Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
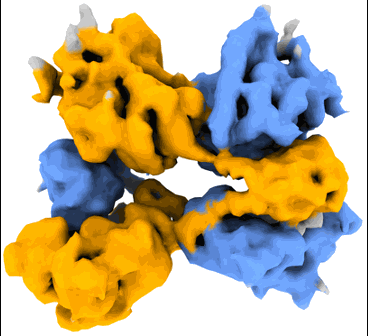
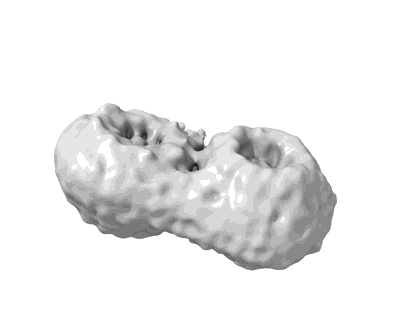
P116 (amino acids 246-818) dimer in the full state
To Be Published
Structure of IgE HMM5 bound to FceRIa cryo-EM class 8
Resolution: 6.9 Å
EM Method: Single-particle
Fitted PDBs: 9eq3
Q-score: 0.136
To Be Published
- Ige hmm5 heavy chain (60 kDa, Protein from Homo sapiens)
- High affinity immunoglobulin epsilon receptor subunit alpha (19 kDa, Protein from Homo sapiens)
- Ige hmm5 light chain (23 kDa, Protein from Homo sapiens)
- Complex of ige hmm5 and the ectodomain of fceria (210 kDa, Complex from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
Resolution: 7.61 Å
EM Method: Single-particle
Fitted PDBs: 8xbv
Q-score: 0.066
Shioi T, Hatazawa S, Oya E, Hosoya N, Kobayashi W, Ogasawara M, Kobayashi T, Takizawa Y, Kurumizaka H
Nature (2024) 628 pp. 212-220 [ Pubmed: 38509361 DOI: doi:10.1038/s41586-024-07196-4 ]
- Dna (5'-d(p*cp*gp*ap*ap*ap*ap*cp*gp*gp*cp*cp*ap*cp*cp*a)-3') (47 kDa, DNA from synthetic construct)
- Dna repair protein rad51 homolog 1 (37 kDa, Protein from Homo sapiens)
- Dna (5'-d(p*tp*gp*gp*cp*cp*gp*tp*tp*tp*tp*cp*g)-3') (47 kDa, DNA from synthetic construct)
- Rad51-nucleosome complex (Complex from Homo sapiens)
Resolution: 7.8 Å
EM Method: Single-particle
Fitted PDBs: 8xby
Q-score: 0.061
Shioi T, Hatazawa S, Oya E, Hosoya N, Kobayashi W, Ogasawara M, Kobayashi T, Takizawa Y, Kurumizaka H
Nature (2024) 628 pp. 212-220 [ Pubmed: 38509361 DOI: doi:10.1038/s41586-024-07196-4 ]
- Dna (5'-d(p*ap*ap*cp*gp*ap*ap*ap*ap*cp*gp*gp*cp*cp*ap*cp*cp*ap*cp*g)-3') (48 kDa, DNA from synthetic construct)
- Dna repair protein rad51 homolog 1 (37 kDa, Protein from Homo sapiens)
- Dna (5'-d(p*cp*gp*tp*gp*gp*tp*gp*gp*cp*cp*gp*tp*tp*tp*tp*cp*gp*tp*t)-3') (48 kDa, DNA from synthetic construct)
- Rad51-nucleosome complex (Complex from Homo sapiens)
Double-motif Single-stranded Paranemic Crossover RNA Triangle (2PXT)
Resolution: 6.56 Å
EM Method: Single-particle
Fitted PDBs: 8bu8
Q-score: 0.182
Sampedro Vallina N, McRae EKS, Geary C, Andersen ES
Small (2023) 19 pp. e2204651-e2204651 [ Pubmed: 36526605 DOI: doi:10.1002/smll.202204651 ]
- Rna (354-mer) (113 kDa, RNA from synthetic construct)
- Double-motif single-stranded paranemic crossover rna triangle (2pxt) (113 MDa, Complex from synthetic construct)
The cryo-EM structure of OsCyc1 dimer state
Resolution: 7.9 Å
EM Method: Single-particle
Fitted PDBs: 8i6u
Q-score: 0.173
Resolution: 6.5 Å
EM Method: Single-particle
Fitted PDBs: 8dme
Q-score: 0.147
To Be Published
- Flavin-adenine dinucleotide (785 Da, Ligand)
- Sulfate ion (96 Da, Ligand)
- Flavin mononucleotide (456 Da, Ligand)
- 6-methoxy-2-{[(4-methoxy-3,5-dimethylpyridin-2-yl)methyl]sulfanyl}-1h-benzimidazole (329 Da, Ligand)
- Tetraethylene glycol (194 Da, Ligand)
- Bifunctional cytochrome p450/nadph--p450 reductase (116 kDa, Protein from Priestia megaterium NBRC 15308 = ATCC 14581)
- Cyp102a1 (240 kDa, Complex from Priestia megaterium NBRC 15308 = ATCC 14581)
- Protoporphyrin ix containing fe (616 Da, Ligand)
LH2-LH3 antenna in antiparallel configuration embedded in a nanodisc
Resolution: 6.4 Å
EM Method: Single-particle
Fitted PDBs: 8fb9
Q-score: 0.164
Wang D, Fiebig OC, Harris D, Toporik H, Ji Y, Chuang C, Nairat M, Tong AL, Ogren JI, Hart SM, Cao J, Sturgis JN, Mazor Y, Schlau-Cohen GS
PNAS (2023) 120 pp. e2220477120-e2220477120 [ Pubmed: 37399405 DOI: doi:10.1073/pnas.2220477120 ]
- Bacteriochlorophyll a (911 Da, Ligand)
- Light-harvesting protein b800-820 alpha chain (6 kDa, Protein from Magnetospirillum molischianum)
- Light-harvesting protein b800-820 beta chain (5 kDa, Protein from Magnetospirillum molischianum)
- Lycopene (536 Da, Ligand)
- Light-harvesting protein b-800/850 beta 1 chain (5 kDa, Protein from Magnetospirillum molischianum)
- Light-harvesting protein b-800/850 alpha chain (5 kDa, Protein from Magnetospirillum molischianum)
- Lhii-lhiii complex in nanodisk (200 kDa, Complex from Magnetospirillum molischianum)
Cryo-EM model of the marine siphophage vB_DshS-R4C stopper-terminator complex
Resolution: 6.6 Å
EM Method: Single-particle
Fitted PDBs: 8gtf
Q-score: 0.16
Huang Y, Sun H, Wei S, Cai L, Liu L, Jiang Y, Xin J, Chen Z, Que Y, Kong Z, Li T, Yu H, Zhang J, Gu Y, Zheng Q, Li S, Zhang R, Xia N
Nat Commun (2023) 14 pp. 3609-3609 [ Pubmed: 37330604 DOI: doi:10.1038/s41467-023-39220-y ]
- Head-to-tail joining protein (10 kDa, Protein from Dinoroseobacter phage vB DshS-R4C)
- Major tail protein (13 kDa, Protein from Dinoroseobacter phage vB DshS-R4C)
- Terminator protein (14 kDa, Protein from Dinoroseobacter phage vB DshS-R4C)
- Dinoroseobacter phage vb_dshs-r4c (Virus from Dinoroseobacter phage vB DshS-R4C)
Complex of RecO-RecR-DNA from Thermus thermophilus.
Resolution: 6.09 Å
EM Method: Single-particle
Fitted PDBs: 8ab0
Q-score: 0.233
Nirwal S, Czarnocki-Cieciura M, Chaudhary A, Zajko W, Skowronek K, Chamera S, Figiel M, Nowotny M
Nat Struct Mol Biol (2023) 30 pp. 650-660 [ DOI: doi:10.1038/s41594-023-00967-z Pubmed: 37081315 ]
- Recombination protein recr (21 kDa, Protein from Thermus thermophilus HB8)
- Oligo1 (7 kDa, DNA from synthetic construct)
- Oligo2 (12 kDa, DNA from synthetic construct)
- Zinc ion (65 Da, Ligand)
- Complex of reco-recr-dna from thermus thermophilus. (130 kDa, Complex from Thermus thermophilus HB8)
- Dna repair protein reco (24 kDa, Protein from Thermus thermophilus HB8)