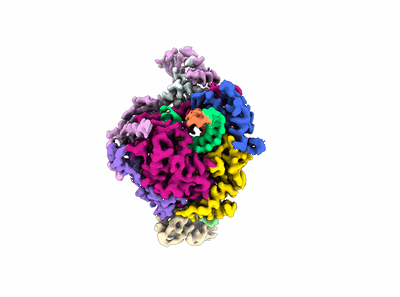Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
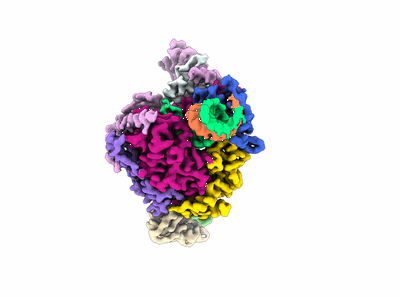
Structure of COVID-19 RNA-dependent RNA polymerase bound to penciclovir.
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 7doi
Q-score: 0.599
Yin W, Luan X, Li Z, Xie Y, Zhou Z, Liu J, Gao M, Wang X, Zhou F, Wang Q, Wang Q, Shen D, Zhang Y, Tian G, Aisa H, Wei D, Jiang Y, Xiao G, Jiang H, Zhang L, Yu X, Shen J, Zhang S, Xu H
To Be Published
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Covid-19 rdrp complex bound to penciclovir (Complex from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*cp*cp*up*ap*up*ap*ap*cp*up*up*ap*ap*up*cp*u)-3') (4 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Pyrophosphate 2- (175 Da, Ligand)
- Rna (5'-r(p*ap*gp*ap*up*up*ap*ap*gp*up*up*ap*u)-3') (3 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- [(2r)-4-(2-azanyl-6-oxidanylidene-3h-purin-9-yl)-2-(hydroxymethyl)butyl] dihydrogen phosphate (333 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (107 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
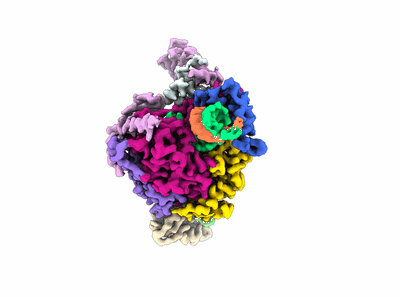
Structure of COVID-19 RNA-dependent RNA polymerase (extended conformation) bound to penciclovir
Resolution: 2.73 Å
EM Method: Single-particle
Fitted PDBs: 7dok
Q-score: 0.554
Yin W, Luan X, Li Z, Xie Y, Zhou Z, Liu J, Gao M, Wang X, Zhou F, Wang Q, Wang Q, Shen D, Zhang Y, Tian G, Aisa H, Wei D, Jiang Y, Xiao G, Jiang H, Zhang L, Yu X, Shen J, Zhang S, Xu H
To Be Published
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Pyrophosphate 2- (175 Da, Ligand)
- Covid-19 rdrp complex (exntended conformation) bound to penciclovir (Complex from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- [(2r)-4-(2-azanyl-6-oxidanylidene-3h-purin-9-yl)-2-(hydroxymethyl)butyl] dihydrogen phosphate (333 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Rna (5'-r(p*gp*cp*up*ap*up*gp*up*gp*ap*gp*ap*up*up*ap*ap*gp*up*up*ap*u)-3') (6 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*cp*cp*cp*up*ap*up*ap*ap*cp*up*up*ap*ap*up*cp*up*cp*ap*cp*ap*up*ap*gp*c)-3') (7 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (107 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
Structure of COVID-19 RNA-dependent RNA polymerase bound to favipiravir
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 7dfg
Q-score: 0.598
Yin W, Luan X, Li Z, Xie Y, Zhou Z, Liu J, Gao M, Wang X, Zhou F, Wang Q, Wang Q, Shen D, Zhang Y, Tian G, Aisa H, Wei D, Jiang Y, Xiao G, Jiang H, Zhang L, Yu X, Shen J, Zhang S, Xu H
To Be Published
- Rna (5'-r(p*cp*cp*cp*up*ap*up*ap*ap*cp*up*up*ap*ap*up*cp*u)-3') (4 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Pyrophosphate 2- (175 Da, Ligand)
- Rna (5'-r(p*ap*gp*ap*up*up*ap*ap*gp*up*up*ap*u)-3') (3 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- 6-fluoro-3-oxo-4-(5-o-phosphono-beta-d-ribofuranosyl)-3,4-dihydropyrazine-2-carboxamide (369 Da, Ligand)
- Covid-19 rdrp complex bound to favipiravir (Complex from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (107 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
Structure of COVID-19 RNA-dependent RNA polymerase bound to ribavirin
Resolution: 2.97 Å
EM Method: Single-particle
Fitted PDBs: 7dfh
Q-score: 0.535
Yin W, Luan X, Li Z, Xie Y, Zhou Z, Liu J, Gao M, Wang X, Zhou F, Wang Q, Wang Q, Shen D, Zhang Y, Tian G, Aisa H, Wei D, Jiang Y, Xiao G, Jiang H, Zhang L, Yu X, Shen J, Zhang S, Xu H
To Be Published
- Rna (5'-r(p*cp*cp*cp*cp*cp*ap*cp*ap*up*ap*gp*c)-3') (3 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*gp*cp*up*ap*up*gp*up*g*(lig))-3') (2 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Pyrophosphate 2- (175 Da, Ligand)
- Covid-19 rdrp complex bound to favipiravir (Complex from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Ribavirin monophosphate (324 Da, Ligand)
- Water (18 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymeras (107 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
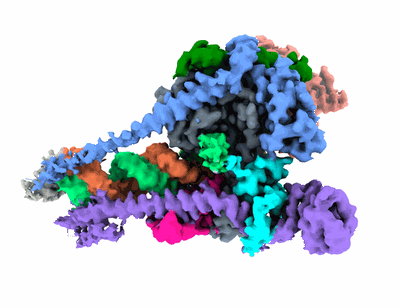
Cryo-EM structure of yeast mitochondrial RNA polymerase transcription initiation complex
Resolution: 3.71 Å
EM Method: Single-particle
Fitted PDBs: 6ymw
Q-score: 0.41
De Wijngaert B, Sultana S, Singh A, Dharia C, Vanbuel H, Shen J, Vasilchuk D, Martinez SE, Kandiah E, Patel SS, Das K
Mol Cell (2021) 81 pp. 268- [ DOI: doi:10.1016/j.molcel.2020.11.016 Pubmed: 33278362 ]
- Dna (Complex from Saccharomyces cerevisiae)
- Chains: t (10 kDa, DNA from Saccharomyces cerevisiae S288C)
- [[(2~{r},3~{s},4~{r},5~{r})-5-[2,4-bis(oxidanylidene)pyrimidin-1-yl]-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-sulfanyl-phosphoryl] phosphono hydrogen phosphate (500 Da, Ligand)
- Mitochondria dna-dependent rna polymerase pre-initiation complex (202 kDa, Complex)
- Dna-directed rna polymerase, mitochondrial (143 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Rna (pppgpg) (Complex from synthetic construct)
- Magnesium ion (24 Da, Ligand)
- Rna (pppgpg) (805 Da, RNA from synthetic construct)
- Mitochondrial transcription factor 1 (41 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Chains: n (10 kDa, DNA from Saccharomyces cerevisiae S288C)
- Mitochondria dna-dependent rna polymerase pre-initiation complex (Complex from Saccharomyces cerevisiae)
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 7bv2
Q-score: 0.607
Yin W, Mao C, Luan X, Shen DD, Shen Q, Su H, Wang X, Zhou F, Zhao W, Gao M, Chang S, Xie YC, Tian G, Jiang HW, Tao SC, Shen J, Jiang Y, Jiang H, Xu Y, Zhang S, Zhang Y, Xu HE
Science (2020) 368 pp. 1499-1504 [ Pubmed: 32358203 DOI: doi:10.1126/science.abc1560 ]
- [(2~{r},3~{s},4~{r},5~{r})-5-(4-azanylpyrrolo[2,1-f][1,2,4]triazin-7-yl)-5-cyano-3,4-bis(oxidanyl)oxolan-2-yl]methyl dihydrogen phosphate (371 Da, Ligand)
- Rna-directed rna polymerase (109 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- The nsp12-nsp7-nsp8 complex bound to the template-primer rna and triphosphate form of remdesivir(rtp) (Complex from SARS bat coronavirus)
- Non-structural protein 8 (23 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Pyrophosphate 2- (175 Da, Ligand)
- Water (18 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Templete (9 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Primer (6 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Non-structural protein 7 (10 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)