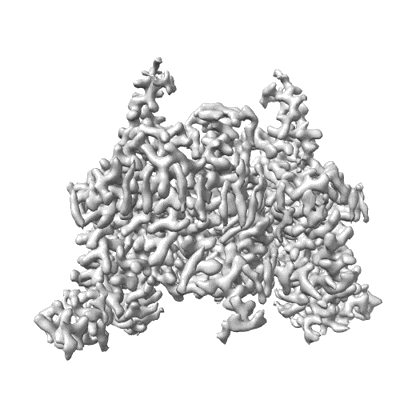Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
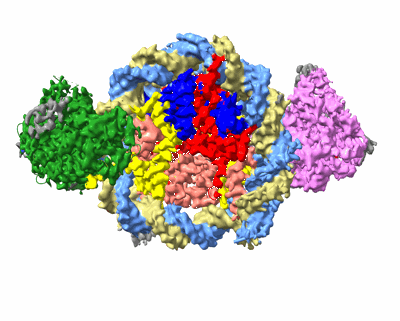
Structure of DDM1-nucleosome complex in the apo state
Liu Y, Zhang Z, Hu H, Chen W, Zhang F, Wang Q, Wang C, Yan K, Du J
Nat Plants (2024) 10 pp. 374-380 [ Pubmed: 38413824 DOI: doi:10.1038/s41477-024-01640-z ]
- Ddm1-nucleosome complex in the apo state (290 kDa, Complex from Arabidopsis thaliana)
- Histone h4 (11 kDa, Protein from Arabidopsis thaliana)
- Histone h3.1 (15 kDa, Protein from Arabidopsis thaliana)
- Dna (sense strand) (51 kDa, DNA from synthetic construct)
- Atp-dependent dna helicase ddm1 (86 kDa, Protein from Arabidopsis thaliana)
- Dna (antisense strand) (51 kDa, DNA from synthetic construct)
- Histone h2b.6 (16 kDa, Protein from Arabidopsis thaliana)
- Histone h2a.6 (13 kDa, Protein from Arabidopsis thaliana)
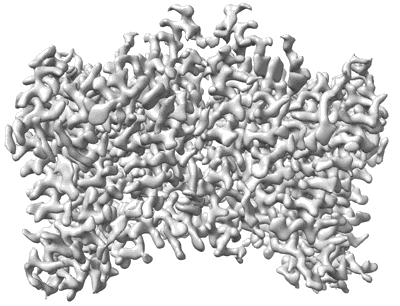
Structure of DDM1-nucleosome complex in ADP state
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 8wh8
Q-score: 0.403
Liu Y, Zhang Z, Hu H, Chen W, Zhang F, Wang Q, Wang C, Yan K, Du J
Nat Plants (2024) 10 pp. 374-380 [ Pubmed: 38413824 DOI: doi:10.1038/s41477-024-01640-z ]
- Histone h4 (11 kDa, Protein from Arabidopsis thaliana)
- Histone h3.1 (15 kDa, Protein from Arabidopsis thaliana)
- Atp-dependent dna helicase ddm1 (86 kDa, Protein from Arabidopsis thaliana)
- Dna (antisense strand) (45 kDa, DNA from synthetic construct)
- Ddm1-nucleosome complex in adp state (290 kDa, Complex from Arabidopsis thaliana)
- Dna (sense strand) (45 kDa, DNA from synthetic construct)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Histone h2b.6 (16 kDa, Protein from Arabidopsis thaliana)
- Histone h2a.6 (13 kDa, Protein from Arabidopsis thaliana)
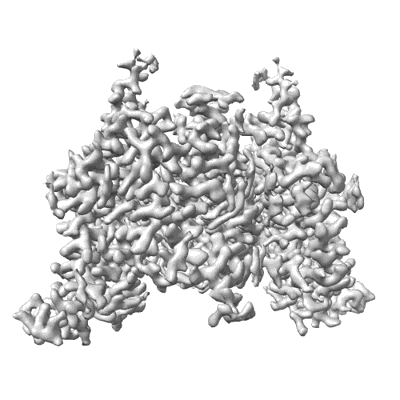
Structure of DDM1-nucleosome complex in ADP-BeFx state
Resolution: 3.31 Å
EM Method: Single-particle
Fitted PDBs: 8wh9
Q-score: 0.336
Liu Y, Zhang Z, Hu H, Chen W, Zhang F, Wang Q, Wang C, Yan K, Du J
Nat Plants (2024) 10 pp. 374-380 [ Pubmed: 38413824 DOI: doi:10.1038/s41477-024-01640-z ]
- Beryllium trifluoride ion (66 Da, Ligand)
- Histone h4 (11 kDa, Protein from Arabidopsis thaliana)
- Histone h3.1 (15 kDa, Protein from Arabidopsis thaliana)
- Atp-dependent dna helicase ddm1 (86 kDa, Protein from Arabidopsis thaliana)
- Dna (antisense strand) (45 kDa, DNA from synthetic construct)
- Dna (sense strand) (45 kDa, DNA from synthetic construct)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Histone h2b.6 (16 kDa, Protein from Arabidopsis thaliana)
- Histone h2a.6 (13 kDa, Protein from Arabidopsis thaliana)
- Ddm1-nucleosome complex in adp-befx state (290 kDa, Complex from Arabidopsis thaliana)
Structure of DDM1-nucleosome complex in the ADP-BeFx state with DDM1 bound to SHL2 and SHL-2
Resolution: 4.05 Å
EM Method: Single-particle
Fitted PDBs: 8wha
Q-score: 0.306
Liu Y, Zhang Z, Hu H, Chen W, Zhang F, Wang Q, Wang C, Yan K, Du J
Nat Plants (2024) 10 pp. 374-380 [ Pubmed: 38413824 DOI: doi:10.1038/s41477-024-01640-z ]
- Beryllium trifluoride ion (66 Da, Ligand)
- Histone h4 (11 kDa, Protein from Arabidopsis thaliana)
- Histone h3.1 (15 kDa, Protein from Arabidopsis thaliana)
- Atp-dependent dna helicase ddm1 (86 kDa, Protein from Arabidopsis thaliana)
- Dna (antisense strand) (45 kDa, DNA from synthetic construct)
- Dna (sense strand) (45 kDa, DNA from synthetic construct)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Ddm1-nucleosome complex in the adp-befx state with ddm1 bound to shl2 and shl-2 (376 kDa, Complex from Arabidopsis thaliana)
- Histone h2b.6 (16 kDa, Protein from Arabidopsis thaliana)
- Histone h2a.6 (13 kDa, Protein from Arabidopsis thaliana)
Structure of nucleosome core particle of Arabidopsis thaliana
Resolution: 3.17 Å
EM Method: Single-particle
Fitted PDBs: 8whb
Q-score: 0.5
Liu Y, Zhang Z, Hu H, Chen W, Zhang F, Wang Q, Wang C, Yan K, Du J
Nat Plants (2024) 10 pp. 374-380 [ Pubmed: 38413824 DOI: doi:10.1038/s41477-024-01640-z ]
- Dna (sense strand) (45 kDa, DNA from Arabidopsis thaliana)
- Histone h4 (11 kDa, Protein from Arabidopsis thaliana)
- Histone h3.1 (15 kDa, Protein from Arabidopsis thaliana)
- Dna (antisense strand) (45 kDa, DNA from synthetic construct)
- Histone h2b.6 (16 kDa, Protein from Arabidopsis thaliana)
- Histone h2a.6 (13 kDa, Protein from Arabidopsis thaliana)
- Structure of the nucleosome core particle of arabidopsis thaliana (200 kDa, Complex from Arabidopsis thaliana)
Cryo-EM structure of Oryza sativa HKT2;1 at 2.5 angstrom
Resolution: 2.53 Å
EM Method: Single-particle
Fitted PDBs: 8k66
Q-score: 0.66
Wang X, Shen X, Qu Y, Zhang H, Wang C, Yang F, Shen H
Nat Plants (2024) 10 pp. 633-644 [ DOI: doi:10.1038/s41477-024-01665-4 Pubmed: 38570642 ]
- Phosphatidylethanolamine (734 Da, Ligand)
- Water (18 Da, Ligand)
- Phosphatidylinositol (887 Da, Ligand)
- Cholesterol hemisuccinate (486 Da, Ligand)
- 1,2-diacyl-sn-glycero-3-phosphocholine (790 Da, Ligand)
- Cholesterol (386 Da, Ligand)
- Oshkt2;1 dimer (Complex from Oryza sativa subsp. japonica)
- Sodium ion (22 Da, Ligand)
- Cation transporter hkt2;1 (60 kDa, Protein from Oryza sativa subsp. japonica)
Cryo-EM structure of Oryza sativa HKT2;2/1 at 2.3 angstrom
Resolution: 2.33 Å
EM Method: Single-particle
Fitted PDBs: 8k69
Q-score: 0.692
Wang X, Shen X, Qu Y, Zhang H, Wang C, Yang F, Shen H
Nat Plants (2024) 10 pp. 633-644 [ DOI: doi:10.1038/s41477-024-01665-4 Pubmed: 38570642 ]
- Phosphatidylethanolamine (734 Da, Ligand)
- Water (18 Da, Ligand)
- Phosphatidylinositol (887 Da, Ligand)
- Cation transporter hkt2;2 (60 kDa, Protein from Oryza sativa subsp. indica)
- Cholesterol hemisuccinate (486 Da, Ligand)
- 1,2-diacyl-sn-glycero-3-phosphocholine (790 Da, Ligand)
- Cholesterol (386 Da, Ligand)
- Sodium ion (22 Da, Ligand)
- Oshkt2;2/1 dimer (Complex from Oryza sativa subsp. indica)
Apo state of Arabidopsis AZG1 at pH 7.4
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 8irl
Q-score: 0.604
Xu L, Jia W, Tao X, Ye F, Zhang Y, Ding ZJ, Zheng SJ, Qiao S, Su N, Zhang Y, Wu S, Guo J
Nat Plants (2024) 10 pp. 180-191 [ Pubmed: 38172575 DOI: doi:10.1038/s41477-023-01590-y ]
- Azg1 dimer (131 kDa, Complex from Arabidopsis thaliana)
- Adenine/guanine permease azg1 (65 kDa, Protein from Arabidopsis thaliana)
Endogenous substrate adenine bound state of Arabidopsis AZG1 at pH 5.5
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8irm
Q-score: 0.601
Xu L, Jia W, Tao X, Ye F, Zhang Y, Ding ZJ, Zheng SJ, Qiao S, Su N, Zhang Y, Wu S, Guo J
Nat Plants (2024) 10 pp. 180-191 [ Pubmed: 38172575 DOI: doi:10.1038/s41477-023-01590-y ]
- Adenine/guanine permease azg1 (65 kDa, Protein from Arabidopsis thaliana)
- Adenine (135 Da, Ligand)
- Complex of azg1 and adenine (131 kDa, Complex from Arabidopsis thaliana)
- Water (18 Da, Ligand)
6-BAP bound state of Arabidopsis AZG1
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 8irn
Q-score: 0.595
Xu L, Jia W, Tao X, Ye F, Zhang Y, Ding ZJ, Zheng SJ, Qiao S, Su N, Zhang Y, Wu S, Guo J
Nat Plants (2024) 10 pp. 180-191 [ Pubmed: 38172575 DOI: doi:10.1038/s41477-023-01590-y ]
- Water (18 Da, Ligand)
- Adenine/guanine permease azg1 (65 kDa, Protein from Arabidopsis thaliana)
- Complex of azg1 and 6-bap (131 kDa, Complex from Arabidopsis thaliana)
- N-benzyl-9h-purin-6-amine (225 Da, Ligand)