Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
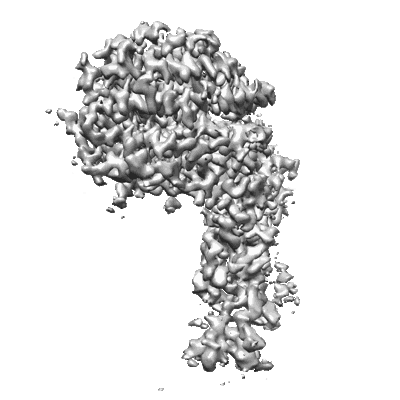
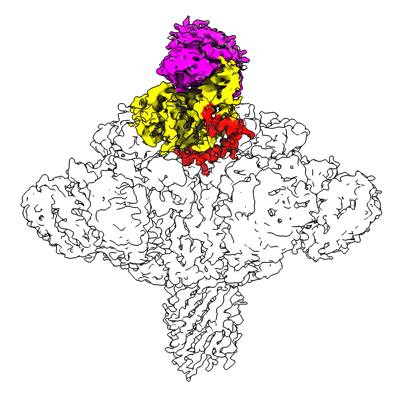
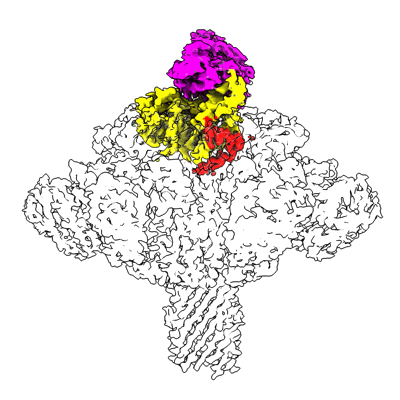
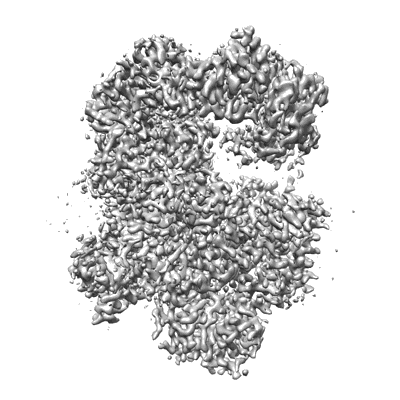
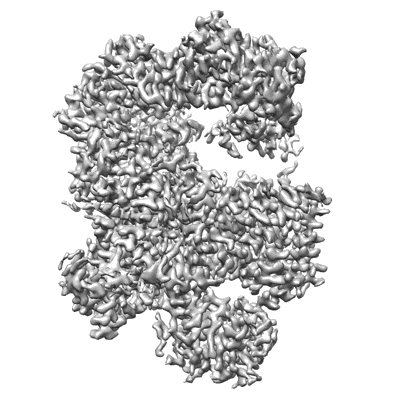
RBD of SARS-CoV2 spike protein with ACE2 decoy
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8jje
Q-score: 0.488
Urano E, Itoh Y, Suzuki T, Sasaki T, Kishikawa JI, Akamatsu K, Higuchi Y, Sakai Y, Okamura T, Mitoma S, Sugihara F, Takada A, Kimura M, Nakao S, Hirose M, Sasaki T, Koketsu R, Tsuji S, Yanagida S, Shioda T, Hara E, Matoba S, Matsuura Y, Kanda Y, Arase H, Okada M, Takagi J, Kato T, Hoshino A, Yasutomi Y, Saito A, Okamoto T
Sci Transl Med (2023) 15 pp. eadi2623-eadi2623 [ DOI: doi:10.1126/scitranslmed.adi2623 Pubmed: 37647387 ]
- Sulfate ion (96 Da, Ligand)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Angiotensin-converting enzyme 2 (73 kDa, Protein from Homo sapiens)
- Water (18 Da, Ligand)
- Spike glycoprotein (138 kDa, Protein from synthetic construct)
- Zinc ion (65 Da, Ligand)
- Ace2 (Complex from Homo sapiens)
- Rbd of sars-cov2 spike protein with ace2 decoy. (460 kDa, Complex)
- Sars-cov2 spike protein (Complex from Severe acute respiratory syndrome coronavirus 2)
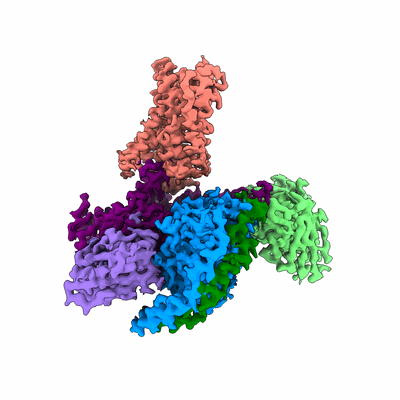
Cryo-EM structure of the histamine-bound histamine H4 receptor and Gq complex
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 7yfc
Q-score: 0.539
Im D, Kishikawa JI, Shiimura Y, Hisano H, Ito A, Fujita-Fujiharu Y, Sugita Y, Noda T, Kato T, Asada H, Iwata S
Nat Commun (2023) 14 pp. 6538-6538 [ DOI: doi:10.1038/s41467-023-42260-z Pubmed: 37863901 ]
- Histamine h4 receptor (62 kDa, Protein from Homo sapiens)
- Guanine nucleotide-binding protein g(i)/g(s)/g(t) subunit beta-1 (42 kDa, Protein from Homo sapiens)
- Histamine h4 receptor (Complex)
- Guanine nucleotide-binding protein g(i)/g(s)/g(t) subunit beta-1 (Complex from Mus musculus)
- Scfv16 (Complex from Homo sapiens)
- Engineered g-alpha-q (41 kDa, Protein from Homo sapiens)
- Nb35 (14 kDa, Protein from Lama glama)
- Histamine (111 Da, Ligand)
- Cholesterol (386 Da, Ligand)
- Scfv16 (27 kDa, Protein from Mus musculus)
- Engineered g-alpha-q (Complex from Homo sapiens)
- Histamine-bound histamine h4 receptor in complex with gq (190 kDa, Complex from Homo sapiens)
- Nb35 (Complex from Homo sapiens)
- Guanine nucleotide-binding protein g(i)/g(s)/g(o) subunit gamma-2 (7 kDa, Protein from Homo sapiens)
- Guanine nucleotide-binding protein g(i)/g(s)/g(o) subunit gamma-2 (Complex from Lama glama)
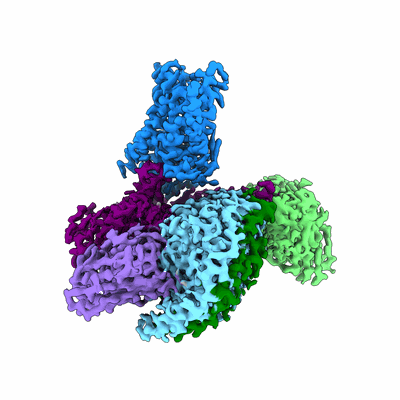
Cryo-EM structure of the imetit-bound histamine H4 receptor and Gq complex
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7yfd
Q-score: 0.552
Im D, Kishikawa JI, Shiimura Y, Hisano H, Ito A, Fujita-Fujiharu Y, Sugita Y, Noda T, Kato T, Asada H, Iwata S
Nat Commun (2023) 14 pp. 6538-6538 [ DOI: doi:10.1038/s41467-023-42260-z Pubmed: 37863901 ]
- Imetit bound histamine h4 receptor in complex with gq (190 kDa, Complex from Homo sapiens)
- Guanine nucleotide-binding protein g(i)/g(s)/g(t) subunit beta-1 (42 kDa, Protein from Homo sapiens)
- Nb35 (Complex)
- Scfv16 (Complex from Homo sapiens)
- Guanine nucleotide-binding protein g(i)/g(s)/g(t) subunit beta-1 (Complex from Mus musculus)
- Engineered g-alpha-q (41 kDa, Protein from Homo sapiens)
- Nb35 (14 kDa, Protein from Lama glama)
- Cholesterol (386 Da, Ligand)
- Scfv16 (27 kDa, Protein from Mus musculus)
- Engineered g-alpha-q (Complex from Homo sapiens)
- 2-(1~{h}-imidazol-5-yl)ethyl carbamimidothioate (170 Da, Ligand)
- Histamine h4 receptor (Complex from Homo sapiens)
- Histamine h4 receptor (62 kDa, Protein from Homo sapiens)
- Guanine nucleotide-binding protein g(i)/g(s)/g(o) subunit gamma-2 (7 kDa, Protein from Homo sapiens)
- Guanine nucleotide-binding protein g(i)/g(s)/g(o) subunit gamma-2 (Complex from Lama glama)
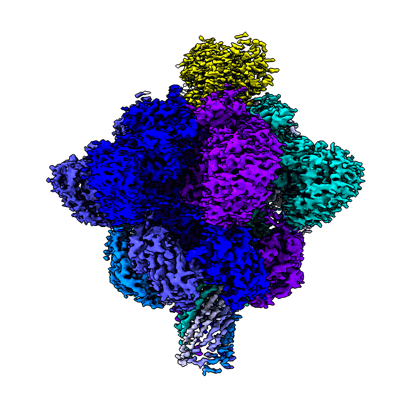
Resolution: 2.56 Å
EM Method: Single-particle
Fitted PDBs: 7vnj
Q-score: 0.536
Kawamoto A, Yamada T, Yoshida T, Sato Y, Kato T, Tsuge H
Nat Commun (2022) 13 pp. 6119-6119 [ DOI: doi:10.1038/s41467-022-33888-4 Pubmed: 36253419 ]
- Adp-ribosyltransferase enzymatic component (49 kDa, Protein from Clostridioides difficile)
- Cdta (Complex)
- Adp-ribosylating binary toxin binding subunit cdtb (75 kDa, Protein from Clostridioides difficile)
- Calcium ion (40 Da, Ligand)
- Cdtb (Complex from Clostridioides difficile)
- Cdta-bound cdtb-pore (long) (Complex from Clostridioides difficile)
Resolution: 2.64 Å
EM Method: Single-particle
Fitted PDBs: 7vnn
Q-score: 0.529
Kawamoto A, Yamada T, Yoshida T, Sato Y, Kato T, Tsuge H
Nat Commun (2022) 13 pp. 6119-6119 [ DOI: doi:10.1038/s41467-022-33888-4 Pubmed: 36253419 ]
- Adp-ribosylating binary toxin binding subunit cdtb (75 kDa, Protein from Clostridioides difficile)
- Cdta (Complex)
- Cdtb-pore (Complex from Clostridioides difficile)
- Calcium ion (40 Da, Ligand)
- Cdta (49 kDa, Protein from Clostridioides difficile)
- Cdta-bound cdtb-pore (long) (Complex from Clostridioides difficile)
Complex structure of Clostridioides difficile binary toxin folded CDTa-bound CDTb-pore (short).
Resolution: 3.18 Å
EM Method: Single-particle
Fitted PDBs: 7yvq
Q-score: 0.494
Kawamoto A, Yamada T, Yoshida T, Sato Y, Kato T, Tsuge H
Nat Commun (2022) 13 pp. 6119-6119 [ DOI: doi:10.1038/s41467-022-33888-4 Pubmed: 36253419 ]
- Adp-ribosylating binary toxin enzymatic subunit cdta (49 kDa, Protein from Clostridioides difficile)
- Clostridioides difficile transferase complex (Complex from Clostridioides difficile)
- Adp-ribosylating binary toxin binding subunit cdtb (75 kDa, Protein from Clostridioides difficile)
- Calcium ion (40 Da, Ligand)
Complex structure of Clostridioides difficile binary toxin unfolded CDTa-bound CDTb-pore (short).
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 7yvs
Q-score: 0.549
Kawamoto A, Yamada T, Yoshida T, Sato Y, Kato T, Tsuge H
Nat Commun (2022) 13 pp. 6119-6119 [ DOI: doi:10.1038/s41467-022-33888-4 Pubmed: 36253419 ]
- Adp-ribosylating binary toxin enzymatic subunit cdta (49 kDa, Protein from Clostridioides difficile)
- Clostridioides difficile transferase complex (Complex from Clostridioides difficile)
- Adp-ribosylating binary toxin binding subunit cdtb (75 kDa, Protein from Clostridioides difficile)
- Calcium ion (40 Da, Ligand)
Cryo-EM structure of Na+-pumping NADH-ubiquinone oxidoreductase from Vibrio cholerae, state 1
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7xk3
Q-score: 0.567
Kishikawa JI, Ishikawa M, Masuya T, Murai M, Kitazumi Y, Butler NL, Kato T, Barquera B, Miyoshi H
Nat Commun (2022) 13 pp. 4082-4082 [ DOI: doi:10.1038/s41467-022-31718-1 Pubmed: 35882843 ]
- Flavin mononucleotide (456 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit e (21 kDa, Protein from Vibrio cholerae O395)
- Na+-pumping nadh-ubiquinone oxidoreductase from vibrio cholerae, state 1 (220 kDa, Complex from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit d (22 kDa, Protein from Vibrio cholerae O395)
- Dodecyl-beta-d-maltoside (510 Da, Ligand)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Calcium ion (40 Da, Ligand)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit f (45 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit c (27 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit b (45 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit a (48 kDa, Protein from Vibrio cholerae O395)
- Riboflavin (376 Da, Ligand)
Cryo-EM structure of Na+-pumping NADH-ubiquinone oxidoreductase from Vibrio cholerae, state 2
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7xk4
Q-score: 0.562
Kishikawa JI, Ishikawa M, Masuya T, Murai M, Kitazumi Y, Butler NL, Kato T, Barquera B, Miyoshi H
Nat Commun (2022) 13 pp. 4082-4082 [ DOI: doi:10.1038/s41467-022-31718-1 Pubmed: 35882843 ]
- Flavin mononucleotide (456 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit f (45 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit e (21 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit d (22 kDa, Protein from Vibrio cholerae O395)
- Dodecyl-beta-d-maltoside (510 Da, Ligand)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Calcium ion (40 Da, Ligand)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Na+-pumping nadh-ubiquinone oxidoreductase from vibrio cholerae, state 2 (220 kDa, Complex from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit c (27 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit b (45 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit a (48 kDa, Protein from Vibrio cholerae O395)
- Riboflavin (376 Da, Ligand)
Cryo-EM structure of Na+-pumping NADH-ubiquinone oxidoreductase from Vibrio cholerae, state 3
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7xk5
Q-score: 0.546
Kishikawa JI, Ishikawa M, Masuya T, Murai M, Kitazumi Y, Butler NL, Kato T, Barquera B, Miyoshi H
Nat Commun (2022) 13 pp. 4082-4082 [ DOI: doi:10.1038/s41467-022-31718-1 Pubmed: 35882843 ]
- Flavin mononucleotide (456 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit e (21 kDa, Protein from Vibrio cholerae O395)
- Na+-pumping nadh-ubiquinone oxidoreductase from vibrio cholerae, state 3 (220 kDa, Complex from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit d (22 kDa, Protein from Vibrio cholerae O395)
- Dodecyl-beta-d-maltoside (510 Da, Ligand)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine (744 Da, Ligand)
- Calcium ion (40 Da, Ligand)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Na(+)-translocating nadh-quinone reductase subunit f (45 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit c (27 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit b (45 kDa, Protein from Vibrio cholerae O395)
- Na(+)-translocating nadh-quinone reductase subunit a (48 kDa, Protein from Vibrio cholerae O395)
- Riboflavin (376 Da, Ligand)