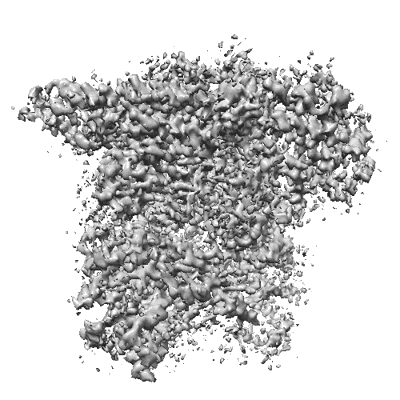Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
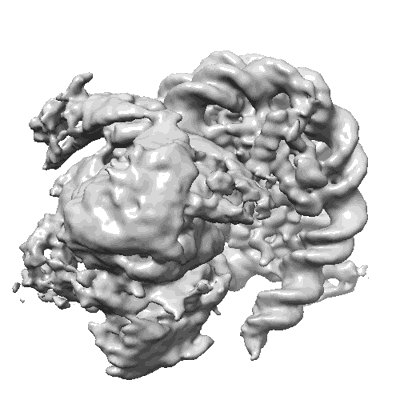
Resolution: 3.72 Å
EM Method: Single-particle
Fitted PDBs: 8iht
Q-score: 0.345
Zhang Y, Xu M, Wang P, Zhou J, Wang G, Han S, Cai G, Wang X
Cell Res (2023) 33 pp. 971-974 [ DOI: doi:10.1038/s41422-023-00884-2 Pubmed: 37845487 ]
- Histone deacetylase rpd3 (48 kDa, Protein from Saccharomyces cerevisiae)
- Histone h2a (13 kDa, Protein from Xenopus laevis)
- Dna (165-mer) (51 kDa, DNA from Xenopus laevis)
- Calcium ion (40 Da, Ligand)
- Histone h4 (11 kDa, Protein from Xenopus laevis)
- Chromatin modification-related protein eaf3 (45 kDa, Protein from Saccharomyces cerevisiae)
- Zinc ion (65 Da, Ligand)
- Histone h2b (13 kDa, Protein from Xenopus laevis)
- Dna (164-mer) (50 kDa, DNA from Xenopus laevis)
- Transcriptional regulatory protein sin3 (175 kDa, Protein from Saccharomyces cerevisiae)
- Eaf3 chd domain bound to the nucleosome (Complex from Xenopus laevis)
- Histone h3 (15 kDa, Protein from Xenopus laevis)
- Rco1 isoform 1 (78 kDa, Protein from Saccharomyces cerevisiae)
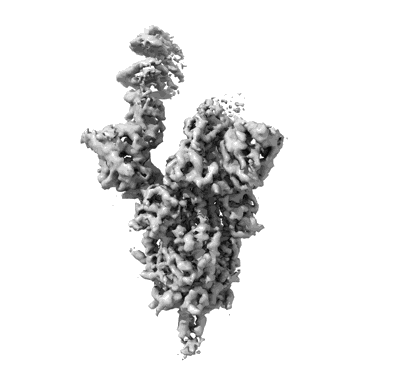
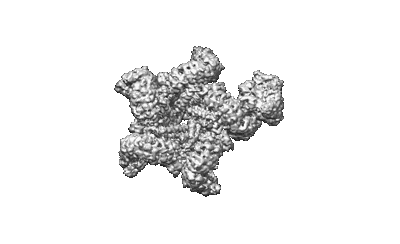
XBB.1.5.10 spike protein in complex with ACE2
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 8wro
Q-score: 0.193
Jian F, Feng L, Yang S, Yu Y, Wang L, Song W, Yisimayi A, Chen X, Xu Y, Wang P, Yu L, Wang J, Liu L, Niu X, Wang J, Xiao T, An R, Wang Y, Gu Q, Shao F, Jin R, Shen Z, Wang Y, Wang X, Cao Y
PLoS Pathog (2023) 19 pp. e1011868-e1011868 [ Pubmed: 38117863 DOI: doi:10.1371/journal.ppat.1011868 ]
- Human ace2 (Complex from Homo sapiens)
- Spike glycoprotein,spike glycoprotein,spike glycoprotein,fusion protein (145 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Spike protein (Complex from Severe acute respiratory syndrome coronavirus 2)
- Processed angiotensin-converting enzyme 2 (68 kDa, Protein from Homo sapiens)
- Xbb.1.5.10 spike protein in complex with ace2 (Complex)
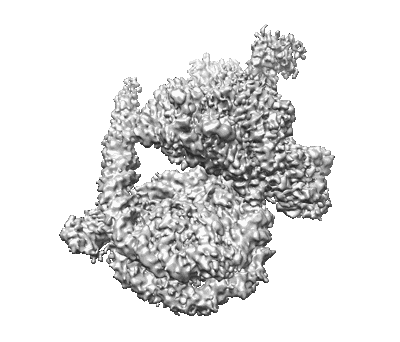
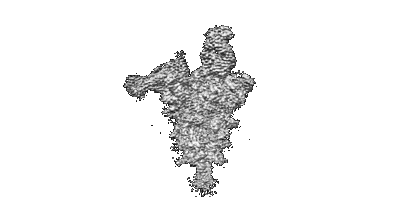
Cryo-EM map of Rpd3S in loose-state Rpd3S-NCP complex
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7yi2
Q-score: 0.483
Guan H, Wang P, Zhang P, Ruan C, Ou Y, Peng B, Zheng X, Lei J, Li B, Yan C, Li H
Nature (2023) 620 pp. 669-675 [ DOI: doi:10.1038/s41586-023-06349-1 Pubmed: 37468628 ]
- Rpd3s complex with h3k36me3 nucleosome (723 MDa, Complex from synthetic construct)
- Chromatin modification-related protein eaf3 (45 kDa, Protein from Saccharomyces cerevisiae S288C)
- Wisdom 601 dna (167-mer) (51 kDa, DNA from synthetic construct)
- Transcriptional regulatory protein sin3 (175 kDa, Protein from Saccharomyces cerevisiae S288C)
- Wisdom 601 dna (167-mer) (51 kDa, DNA from synthetic construct)
- Dna (Complex from Saccharomyces cerevisiae S288C)
- Transcriptional regulatory protein rco1 (78 kDa, Protein from Saccharomyces cerevisiae S288C)
- Histone deacetylase rpd3 (48 kDa, Protein from Saccharomyces cerevisiae S288C)
- Zinc ion (65 Da, Ligand)
- Sin3/rpd3/eaf3/rco1 (Complex)
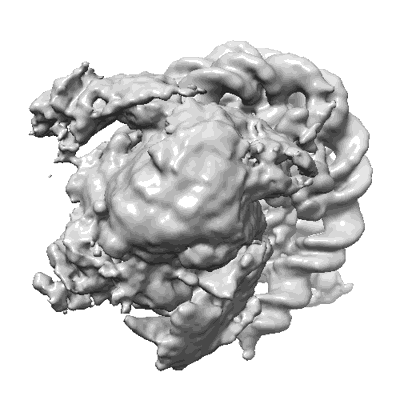
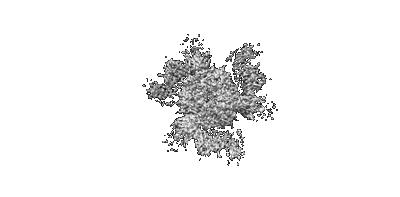
Cryo-EM map of Rpd3S complex bound to H3K36me3 nucleosome in close state
Resolution: 3.96 Å
EM Method: Single-particle
Fitted PDBs: 7yi4
Q-score: 0.23
Guan H, Wang P, Zhang P, Ruan C, Ou Y, Peng B, Zheng X, Lei J, Li B, Yan C, Li H
Nature (2023) 620 pp. 669-675 [ DOI: doi:10.1038/s41586-023-06349-1 Pubmed: 37468628 ]
- Chromatin modification-related protein eaf3 (45 kDa, Protein from Saccharomyces cerevisiae S288C)
- Wisdom 601 dna (167-mer) (51 kDa, DNA from synthetic construct)
- Transcriptional regulatory protein sin3 (175 kDa, Protein from Saccharomyces cerevisiae S288C)
- Histone h2a (13 kDa, Protein from Xenopus laevis)
- Histone h4 (11 kDa, Protein from Xenopus laevis)
- Wisdom 601 dna (167-mer) (51 kDa, DNA from synthetic construct)
- Dna (Complex from Xenopus laevis)
- Sin3/rpd3/eaf3/rco1 (Complex from Xenopus laevis)
- Histone deacetylase rpd3 (48 kDa, Protein from Saccharomyces cerevisiae S288C)
- Histone h3 (Complex)
- Histone h4/h2a/h2b 1.1 (Complex from synthetic construct)
- Transcriptional regulatory protein rco1 (78 kDa, Protein from Saccharomyces cerevisiae S288C)
- Histone h2b 1.1 (13 kDa, Protein from Xenopus laevis)
- Zinc ion (65 Da, Ligand)
- Histone h3 (15 kDa, Protein from Xenopus laevis)
- Rpd3s complex with h3k36me3 nucleosome (723 MDa, Complex from Saccharomyces cerevisiae S288C)
Cryo-EM msp of Rpd3S complex bound to H3K36me3 nucleosome in loose state
Resolution: 3.96 Å
EM Method: Single-particle
Fitted PDBs: 7yi5
Q-score: 0.254
Guan H, Wang P, Zhang P, Ruan C, Ou Y, Peng B, Zheng X, Lei J, Li B, Yan C, Li H
Nature (2023) 620 pp. 669-675 [ DOI: doi:10.1038/s41586-023-06349-1 Pubmed: 37468628 ]
- Chromatin modification-related protein eaf3 (45 kDa, Protein from Saccharomyces cerevisiae S288C)
- Wisdom 601 dna (167-mer) (51 kDa, DNA from synthetic construct)
- Transcriptional regulatory protein sin3 (175 kDa, Protein from Saccharomyces cerevisiae S288C)
- Histone h2a (13 kDa, Protein from Xenopus laevis)
- Histone h4 (11 kDa, Protein from Xenopus laevis)
- Wisdom 601 dna (167-mer) (51 kDa, DNA from synthetic construct)
- Dna (Complex from Xenopus laevis)
- Sin3/rpd3/eaf3/rco1 (Complex from Xenopus laevis)
- Histone deacetylase rpd3 (48 kDa, Protein from Saccharomyces cerevisiae S288C)
- Histone h3 (Complex)
- Histone h4/h2a/h2b 1.1 (Complex from synthetic construct)
- Transcriptional regulatory protein rco1 (78 kDa, Protein from Saccharomyces cerevisiae S288C)
- Histone h2b 1.1 (13 kDa, Protein from Xenopus laevis)
- Zinc ion (65 Da, Ligand)
- Histone h3 (15 kDa, Protein from Xenopus laevis)
- Rpd3s complex with h3k36me3 nucleosome (723 MDa, Complex from Saccharomyces cerevisiae S288C)
Cryo-EM structure of Sr35 resistosome induced by AvrSr35 R381A
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 7xx2
Q-score: 0.418
Zhao YB, Liu MX, Chen TT, Ma X, Li ZK, Zheng Z, Zheng SR, Chen L, Li YZ, Tang LR, Chen Q, Wang P, Ouyang S
Sci Adv (2022) 8 pp. eabq5108-eabq5108 [ DOI: doi:10.1126/sciadv.abq5108 Pubmed: 36083908 ]
- Sr35 (Complex from Triticum monococcum)
- Avrsr35 (66 kDa, Protein from Puccinia graminis f. sp. tritici)
- Sr35 resistosome (Complex)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Cnl9 (105 kDa, Protein from Triticum monococcum)
- Avrsr35 (Complex from Puccinia graminis f.sp.tritici)
SARS-CoV-2 BA.2.75 S Trimer in complex with S309 (state1)
Resolution: 3.72 Å
EM Method: Single-particle
Fitted PDBs: 7yqx
Q-score: 0.353
Cao Y, Song W, Wang L, Liu P, Yue C, Jian F, Yu Y, Yisimayi A, Wang P, Wang Y, Zhu Q, Deng J, Fu W, Yu L, Zhang N, Wang J, Xiao T, An R, Wang J, Liu L, Yang S, Niu X, Gu Q, Shao F, Hao X, Meng B, Gupta RK, Jin R, Wang Y, Xie XS, Wang X
Cell Host Microbe (2022) 30 pp. 1527- [ DOI: doi:10.1016/j.chom.2022.09.018 Pubmed: 36270286 ]
- Ba.2.75 s trimer (Complex from Homo sapiens)
- S309 heavy chain (13 kDa, Protein from Homo sapiens)
- Ba.2.75 s trimer in complex with s309 (Complex from Severe acute respiratory syndrome coronavirus 2)
- S309 light chain (23 kDa, Protein from Homo sapiens)
- S309 (Complex)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
SARS-CoV-2 BA.2.75 S Trimer in complex with S309 (state2)
Resolution: 3.74 Å
EM Method: Single-particle
Fitted PDBs: 7yqy
Q-score: 0.352
Cao Y, Song W, Wang L, Liu P, Yue C, Jian F, Yu Y, Yisimayi A, Wang P, Wang Y, Zhu Q, Deng J, Fu W, Yu L, Zhang N, Wang J, Xiao T, An R, Wang J, Liu L, Yang S, Niu X, Gu Q, Shao F, Hao X, Meng B, Gupta RK, Jin R, Wang Y, Xie XS, Wang X
Cell Host Microbe (2022) 30 pp. 1527- [ DOI: doi:10.1016/j.chom.2022.09.018 Pubmed: 36270286 ]
- Ba.2.75 s trimer in complex with s309 (Complex from Homo sapiens)
- S309 heavy chain (13 kDa, Protein from Homo sapiens)
- S309 light chain (23 kDa, Protein from Homo sapiens)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
SARS-CoV-2 BA.2.75 S Trimer in complex with S309 (state3)
Resolution: 3.84 Å
EM Method: Single-particle
Fitted PDBs: 7yqz
Q-score: 0.382
Cao Y, Song W, Wang L, Liu P, Yue C, Jian F, Yu Y, Yisimayi A, Wang P, Wang Y, Zhu Q, Deng J, Fu W, Yu L, Zhang N, Wang J, Xiao T, An R, Wang J, Liu L, Yang S, Niu X, Gu Q, Shao F, Hao X, Meng B, Gupta RK, Jin R, Wang Y, Xie XS, Wang X
Cell Host Microbe (2022) 30 pp. 1527- [ DOI: doi:10.1016/j.chom.2022.09.018 Pubmed: 36270286 ]
- Ba.2.75 s trimer (Complex from Homo sapiens)
- S309 heavy chain (13 kDa, Protein from Homo sapiens)
- Ba.2.75 s trimer in complex with s309 (Complex from Severe acute respiratory syndrome coronavirus 2)
- S309 light chain (23 kDa, Protein from Homo sapiens)
- S309 (Complex)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
SARS-CoV-2 BA.2.75 S Trimer in complex with XG2v024
Resolution: 3.62 Å
EM Method: Single-particle
Fitted PDBs: 7yr1
Q-score: 0.459
Cao Y, Song W, Wang L, Liu P, Yue C, Jian F, Yu Y, Yisimayi A, Wang P, Wang Y, Zhu Q, Deng J, Fu W, Yu L, Zhang N, Wang J, Xiao T, An R, Wang J, Liu L, Yang S, Niu X, Gu Q, Shao F, Hao X, Meng B, Gupta RK, Jin R, Wang Y, Xie XS, Wang X
Cell Host Microbe (2022) 30 pp. 1527- [ DOI: doi:10.1016/j.chom.2022.09.018 Pubmed: 36270286 ]
- Ba.2.75 s trimer (Complex from Homo sapiens)
- Ba.2.75 s trimer in complex with xg2v024 (Complex from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Xg2v024 light chain (11 kDa, Protein from Homo sapiens)
- Xg2v024 heavy/light chains (Complex)
- Xg2v024 heavy chain (13 kDa, Protein from Homo sapiens)