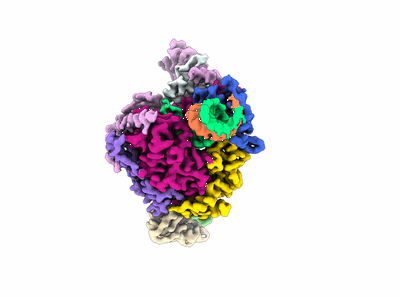
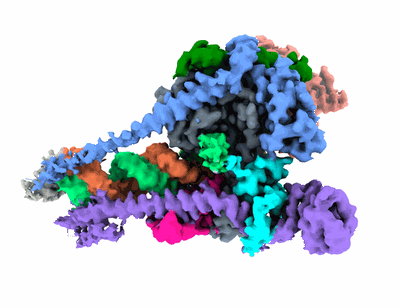
Cryo-EM structure of human NatB in complex with CoA-Alpha-Synuclein
Resolution: 3.14 Å
EM Method: Single-particle
Fitted PDBs: 7stx
Q-score: 0.4
To Be Published
- Mdvfm peptide (Complex from Homo sapiens)
- N-alpha-acetyltransferase 25, natb auxiliary subunit (109 kDa, Protein from Homo sapiens)
- N-alpha-acetyltransferase 20, n-alpha-acetyltransferase 25, natb auxiliary subunit (Complex from Homo sapiens)
- N-alpha-acetyltransferase 20 (20 kDa, Protein from Homo sapiens)
- Acetyl group (44 Da, Ligand)
- Coenzyme a (767 Da, Ligand)
- Human natb complex (Complex)
- Alpha-synuclein (641 Da, Protein from Homo sapiens)
Cfr-modified 50S subunit from Escherichia coli
Resolution: 2.2 Å
EM Method: Single-particle
Fitted PDBs: 7lvk
Q-score: 0.646
Tsai K, Stojkovic V, Noda-Garcia L, Myasnikov AG, Young ID, Palla A, Venkataramanan S, Floor S, Fraser JS, Frost A, Tawfik DS, Fujimori DG
To Be Published
- 50s ribosomal protein l20 (13 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l23 (11 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l22 (12 kDa, Protein from Escherichia coli)
- 23s rrna (941 kDa, RNA from Escherichia coli)
- 50s ribosomal protein l36 (4 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l13 (16 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l14 (13 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l27 (9 kDa, Protein from Escherichia coli)
- Sodium ion (22 Da, Ligand)
- 50s ribosomal protein l4 (22 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l17 (14 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l5 (20 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l28 (9 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l9 (15 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l18 (12 kDa, Protein from Escherichia coli)
- Zinc ion (65 Da, Ligand)
- Escherichia coli 50s subunit (Complex from Escherichia coli)
- 50s ribosomal protein l24 (11 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l34 (5 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l21 (11 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l19 (13 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l35 (7 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l33 (6 kDa, Protein from Escherichia coli)
- Water (18 Da, Ligand)
- 50s ribosomal protein l3 (22 kDa, Protein from Escherichia coli)
- Magnesium ion (24 Da, Ligand)
- 50s ribosomal protein l6 (18 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l32 (6 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l15 (15 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l2 (29 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l30 (6 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l25 (10 kDa, Protein from Escherichia coli)
- 5s rrna (38 kDa, RNA from Escherichia coli)
- 50s ribosomal protein l29 (7 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l16 (15 kDa, Protein from Escherichia coli)
Capsid structure of human sapovirus
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 7dod
Q-score: 0.525
Miyazaki N, Murakami K, Oka T, Iwasaki K, Katayama K, Murata K
To Be Published
Structure of COVID-19 RNA-dependent RNA polymerase bound to penciclovir.
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 7doi
Q-score: 0.599
Yin W, Luan X, Li Z, Xie Y, Zhou Z, Liu J, Gao M, Wang X, Zhou F, Wang Q, Wang Q, Shen D, Zhang Y, Tian G, Aisa H, Wei D, Jiang Y, Xiao G, Jiang H, Zhang L, Yu X, Shen J, Zhang S, Xu H
To Be Published
- Covid-19 rdrp complex bound to penciclovir (Complex from Severe acute respiratory syndrome coronavirus 2)
- Pyrophosphate 2- (175 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (107 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*cp*cp*up*ap*up*ap*ap*cp*up*up*ap*ap*up*cp*u)-3') (4 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- [(2r)-4-(2-azanyl-6-oxidanylidene-3h-purin-9-yl)-2-(hydroxymethyl)butyl] dihydrogen phosphate (333 Da, Ligand)
- Water (18 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Rna (5'-r(p*ap*gp*ap*up*up*ap*ap*gp*up*up*ap*u)-3') (3 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
Structure of COVID-19 RNA-dependent RNA polymerase (extended conformation) bound to penciclovir
Resolution: 2.73 Å
EM Method: Single-particle
Fitted PDBs: 7dok
Q-score: 0.554
Yin W, Luan X, Li Z, Xie Y, Zhou Z, Liu J, Gao M, Wang X, Zhou F, Wang Q, Wang Q, Shen D, Zhang Y, Tian G, Aisa H, Wei D, Jiang Y, Xiao G, Jiang H, Zhang L, Yu X, Shen J, Zhang S, Xu H
To Be Published
- Covid-19 rdrp complex (exntended conformation) bound to penciclovir (Complex from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*gp*cp*up*ap*up*gp*up*gp*ap*gp*ap*up*up*ap*ap*gp*up*up*ap*u)-3') (6 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Pyrophosphate 2- (175 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (107 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*cp*cp*cp*up*ap*up*ap*ap*cp*up*up*ap*ap*up*cp*up*cp*ap*cp*ap*up*ap*gp*c)-3') (7 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- [(2r)-4-(2-azanyl-6-oxidanylidene-3h-purin-9-yl)-2-(hydroxymethyl)butyl] dihydrogen phosphate (333 Da, Ligand)
- Water (18 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Zinc ion (65 Da, Ligand)
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 6x96
Q-score: 0.45
Ozorowski G, Cottrell CA, de Val N, Scaringi N, Polveroni TM, Ward AB
To Be Published
- Bg505 hiv-1 env gp120 (57 kDa, Protein from Human immunodeficiency virus 1)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Monoclonal antibody 10a kappa chain (25 kDa, Protein from Oryctolagus cuniculus)
- Bg505 hiv-1 env gp41 (17 kDa, Protein from Human immunodeficiency virus 1)
- Hiv-1 env bg505 sosip.664 in complex with rabbit monoclonal antibody 10a fragment antigen binding (570 kDa, Complex)
- Hiv-1 env bg505 sosip.664 (Complex from Human immunodeficiency virus 1)
- Rabbit monoclonal antibody 10a fragment antigen (Complex from Oryctolagus cuniculus)
- Monoclonal antibody 10a fragment antigen binding heavy chain (24 kDa, Protein from Oryctolagus cuniculus)
Resolution: 3.65 Å
EM Method: Single-particle
Fitted PDBs: 6x97
Q-score: 0.427
Ozorowski G, Cottrell CA, de Val N, Scaringi N, Polveroni TM, Ward AB
To Be Published
- Bg505 hiv-1 env gp120 (57 kDa, Protein from Human immunodeficiency virus 1)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Bg505 hiv-1 env gp41 (17 kDa, Protein from Human immunodeficiency virus 1)
- Rabbit monoclonal antibody 11a fragment antigen (Complex from Oryctolagus cuniculus)
- Hiv-1 env bg505 sosip.664 (Complex from Human immunodeficiency virus 1)
- Hiv-1 env bg505 sosip.664 in complex with rabbit monoclonal antibody 11a fragment antigen binding (570 kDa, Complex)
- Monoclonal antibody 11a fragment antigen binding heavy chain (25 kDa, Protein from Oryctolagus cuniculus)
- Monoclonal antibody 11a kappa chain (24 kDa, Protein from Oryctolagus cuniculus)
Resolution: 3.38 Å
EM Method: Single-particle
Fitted PDBs: 6x98
Q-score: 0.475
Ozorowski G, Cottrell CA, de Val N, Scaringi N, Polveroni TM, Ward AB
To Be Published
- Bg505 hiv-1 env gp120 (57 kDa, Protein from Human immunodeficiency virus 1)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Rabbit monoclonal antibody 11b fragment antigen (Complex from Oryctolagus cuniculus)
- Bg505 hiv-1 env gp41 (17 kDa, Protein from Human immunodeficiency virus 1)
- Hiv-1 env bg505 sosip.664 (Complex from Human immunodeficiency virus 1)
- Monoclonal antibody 11b kappa chain (25 kDa, Protein from Oryctolagus cuniculus)
- Hiv-1 env bg505 sosip.664 in complex with rabbit monoclonal antibody 11b fragment antigen binding (570 kDa, Complex)
- Monoclonal antibody 11b fragment antigen binding heavy chain (25 kDa, Protein from Oryctolagus cuniculus)
Cytochrome c oxidase structure in P-state
Resolution: 2.4 Å
EM Method: Single-particle
Fitted PDBs: 7ate
Q-score: 0.749
To Be Published
- Cytochrome c oxidase subunit 3 (30 kDa, Protein from Paracoccus denitrificans)
- Dinuclear copper ion (127 Da, Ligand)
- Cytochrome c oxidase subunit 4 (5 kDa, Protein from Paracoccus denitrificans)
- Manganese (ii) ion (54 Da, Ligand)
- 1,2-diacyl-sn-glycero-3-phosphocholine (790 Da, Ligand)
- Cytochrome c oxidase with four subunits reconstituted in lipid nanodisc (127 kDa, Complex from Paracoccus denitrificans)
- Water (18 Da, Ligand)
- Copper (ii) ion (63 Da, Ligand)
- Calcium ion (40 Da, Ligand)
- (1r)-2-{[{[(2s)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(palmitoyloxy)methyl]ethyl (11e)-octadec-11-enoate (749 Da, Ligand)
- Cytochrome c oxidase subunit 2 (32 kDa, Protein from Paracoccus denitrificans)
- Cytochrome c oxidase subunit 1-beta (62 kDa, Protein from Paracoccus denitrificans)
- Heme-a (852 Da, Ligand)
- Hydrogen peroxide (34 Da, Ligand)
Cytochrome c oxidase structure in R-state
Resolution: 2.66 Å
EM Method: Single-particle
Fitted PDBs: 7atn
Q-score: 0.665
To Be Published
- Cytochrome c oxidase subunit 3 (30 kDa, Protein from Paracoccus denitrificans)
- Dinuclear copper ion (127 Da, Ligand)
- Cytochrome c oxidase subunit 4 (5 kDa, Protein from Paracoccus denitrificans)
- Manganese (ii) ion (54 Da, Ligand)
- 1,2-diacyl-sn-glycero-3-phosphocholine (790 Da, Ligand)
- Cytochrome c oxidase with four subunits reconstituted in lipid nanodisc (127 kDa, Complex from Paracoccus denitrificans)
- Water (18 Da, Ligand)
- Copper (ii) ion (63 Da, Ligand)
- Calcium ion (40 Da, Ligand)
- Cytochrome c oxidase subunit 2 (32 kDa, Protein from Paracoccus denitrificans)
- Cytochrome c oxidase subunit 1-beta (62 kDa, Protein from Paracoccus denitrificans)
- Heme-a (852 Da, Ligand)