1 heavy chain (23 kDa, Protein from Homo sapiens)
- Dh898.1 light chain (23 kDa, Protein from Homo sapiens)
- Cryo-em structure of dh898.1 fab-dimer bound near the cd4 binding site of hiv-1 env ch848 sosip trimer. (Complex from Human immunodeficiency virus 1)
- Env polyprotein (16 kDa, Protein from Simian-Human immunodeficiency virus)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)