Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 3.37 Å
EM Method: Single-particle
Fitted PDBs: 8gwg
Q-score: 0.007
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ Pubmed: 36335936 DOI: doi:10.1016/j.cell.2022.09.037 ]
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Rna (Complex)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_smp-nsp9_gmppnp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Protein from sars-cov-2 (Complex)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Template (10 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 3.18 Å
EM Method: Single-particle
Fitted PDBs: 8gwi
Q-score: 0.185
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ Pubmed: 36335936 DOI: doi:10.1016/j.cell.2022.09.037 ]
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_smp-nsp9_gtp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (27-mer) (10 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Water (18 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 3.38 Å
EM Method: Single-particle
Fitted PDBs: 8gwn
Q-score: 0.407
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ Pubmed: 36335936 DOI: doi:10.1016/j.cell.2022.09.037 ]
- Template (10 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_atmp-nsp9_gmppnp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
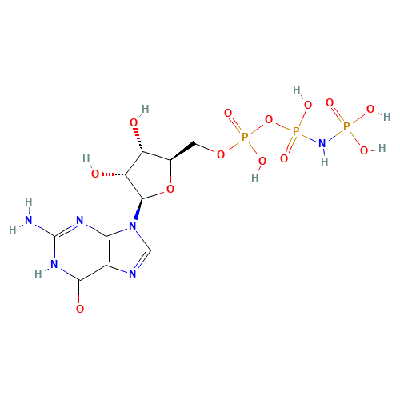
- Phosphoaminophosphonic acid-guanylate ester (522 Da, Ligand)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 3.8 Å
EM Method: Single-particle
Fitted PDBs: 8gwo
Q-score: 0.362
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ Pubmed: 36335936 DOI: doi:10.1016/j.cell.2022.09.037 ]
- Rna (25-mer) (8 kDa, RNA from synthetic construct)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_ump-nsp9_gmppnp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Replicase polyprotein (Complex)
- Rna (Complex)
- Template (10 kDa, RNA from synthetic construct)
- Uridine-5'-monophosphate (324 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Phosphoaminophosphonic acid-guanylate ester (522 Da, Ligand)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)