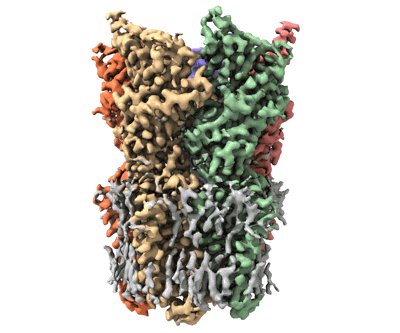
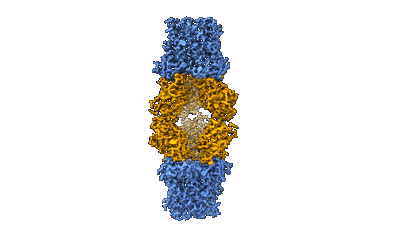
FCP trimer in diatom Thalassiosira pseudonana
Feng Y, Li Z, Zhou CC, Wang W, Shen JR
To Be Published
- 1,2-distearoyl-monogalactosyl-diglyceride (787 Da, Ligand)
- Chlorophyll c1 (610 Da, Ligand)
- Chlorophyll c2 (608 Da, Ligand)
- Fcp trimer (Complex from Thalassiosira pseudonana CCMP1335)
- Fucoxanthin chlorophyll a/c protein 8 (19 kDa, Protein from Thalassiosira pseudonana CCMP1335)
- (3s,3's,5r,5'r,6s,6'r,8'r)-3,5'-dihydroxy-8-oxo-6',7'-didehydro-5,5',6,6',7,8-hexahydro-5,6-epoxy-beta,beta-caroten-3'- yl acetate (658 Da, Ligand)
- Chlorophyll a (893 Da, Ligand)
Structure of CbCas9 bound to 20-nucleotide complementary DNA substrate
Resolution: 2.46 Å
EM Method: Single-particle
Fitted PDBs: 8iyq
Q-score: 0.513
Zhang S, Sun A, Qian JM, Lin S, Xing W, Yang Y, Zhu HZ, Zhou XY, Guo YS, Liu Y, Meng Y, Jin SL, Song W, Li CP, Li Z, Jin S, Wang JH, Dong MQ, Gao C, Chen C, Bai Y, Liu JG
Nature (2024) [ Pubmed: 38811729 DOI: doi:10.1038/s41586-024-07486-x ]
- Nts (8 kDa, DNA from Chryseobacterium)
- Deadcbcas9 (170 kDa, Protein from Chryseobacterium)
- Ts (8 kDa, DNA from Chryseobacterium)
- Sgrna (40 kDa, RNA from Chryseobacterium)
- Structure of cbcas9 bound to 20-nucleotide complementary dna substrate (228 kDa, Complex from Chryseobacterium)
Structure of CbCas9 bound to 6-nucleotide complementary DNA substrate
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8wmh
Q-score: 0.279
Zhang S, Sun A, Qian JM, Lin S, Xing W, Yang Y, Zhu HZ, Zhou XY, Guo YS, Liu Y, Meng Y, Jin SL, Song W, Li CP, Li Z, Jin S, Wang JH, Dong MQ, Gao C, Chen C, Bai Y, Liu JG
Nature (2024) [ Pubmed: 38811729 DOI: doi:10.1038/s41586-024-07486-x ]
- Nts (8 kDa, DNA from Chryseobacterium)
- Ts (8 kDa, DNA from Chryseobacterium)
- Deadcbcas9 (170 kDa, Protein from Chryseobacterium)
- Sgrna (40 kDa, RNA from Chryseobacterium)
- Structure of cbcas9 bound to 6-nucleotide complementary dna substrate (228 kDa, Complex from Chryseobacterium)
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 5.5
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 8i47
Q-score: 0.519
Bharambe N, Li Z, Seiferth D, Balakrishna AM, Biggin PC, Basak S
Nat Commun (2024) 15 pp. 2967-2967 [ DOI: doi:10.1038/s41467-024-47370-w Pubmed: 38580666 ]
- Water (18 Da, Ligand)
- Pentameric ligand-gated ion channel (Complex from Gloeobacter violaceus)
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Proton-gated ion channel (35 kDa, Protein from Gloeobacter violaceus)
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in closed state
Resolution: 2.74 Å
EM Method: Single-particle
Fitted PDBs: 8i48
Q-score: 0.547
Bharambe N, Li Z, Seiferth D, Balakrishna AM, Biggin PC, Basak S
Nat Commun (2024) 15 pp. 2967-2967 [ DOI: doi:10.1038/s41467-024-47370-w Pubmed: 38580666 ]
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Chloride ion (35 Da, Ligand)
- Water (18 Da, Ligand)
- Proton-gated ion channel (35 kDa, Protein from Gloeobacter violaceus)
- Pentameric ligand-gated ion channel (Complex from Gloeobacter violaceus)
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 2.5
Resolution: 2.6476 Å
EM Method: Single-particle
Fitted PDBs: 8jj3
Q-score: 0.624
Bharambe N, Li Z, Seiferth D, Balakrishna AM, Biggin PC, Basak S
Nat Commun (2024) 15 pp. 2967-2967 [ DOI: doi:10.1038/s41467-024-47370-w Pubmed: 38580666 ]
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Chloride ion (35 Da, Ligand)
- Water (18 Da, Ligand)
- Proton-gated ion channel (35 kDa, Protein from Gloeobacter violaceus)
- Pentameric ligand-gated ion channel (Complex from Gloeobacter violaceus)
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in intermediate state
Resolution: 3.35 Å
EM Method: Single-particle
Fitted PDBs: 8wcq
Q-score: 0.378
Bharambe N, Li Z, Seiferth D, Balakrishna AM, Biggin PC, Basak S
Nat Commun (2024) 15 pp. 2967-2967 [ DOI: doi:10.1038/s41467-024-47370-w Pubmed: 38580666 ]
- Pentameric ligand-gated ion channel (Complex from Gloeobacter violaceus)
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Chloride ion (35 Da, Ligand)
- Proton-gated ion channel (35 kDa, Protein from Gloeobacter violaceus)
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in open state
Resolution: 2.74 Å
EM Method: Single-particle
Fitted PDBs: 8wcr
Q-score: 0.625
Bharambe N, Li Z, Seiferth D, Balakrishna AM, Biggin PC, Basak S
Nat Commun (2024) 15 pp. 2967-2967 [ DOI: doi:10.1038/s41467-024-47370-w Pubmed: 38580666 ]
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Chloride ion (35 Da, Ligand)
- Water (18 Da, Ligand)
- Proton-gated ion channel (35 kDa, Protein from Gloeobacter violaceus)
- Pentameric ligand-gated ion channel (Complex from Gloeobacter violaceus)
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 7.5
Resolution: 2.92 Å
EM Method: Single-particle
Fitted PDBs: 8i42
Q-score: 0.503
Bharambe N, Li Z, Seiferth D, Balakrishna AM, Biggin PC, Basak S
Nat Commun (2024) 15 pp. 2967-2967 [ DOI: doi:10.1038/s41467-024-47370-w Pubmed: 38580666 ]
Cryo-EM structure of styrene oxide isomerase bound to benzylamine inhibitor
Resolution: 2.12 Å
EM Method: Single-particle
Fitted PDBs: 8pnu
Q-score: 0.708
Khanppnavar B, Choo JPS, Hagedoorn PL, Smolentsev G, Stefanic S, Kumaran S, Tischler D, Winkler FK, Korkhov VM, Li Z, Kammerer RA, Li X
Nat Chem (2024) [ DOI: doi:10.1038/s41557-024-01523-y Pubmed: 38744914 ]
- Protoporphyrin ix containing fe (616 Da, Ligand)
- Benzylamine (107 Da, Ligand)
- Nanobody (14 kDa, Protein from Vicugna pacos)
- Styrene oxide isomerase (19 kDa, Protein from Pseudomonas sp. VLB120)
- Water (18 Da, Ligand)
- Styrene oxide isomerase-nanobody-benzylamine complex (201 MDa, Complex from Pseudomonas sp. VLB120)