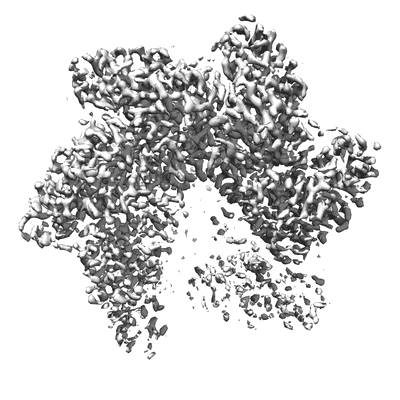Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
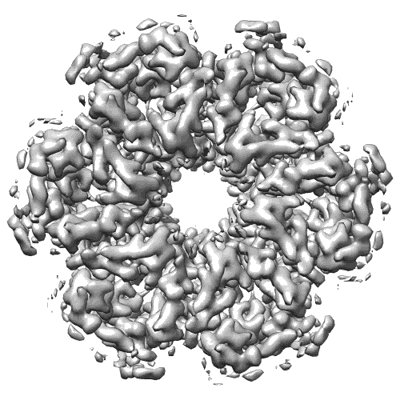
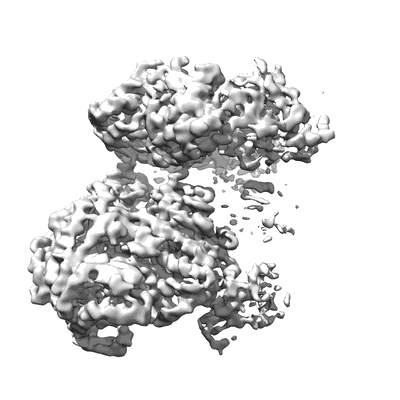
CryoEM structure of Helicobacter pylori urea channel in closed state
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 6nsj
Q-score: 0.509
Cui Y, Zhou K, Strugatsky D, Wen Y, Sachs G, Zhou ZH, Munson K
Sci Adv (2019) 5 pp. eaav8423-eaav8423 [ Pubmed: 30906870 DOI: doi:10.1126/sciadv.aav8423 ]
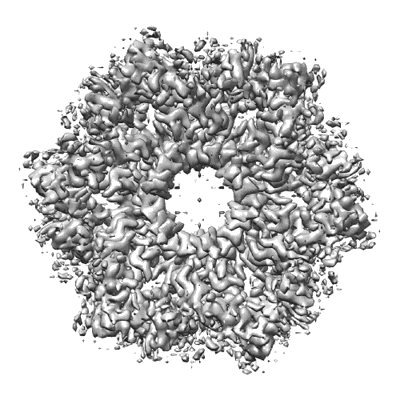
CryoEM structure of Helicobacter pylori urea channel in open state.
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 6nsk
Q-score: 0.591
Cui Y, Zhou K, Strugatsky D, Wen Y, Sachs G, Zhou ZH, Munson K
Sci Adv (2019) 5 pp. eaav8423-eaav8423 [ Pubmed: 30906870 DOI: doi:10.1126/sciadv.aav8423 ]
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 6n7i
Q-score: 0.483
Gao Y, Cui Y, Fox T, Lin S, Wang H, de Val N, Zhou ZH, Yang W
Science (2019) 363 [ Pubmed: 30679383 DOI: doi:10.1126/science.aav7003 ]
- Dna primase/helicase (62 kDa, Protein from Enterobacteria phage T7)
- Thymidine-5'-triphosphate (482 Da, Ligand)
- Water (18 Da, Ligand)
- Bacteriophage t7 gene product 4 (gp4) helicase primase dna complex with five helicase subunits ordered (Complex from Enterobacteria phage T7)
- Dna (25-mer) (7 kDa, DNA from Enterobacteria phage T7)
- Magnesium ion (24 Da, Ligand)
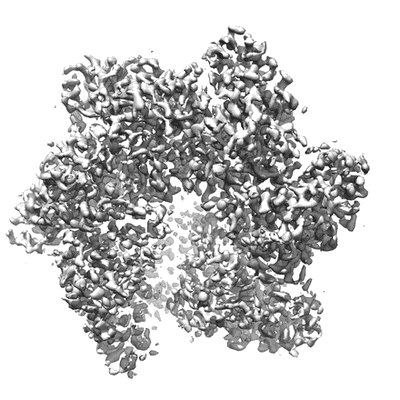
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 6n7n
Q-score: 0.421
Gao Y, Cui Y, Fox T, Lin S, Wang H, de Val N, Zhou ZH, Yang W
Science (2019) 363 [ Pubmed: 30679383 DOI: doi:10.1126/science.aav7003 ]
- Dna primase/helicase (62 kDa, Protein from Enterobacteria phage T7)
- Thymidine-5'-triphosphate (482 Da, Ligand)
- Dna (5'-d(p*tp*tp*tp*tp*tp*tp*tp*tp*tp*tp*tp*tp*tp*tp*t)-3') (4 kDa, DNA from Enterobacteria phage T7)
- Magnesium ion (24 Da, Ligand)
- Bacteriophage t7 gene product 4 (gp4) helicase primase dna complex i (Complex from Enterobacteria phage T7)
Resolution: 4.0 Å
EM Method: Single-particle
Fitted PDBs: 6n9w
Q-score: 0.334
Gao Y, Cui Y, Fox T, Lin S, Wang H, de Val N, Zhou ZH, Yang W
Science (2019) 363 [ Pubmed: 30679383 DOI: doi:10.1126/science.aav7003 ]
- Dna primase/helicase (62 kDa, Protein from Enterobacteria phage T7)
- Dna-directed dna polymerase (79 kDa, Protein from Enterobacteria phage T7)
- Thymidine-5'-triphosphate (482 Da, Ligand)
- Primer (1 kDa, RNA from Enterobacteria phage T7)
- Zinc ion (65 Da, Ligand)
- Template (13 kDa, DNA from Enterobacteria phage T7)
- Magnesium ion (24 Da, Ligand)
- Gp5 dna polymerase binding to a and b subunits of gp4 primase-helicase hexamer complexed with trx, rna/dna hybrid, and incoming dttp (lags2) (Complex from Enterobacteria phage T7)
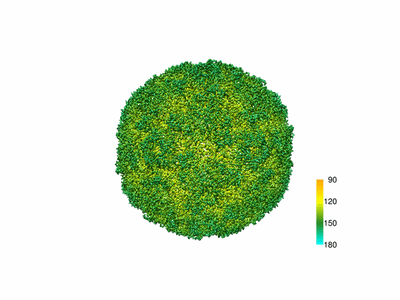
The structure of CVA10 virus mature virion
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 6acu
Q-score: 0.086
Zhu R, Xu L, Zheng Q, Cui Y, Li S, He M, Yin Z, Liu D, Li S, Li Z, Chen Z, Yu H, Que Y, Liu C, Kong Z, Zhang J, Baker TS, Yan X, Zhou ZH, Cheng T, Xia N
Sci Adv (2018) 4 pp. eaat7459-eaat7459 [ Pubmed: 30255146 DOI: doi:10.1126/sciadv.aat7459 ]
- Vp1 (33 kDa, Protein from Coxsackievirus A10)
- Vp3 (26 kDa, Protein from Coxsackievirus A10)
- Sphingosine (299 Da, Ligand)
- Coxsackievirus a10 (Virus from Coxsackievirus A10)
- Vp2 (27 kDa, Protein from Coxsackievirus A10)
- Vp4 (7 kDa, Protein from Coxsackievirus A10)