Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
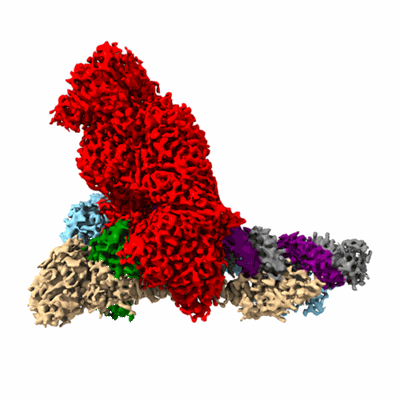
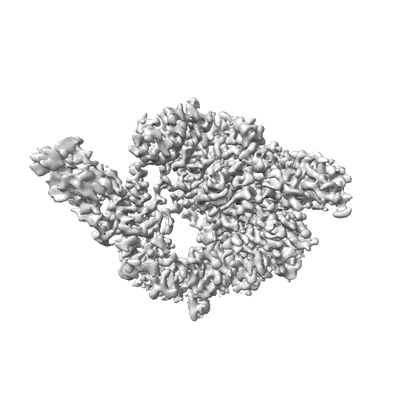
SARS-CoV-2 RdRP catalytic complex with T33-1 RNA
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 7dte
Q-score: 0.528
Wu J, Wang H, Liu Q, Li R, Gao Y, Fang X, Zhong Y, Wang M, Wang Q, Rao Z, Gong P
Cell Rep (2021) 37 pp. 109882-109882 [ Pubmed: 34653416 DOI: doi:10.1016/j.celrep.2021.109882 ]
- Rna (Complex)
- Rna (33-mer) (10 kDa, RNA from Foot-and-mouth disease virus)
- Sars-cov-2 rna-dependent rna polymerase catalytic complex (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (57-mer) (18 kDa, RNA from Foot-and-mouth disease virus)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 rna-dependent rna polymerase (Complex)
- Rna-directed rna polymerase (108 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
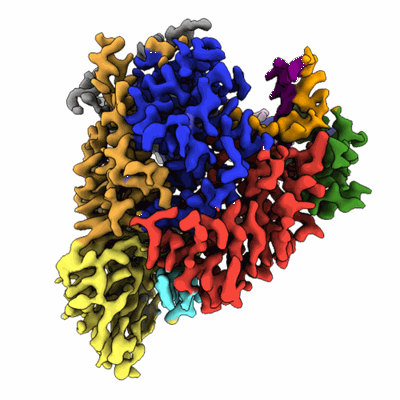
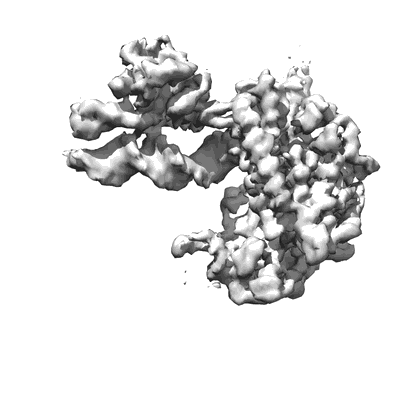
COVID-19 RNA-dependent RNA polymerase post-translocated catalytic complex
Resolution: 3.26 Å
EM Method: Single-particle
Fitted PDBs: 7bzf
Q-score: 0.47
Wang Q, Wu J, Wang H, Gao Y, Liu Q, Mu A, Ji W, Yan L, Zhu Y, Zhu C, Fang X, Yang X, Huang Y, Gao H, Liu F, Ge J, Sun Q, Yang X, Xu W, Liu Z, Yang H, Lou Z, Jiang B, Guddat LW, Gong P, Rao Z
Cell (2020) 182 pp. 417-428.e13 [ Pubmed: 32526208 DOI: doi:10.1016/j.cell.2020.05.034 ]
- Structure of covid-19 rna-dependent rna polymerase in complex with rna (155 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (108 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Rna (31-mer) (10 kDa, RNA from Foot-and-mouth disease virus)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(*up*gp*up*up*cp*gp*ap*cp*gp*ap*cp*ap*cp*a)-3') (4 kDa, RNA from Foot-and-mouth disease virus)
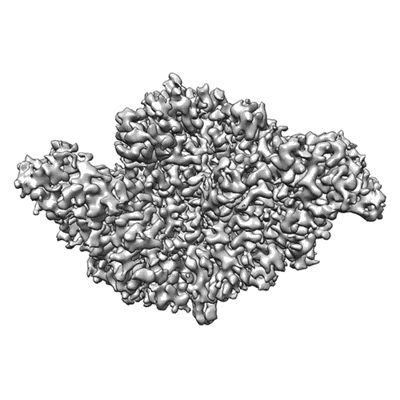
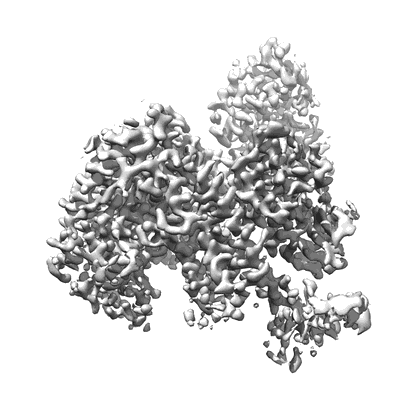
COVID-19 RNA-dependent RNA polymerase pre-translocated catalytic complex
Resolution: 2.93 Å
EM Method: Single-particle
Fitted PDBs: 7c2k
Q-score: 0.581
Wang Q, Wu J, Wang H, Gao Y, Liu Q, Mu A, Ji W, Yan L, Zhu Y, Zhu C, Fang X, Yang X, Huang Y, Gao H, Liu F, Ge J, Sun Q, Yang X, Xu W, Liu Z, Yang H, Lou Z, Jiang B, Guddat LW, Gong P, Rao Z
Cell (2020) 182 pp. 417-428.e13 [ Pubmed: 32526208 DOI: doi:10.1016/j.cell.2020.05.034 ]
- Rna (5'-r(*up*gp*up*up*cp*gp*ap*cp*gp*ap*cp*ap*cp*ap*gp*g*(f86)p*g)-3') (5 kDa, RNA from Foot-and-mouth disease virus)
- Covid-19 rna-dependent rna polymerase pre-translocated catalytic complex (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (29-mer) (9 kDa, RNA from Foot-and-mouth disease virus)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna-directed rna polymerase (108 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
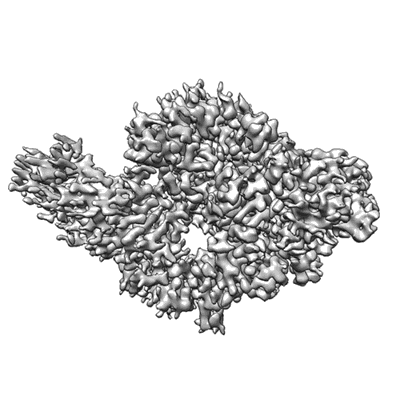
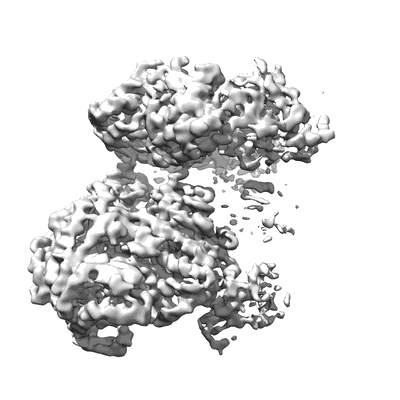
SARS-Cov-2 RNA-dependent RNA polymerase in complex with cofactors in reduced conditions
Resolution: 2.95 Å
EM Method: Single-particle
Fitted PDBs: 7btf
Q-score: 0.569
Gao Y, Yan L, Huang Y, Liu F, Zhao Y, Cao L, Wang T, Sun Q, Ming Z, Zhang L, Ge J, Zheng L, Zhang Y, Wang H, Zhu Y, Zhu C, Hu T, Hua T, Zhang B, Yang X, Li J, Yang H, Liu Z, Xu W, Guddat LW, Wang Q, Lou Z, Rao Z
Science (2020) 368 pp. 779-782 [ Pubmed: 32277040 DOI: doi:10.1126/science.abb7498 ]
- Rna-directed rna polymerase (108 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Structure of sars-cov-2 rna-dependent rna polymerase in complex with cofactors in reduced condition (155 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
SARS-Cov-2 RNA-dependent RNA polymerase in complex with cofactors
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 6m71
Q-score: 0.519
Gao Y, Yan L, Huang Y, Liu F, Zhao Y, Cao L, Wang T, Sun Q, Ming Z, Zhang L, Ge J, Zheng L, Zhang Y, Wang H, Zhu Y, Zhu C, Hu T, Hua T, Zhang B, Yang X, Li J, Yang H, Liu Z, Xu W, Guddat LW, Wang Q, Lou Z, Rao Z
Science (2020) 368 pp. 779-782 [ Pubmed: 32277040 DOI: doi:10.1126/science.abb7498 ]
- Rna-directed rna polymerase (108 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 rna-dependent rna polymerase in complex with cofactors (155 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
Resolution: 4.5 Å
EM Method: Single-particle
Fitted PDBs: 6n7w
Q-score: 0.324
Gao Y, Cui Y, Fox T, Lin S, Wang H, de Val N, Zhou ZH, Yang W
Science (2019) 363 [ Pubmed: 30679383 DOI: doi:10.1126/science.aav7003 ]
- Thymidine-5'-triphosphate (482 Da, Ligand)
- Dna (77-mer) (23 kDa, DNA from Enterobacteria phage T7)
- Trxa (11 kDa, Protein from Escherichia coli)
- Dna-directed dna polymerase (79 kDa, Protein from Enterobacteria phage T7)
- Dna (25-mer) (7 kDa, DNA from Enterobacteria phage T7)
- Magnesium ion (24 Da, Ligand)
- D5ae7a mutant gp5 dna polymerase complexed with trx, fork dna, and incoming dttp (from gp4-gp5 leading-strand complexes) (Complex from Enterobacteria phage T7)
Resolution: 3.7 Å
EM Method: Single-particle
Fitted PDBs: 6n9u
Q-score: 0.477
Gao Y, Cui Y, Fox T, Lin S, Wang H, de Val N, Zhou ZH, Yang W
Science (2019) 363 [ Pubmed: 30679383 DOI: doi:10.1126/science.aav7003 ]
- Dna primase/helicase (62 kDa, Protein from Enterobacteria phage T7)
- Dna-directed dna polymerase (79 kDa, Protein from Enterobacteria phage T7)
- Thymidine-5'-triphosphate (482 Da, Ligand)
- Rna (5'-r(*ap*cp*cp*ap*g)-d(p*(doc))-3') (1 kDa, RNA from Enterobacteria phage T7)
- Dna (44-mer) (13 kDa, DNA from Enterobacteria phage T7)
- Zinc ion (65 Da, Ligand)
- Gp5 dna polymerase and two gp4 primase subunits complexed with trx, rna/dna hybrid, and incoming dttp (from gp4-gp5 lagging-strand complex lags1) (Complex from Enterobacteria phage T7)
- Magnesium ion (24 Da, Ligand)
Resolution: 4.0 Å
EM Method: Single-particle
Fitted PDBs: 6n9v
Q-score: 0.408
Gao Y, Cui Y, Fox T, Lin S, Wang H, de Val N, Zhou ZH, Yang W
Science (2019) 363 [ Pubmed: 30679383 DOI: doi:10.1126/science.aav7003 ]
- Dna primase/helicase (62 kDa, Protein from Enterobacteria phage T7)
- Thymidine-5'-triphosphate (482 Da, Ligand)
- Primer (1 kDa, RNA from Enterobacteria phage T7)
- Zinc ion (65 Da, Ligand)
- Gp5 dna polymerase and two gp4 primase subunits complexed with trx, rna/dna hybrid, and incoming dttp (from gp4-gp5 lagging-strand complex lags1) (Complex from Enterobacteria phage T7)
- Template (13 kDa, DNA from Enterobacteria phage T7)
- Magnesium ion (24 Da, Ligand)
- Dna-directed dna polymerase (79 kDa, Protein from Enterobacteria phage T7)
Resolution: 4.0 Å
EM Method: Single-particle
Fitted PDBs: 6n9w
Q-score: 0.334
Gao Y, Cui Y, Fox T, Lin S, Wang H, de Val N, Zhou ZH, Yang W
Science (2019) 363 [ Pubmed: 30679383 DOI: doi:10.1126/science.aav7003 ]
- Dna primase/helicase (62 kDa, Protein from Enterobacteria phage T7)
- Dna-directed dna polymerase (79 kDa, Protein from Enterobacteria phage T7)
- Thymidine-5'-triphosphate (482 Da, Ligand)
- Primer (1 kDa, RNA from Enterobacteria phage T7)
- Zinc ion (65 Da, Ligand)
- Template (13 kDa, DNA from Enterobacteria phage T7)
- Magnesium ion (24 Da, Ligand)
- Gp5 dna polymerase binding to a and b subunits of gp4 primase-helicase hexamer complexed with trx, rna/dna hybrid, and incoming dttp (lags2) (Complex from Enterobacteria phage T7)
Resolution: 4.1 Å
EM Method: Single-particle
Fitted PDBs: 6n9x
Q-score: 0.371
Gao Y, Cui Y, Fox T, Lin S, Wang H, de Val N, Zhou ZH, Yang W
Science (2019) 363 [ Pubmed: 30679383 DOI: doi:10.1126/science.aav7003 ]
- Dna primase/helicase (62 kDa, Protein from Enterobacteria phage T7)
- Thymidine-5'-triphosphate (482 Da, Ligand)
- Primer (1 kDa, RNA from Enterobacteria phage T7)
- Gp5 dna polymerase binding to b and c subunits of gp4 primase-helicase hexamer complexed with trx, rna/dna hybrid, and incoming dttp (lags3) (Complex from Enterobacteria phage T7)
- Zinc ion (65 Da, Ligand)
- Template (13 kDa, DNA from Enterobacteria phage T7)
- Magnesium ion (24 Da, Ligand)
- Dna-directed dna polymerase (79 kDa, Protein from Enterobacteria phage T7)