Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
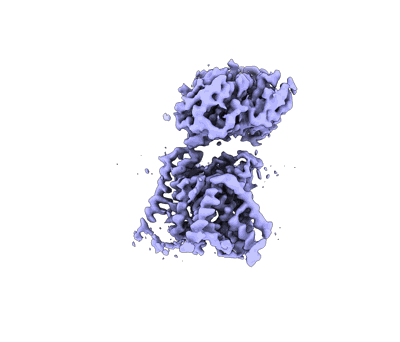
F8-A22-E4 complex of MPXV in complex with DNA and Ara-CTP
Resolution: 3.06 Å
EM Method: Single-particle
Fitted PDBs: 8k8s
Q-score: 0.402
Structure (2024) [ DOI: doi:10.1016/j.str.2024.03.004 Pubmed: 38579705 ]
- Magnesium ion (24 Da, Ligand)
- Dna (5'-d(*ap*tp*cp*cp*tp*cp*cp*cp*cp*tp*ap*c)-3') (3 kDa, DNA from DNA molecule)
- F8-a22-e4 complex of mpxv in complex with dna and ara-ctp (Complex from Monkeypox virus)
- Uracil-dna glycosylase e4 (25 kDa, Protein from Monkeypox virus)
- Water (18 Da, Ligand)
- Dna polymerase f8 (117 kDa, Protein from Monkeypox virus)
- Dna polymerase processivity factor component a20 (49 kDa, Protein from Monkeypox virus)
- Dna (5'-d(p*tp*ap*gp*gp*tp*ap*gp*gp*gp*gp*ap*gp*gp*ap*t)-3') (5 kDa, DNA from DNA molecule)
- 4-amino-1-{5-o-[(s)-hydroxy{[(r)-hydroxy(phosphonooxy)phosphoryl]oxy}phosphoryl]-beta-d-arabinofuranosyl}pyrimidin-2(1h)-one (483 Da, Ligand)
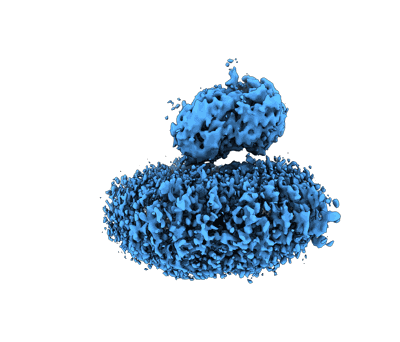
F8-A22-E4 complex of MPXV in complex with DNA and dCTP
Resolution: 3.05 Å
EM Method: Single-particle
Fitted PDBs: 8k8u
Q-score: 0.347
Structure (2024) [ DOI: doi:10.1016/j.str.2024.03.004 Pubmed: 38579705 ]
- Magnesium ion (24 Da, Ligand)
- Dna (5'-d(*ap*tp*cp*cp*tp*cp*cp*cp*cp*tp*ap*c)-3') (3 kDa, DNA from DNA molecule)
- Cytidine-5'-triphosphate (483 Da, Ligand)
- F8-a22-e4 complex of mpxv in complex with dna and ara-ctp (Complex from Monkeypox virus)
- Uracil-dna glycosylase e4 (25 kDa, Protein from Monkeypox virus)
- Water (18 Da, Ligand)
- Dna polymerase f8 (117 kDa, Protein from Monkeypox virus)
- Dna polymerase processivity factor component a20 (49 kDa, Protein from Monkeypox virus)
- Dna (5'-d(p*ap*ap*gp*gp*tp*ap*gp*gp*gp*gp*ap*gp*gp*ap*t)-3') (5 kDa, DNA from DNA molecule)
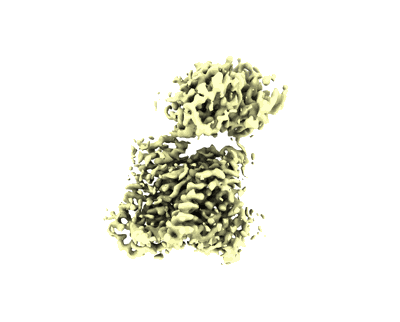
the local map of DNA and Ara-CTP binding site
Structure (2024) [ DOI: doi:10.1016/j.str.2024.03.004 Pubmed: 38579705 ]
the local map of DNA and dCTP binding site
Structure (2024) [ DOI: doi:10.1016/j.str.2024.03.004 Pubmed: 38579705 ]
Cryo EM map of SLC7A10 in the apo state
Resolution: 3.42 Å
EM Method: Single-particle
Fitted PDBs: 8wns
Q-score: 0.406
Li Y, Guo Y, Broer A, Dai L, Yan R
Nat Commun (2024) 15 pp. 3036-3036 [ DOI: doi:10.1038/s41467-024-47468-1 Pubmed: 38589439 ]
- 4f2hc-asc-1 complex (Complex from Homo sapiens)
- Asc-type amino acid transporter 1 (56 kDa, Protein from Homo sapiens)
- Amino acid transporter heavy chain slc3a2 (68 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
Cryo EM map of SLC7A10 with L-Alanine substrate
Resolution: 3.42 Å
EM Method: Single-particle
Fitted PDBs: 8wnt
Q-score: 0.425
Li Y, Guo Y, Broer A, Dai L, Yan R
Nat Commun (2024) 15 pp. 3036-3036 [ DOI: doi:10.1038/s41467-024-47468-1 Pubmed: 38589439 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- 4f2hc-asc-1 complex (Complex from Homo sapiens)
- Alanine (89 Da, Ligand)
- Asc-type amino acid transporter 1 (56 kDa, Protein from Homo sapiens)
- Amino acid transporter heavy chain slc3a2 (68 kDa, Protein from Homo sapiens)
Cryo EM map of SLC7A10-SLC3A2 complex in the D-serine bound state
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 8wny
Q-score: 0.456
Li Y, Guo Y, Broer A, Dai L, Yan R
Nat Commun (2024) 15 pp. 3036-3036 [ DOI: doi:10.1038/s41467-024-47468-1 Pubmed: 38589439 ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- 4f2hc-asc-1 complex (Complex from Homo sapiens)
- Asc-type amino acid transporter 1 (56 kDa, Protein from Homo sapiens)
- Amino acid transporter heavy chain slc3a2 (68 kDa, Protein from Homo sapiens)
- D-serine (105 Da, Ligand)
CryoEM structure of NaDC1 with Citrate
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 8w6c
Q-score: 0.631
Chi X, Chen Y, Li Y, Dai L, Zhang Y, Shen Y, Chen Y, Shi T, Yang H, Wang Z, Yan R
Sci Adv (2024) 10 pp. eadl3685-eadl3685 [ Pubmed: 38552027 DOI: doi:10.1126/sciadv.adl3685 ]
- Structure of nadc1 in complex with citrate (Complex from Homo sapiens)
- Citric acid (192 Da, Ligand)
- 1,2-diacyl-glycerol-3-sn-phosphate (704 Da, Ligand)
- Sodium ion (22 Da, Ligand)
- Solute carrier family 13 member 2 (65 kDa, Protein from Homo sapiens)
- Water (18 Da, Ligand)
- Cholesterol hemisuccinate (486 Da, Ligand)
CryoEM structure of NaDC1 in apo state
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 8w6d
Q-score: 0.662
Chi X, Chen Y, Li Y, Dai L, Zhang Y, Shen Y, Chen Y, Shi T, Yang H, Wang Z, Yan R
Sci Adv (2024) 10 pp. eadl3685-eadl3685 [ Pubmed: 38552027 DOI: doi:10.1126/sciadv.adl3685 ]
- Structure of nadc1 in apo state (Complex from Homo sapiens)
- 1,2-diacyl-glycerol-3-sn-phosphate (704 Da, Ligand)
- Sodium ion (22 Da, Ligand)
- Solute carrier family 13 member 2 (65 kDa, Protein from Homo sapiens)
- Cholesterol hemisuccinate (486 Da, Ligand)
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8w6g
Q-score: 0.552
Chi X, Chen Y, Li Y, Dai L, Zhang Y, Shen Y, Chen Y, Shi T, Yang H, Wang Z, Yan R
Sci Adv (2024) 10 pp. eadl3685-eadl3685 [ Pubmed: 38552027 DOI: doi:10.1126/sciadv.adl3685 ]
- 2-[[(~{e})-3-(4-pentylphenyl)prop-2-enoyl]amino]benzoic acid (337 Da, Ligand)
- 1,2-diacyl-glycerol-3-sn-phosphate (704 Da, Ligand)
- Solute carrier family 13 member 2 (65 kDa, Protein from Homo sapiens)
- Nadc1 in complex with inhibitor n-(p-amylcinnamoyl)anthranilic acid (aca) (Complex from Homo sapiens)
- Cholesterol hemisuccinate (486 Da, Ligand)