Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
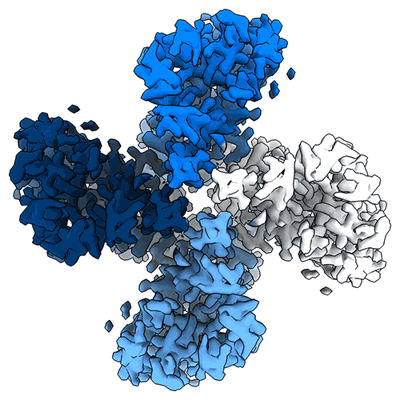
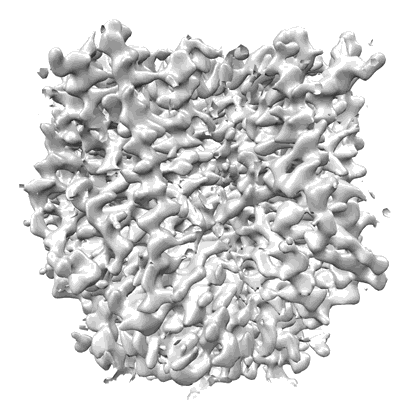
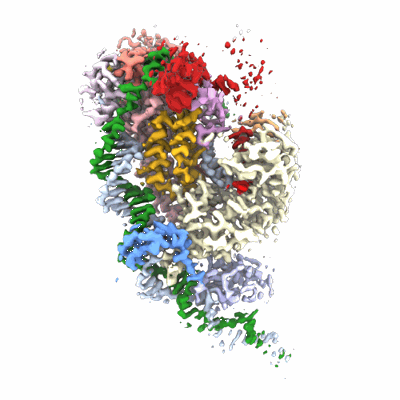
Structure of the insect gustatory receptor Gr9 from Bombyx mori
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 8uvt
Q-score: 0.547
Gomes JV, Singh-Bhagania S, Cenci M, Chacon Cordon C, Singh M, Butterwick JA
Nature (2024) 629 pp. 228-234 [ Pubmed: 38447670 DOI: doi:10.1038/s41586-024-07255-w ]
- Gustatory receptor (50 kDa, Protein from Bombyx mori)
- Homotetramer of gr9 in the absence of ligand (Complex from Bombyx mori)
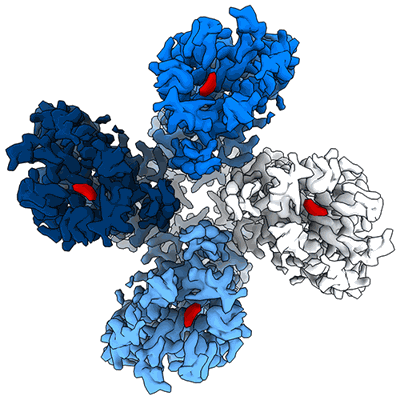
Structure of the insect gustatory receptor Gr9 from Bombyx mori in complex with D-fructose
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 8uvu
Q-score: 0.551
Gomes JV, Singh-Bhagania S, Cenci M, Chacon Cordon C, Singh M, Butterwick JA
Nature (2024) 629 pp. 228-234 [ Pubmed: 38447670 DOI: doi:10.1038/s41586-024-07255-w ]
- Beta-d-fructofuranose (180 Da, Ligand)
- Gustatory receptor (50 kDa, Protein from Bombyx mori)
- Beta-d-fructopyranose (180 Da, Ligand)
- Homotetramer of gr9 bound to d-fructose (Complex from Bombyx mori)
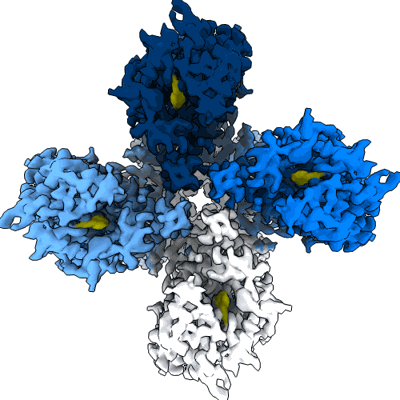
Structure of the insect gustatory receptor Gr9 from Bombyx mori in complex with L-sorbose
Resolution: 2.61 Å
EM Method: Single-particle
Fitted PDBs: 8vv3
Q-score: 0.563
Gomes JV, Singh-Bhagania S, Cenci M, Chacon Cordon C, Singh M, Butterwick JA
Nature (2024) 629 pp. 228-234 [ Pubmed: 38447670 DOI: doi:10.1038/s41586-024-07255-w ]
- Gustatory receptor (50 kDa, Protein from Bombyx mori)
- Homotetramer of gr9 bound to l-sorbose (Complex from Bombyx mori)
- Alpha-l-sorbopyranose (180 Da, Ligand)
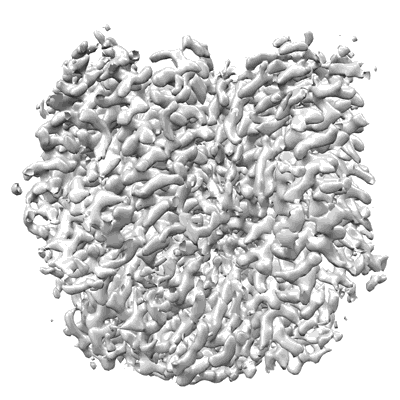
CryoEM structure of insect gustatory receptor BmGr9
Resolution: 2.85 Å
EM Method: Single-particle
Fitted PDBs: 8vc1
Q-score: 0.539
Frank HM, Walujkar S, Walsh RM, Laursen WJ, Theobald DL, Garrity PA, Gaudet R
bioRxiv (2023) [ Pubmed: 38187590 DOI: doi:10.1101/2023.12.19.572336 ]
- Agonist-free bmgr9 homotetramer (218 kDa, Complex from Bombyx mori)
- (3beta,14beta,17beta,25r)-3-[4-methoxy-3-(methoxymethyl)butoxy]spirost-5-en (544 Da, Ligand)
- (7s)-4-hydroxy-n,n,n-trimethyl-9-oxo-7-[(palmitoyloxy)methyl]-3,5,8-trioxa-4-phosphahexacosan-1-aminium 4-oxide (763 Da, Ligand)
- Gustatory receptor (54 kDa, Protein from Bombyx mori)
- (7r,17e,20e)-4-hydroxy-n,n,n-trimethyl-9-oxo-7-[(palmitoyloxy)methyl]-3,5,8-trioxa-4-phosphahexacosa-17,20-dien-1-aminium 4-oxide (759 Da, Ligand)
CryoEM structure of insect gustatory receptor BmGr9 in the presence of fructose
Resolution: 3.98 Å
EM Method: Single-particle
Fitted PDBs: 8vc2
Q-score: 0.376
Frank HM, Walujkar S, Walsh RM, Laursen WJ, Theobald DL, Garrity PA, Gaudet R
bioRxiv (2023) [ Pubmed: 38187590 DOI: doi:10.1101/2023.12.19.572336 ]
Structure of R2 with 3'UTR and DNA in binding state
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 8ibw
Q-score: 0.385
Deng P, Tan SQ, Yang QY, Fu L, Wu Y, Zhu HZ, Sun L, Bao Z, Lin Y, Zhang QC, Wang H, Wang J, Liu JG
Cell (2023) 186 pp. 2865-2879.e20 [ Pubmed: 37301196 DOI: doi:10.1016/j.cell.2023.05.032 ]
- Reverse transcriptase-like protein (123 kDa, Protein from Bombyx mori)
- R2 complex (177 kDa, Complex from Bombyx mori)
- Dna (60-mer) (18 kDa, DNA from Bombyx mori)
- 3'utr (16 kDa, RNA from Bombyx mori)
- Dna (60-mer) (18 kDa, DNA from Bombyx mori)
- Zinc ion (65 Da, Ligand)
Structure of R2 with 3'UTR and DNA in unwinding state
Resolution: 3.74 Å
EM Method: Single-particle
Fitted PDBs: 8ibx
Q-score: 0.385
Deng P, Tan SQ, Yang QY, Fu L, Wu Y, Zhu HZ, Sun L, Bao Z, Lin Y, Zhang QC, Wang H, Wang J, Liu JG
Cell (2023) 186 pp. 2865-2879.e20 [ Pubmed: 37301196 DOI: doi:10.1016/j.cell.2023.05.032 ]
- Reverse transcriptase-like protein (123 kDa, Protein from Bombyx mori)
- R2 complex (177 kDa, Complex from Bombyx mori)
- Dna (60-mer) (18 kDa, DNA from Bombyx mori)
- 3'utr (16 kDa, RNA from Bombyx mori)
- Dna (60-mer) (18 kDa, DNA from Bombyx mori)
- Zinc ion (65 Da, Ligand)
Resolution: 3.47 Å
EM Method: Single-particle
Fitted PDBs: 8iby
Q-score: 0.342
Deng P, Tan SQ, Yang QY, Fu L, Wu Y, Zhu HZ, Sun L, Bao Z, Lin Y, Zhang QC, Wang H, Wang J, Liu JG
Cell (2023) 186 pp. 2865-2879.e20 [ Pubmed: 37301196 DOI: doi:10.1016/j.cell.2023.05.032 ]
- Zinc ion (65 Da, Ligand)
- Reverse transcriptase-like protein (123 kDa, Protein from Bombyx mori)
- R2 complex (177 kDa, Complex from Bombyx mori)
- 5'orf rna (72 kDa, RNA from Bombyx mori)
Structure of R2 with 5'ORF and 3'UTR
Resolution: 3.04 Å
EM Method: Single-particle
Fitted PDBs: 8ibz
Q-score: 0.435
Deng P, Tan SQ, Yang QY, Fu L, Wu Y, Zhu HZ, Sun L, Bao Z, Lin Y, Zhang QC, Wang H, Wang J, Liu JG
Cell (2023) 186 pp. 2865-2879.e20 [ Pubmed: 37301196 DOI: doi:10.1016/j.cell.2023.05.032 ]
- Zinc ion (65 Da, Ligand)
- Reverse transcriptase-like protein (123 kDa, Protein from Bombyx mori)
- R2 complex (177 kDa, Complex from Bombyx mori)
- 5orf-linker-3utr (65 kDa, RNA from Bombyx mori)
Bombyx mori R2 retrotransposon initiating target-primed reverse transcription
Resolution: 3.08 Å
EM Method: Single-particle
Fitted PDBs: 8gh6
Q-score: 0.49
Wilkinson ME, Frangieh CJ, Macrae RK, Zhang F
Science (2023) 380 pp. 301-308 [ DOI: doi:10.1126/science.adg7883 Pubmed: 37023171 ]
- 28s dna top strand (21 kDa, DNA from Bombyx mori)
- Thymidine-5'-triphosphate (482 Da, Ligand)
- 28s dna bottom strand, 5' side (priming strand) (7 kDa, DNA from Bombyx mori)
- Magnesium ion (24 Da, Ligand)
- R2bm 3'utr rna (81 kDa, RNA from Bombyx mori)
- Reverse transcriptase-like protein (123 kDa, Protein from Bombyx mori)
- Bombyx mori r2 retrotransposon initiating target-primed reverse transcription (230 kDa, Complex from Bombyx mori)
- 28s dna bottom strand, 3' side (21 kDa, DNA from Bombyx mori)
- Zinc ion (65 Da, Ligand)