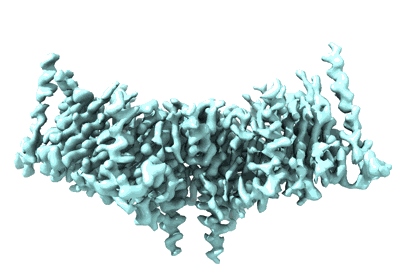
Ghanaian virus fusion glycoprotein (GhV F)
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 8tvb
Q-score: 0.56
Wang Z, McCallum M, Yan L, Gibson CA, Sharkey W, Park YJ, Dang HV, Amaya M, Person A, Broder CC, Veesler D
PNAS (2024) 121 pp. e2314990121-e2314990121 [ DOI: doi:10.1073/pnas.2314990121 Pubmed: 38593070 ]
- Fusion glycoprotein f0,2-dehydro-3-deoxyphosphogluconate aldolase/4-hydroxy-2-oxoglutarate aldolase (96 kDa, Protein from Thermotoga maritima MSB8)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Prefusion ghanian henipavirus f fused to the i53-50a trimer via an intervening 16 gs linker (Complex from Henipavirus ghanaense)
Cryo-EM structure of the yeast TOM core complex crosslinked by BS3 (from TOM-TIM23 complex)
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 8w5k
Q-score: 0.436
Wang Q, Zhuang J, Huang R, Guan Z, Yan L, Hong S, Zhang L, Huang C, Liu Z, Yin P
Cell Discov (2024) 10 pp. 19-19 [ DOI: doi:10.1038/s41421-023-00643-y Pubmed: 38360717 ]
- (2r)-3-{[(s)-(2-aminoethoxy)(hydroxy)phosphoryl]oxy}-2-(tetradecanoyloxy)propyl tetradecanoate (635 Da, Ligand)
- Mitochondrial import receptor subunit tom40 (42 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Mitochondrial import receptor subunit tom5 (5 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Tom-tim23 complex (Complex from Saccharomyces cerevisiae S288C)
- Mitochondrial import receptor subunit tom6 (6 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Mitochondrial import receptor subunit tom7 (6 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Mitochondrial import receptor subunit tom22 (16 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
SARS-CoV-2 RNA E-RTC complex with RMP-nsp9 and GMPPNP
Resolution: 2.72 Å
EM Method: Single-particle
Fitted PDBs: 8gwk
Q-score: 0.423
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ DOI: doi:10.1016/j.cell.2022.09.037 Pubmed: 36335936 ]
- E-rtc_rmp-nsp9_gmppnp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Template (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- [(2~{r},3~{s},4~{r},5~{r})-5-(4-azanylpyrrolo[2,1-f][1,2,4]triazin-7-yl)-5-cyano-3,4-bis(oxidanyl)oxolan-2-yl]methyl dihydrogen phosphate (371 Da, Ligand)
- Water (18 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Phosphoaminophosphonic acid-guanylate ester (522 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
SARS-CoV-2 E-RTC bound with MMP-nsp9 and GMPPNP
Resolution: 2.64 Å
EM Method: Single-particle
Fitted PDBs: 8gwm
Q-score: 0.41
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ DOI: doi:10.1016/j.cell.2022.09.037 Pubmed: 36335936 ]
- 2'-deoxy-2'-fluoro-2'-methyluridine 5'-(trihydrogen diphosphate) (420 Da, Ligand)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_mmp-nsp9_gmppnp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Phosphoaminophosphonic acid-guanylate ester (522 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Rna (25-mer) (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Rna (26-mer) (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
Cryo-EM structure of the yeast TOM core complex (from TOM-TIM23 complex)
Resolution: 4.4 Å
EM Method: Single-particle
Fitted PDBs: 8w5j
Q-score: 0.281
Wang Q, Zhuang J, Huang R, Guan Z, Yan L, Hong S, Zhang L, Huang C, Liu Z, Yin P
Cell Discov (2024) 10 pp. 19-19 [ DOI: doi:10.1038/s41421-023-00643-y Pubmed: 38360717 ]
- (2r)-3-{[(s)-(2-aminoethoxy)(hydroxy)phosphoryl]oxy}-2-(tetradecanoyloxy)propyl tetradecanoate (635 Da, Ligand)
- Tom-tim23 complex (Complex from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Mitochondrial import receptor subunit tom40 (42 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Mitochondrial import receptor subunit tom5 (5 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Mitochondrial import receptor subunit tom6 (6 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Mitochondrial import receptor subunit tom7 (6 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
- Mitochondrial import receptor subunit tom22 (16 kDa, Protein from Saccharomyces cerevisiae (strain ATCC 204508 / S288c))
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 3.31 Å
EM Method: Single-particle
Fitted PDBs: 8gw1
Q-score: 0.332
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ DOI: doi:10.1016/j.cell.2022.09.037 Pubmed: 36335936 ]
- Template (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_ump-nsp9 (350 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Manganese (ii) ion (54 Da, Ligand)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Uridine-5'-monophosphate (324 Da, Ligand)
- Water (18 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Replicase polyprotein 1ab (108 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Rna (25-mer) (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
lymphocytic choriomeningitis virus polymerase- Matrix Z Protein Complex (LCMV L-Z Complex)
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 7x6v
Q-score: 0.393
Liu L, Wang P, Liu A, Zhang L, Yan L, Guo Y, Xiao G, Rao Z, Lou Z
Protein Cell (2023) 14 pp. 703-707 [ Pubmed: 37038286 DOI: doi:10.1093/procel/pwad018 ]
- Manganese (ii) ion (54 Da, Ligand)
- Ring finger protein z (10 kDa, Protein from Lymphocytic choriomeningitis virus (strain Armstrong))
- Lcmv polymerase l protein (Complex from Lymphocytic choriomeningitis virus (strain Armstrong))
- Matrix protein z (Complex from Lymphocytic choriomeningitis virus (strain Armstrong))
- The complex of lcmv polymerase with matrix protein z (Complex)
- Zinc ion (65 Da, Ligand)
- Rna-directed rna polymerase l (254 kDa, Protein from Lymphocytic choriomeningitis virus (strain Armstrong))
Cryo-EM structure of CD38 in complex with FTL004
Resolution: 3.86 Å
EM Method: Single-particle
Fitted PDBs: 8il3
Q-score: 0.387
Zhang G, Guo C, Wang Y, Zhang X, Liu S, Qu W, Chen C, Yan L, Yang Z, Zhang Z, Jiang X, Chen X, Liu H, Lai Q, Wei X, Lu Y, Zhao S, Deng H, Wang Y, Yu L, Yu H, Wu Y, Su Z, Chen P, Ren Z, Yu M, Qu F, Luo Y, Gou L, Li Q, Huang Y, Ma F, Yang J
J Hematol Oncol (2022) 15 pp. 177-177 [ Pubmed: 36581954 DOI: doi:10.1186/s13045-022-01395-0 ]
- Heavy chain (22 kDa, Protein from Homo sapiens)
- Adp-ribosyl cyclase/cyclic adp-ribose hydrolase 1 (26 kDa, Protein from Homo sapiens)
- Cryo-em structure of cd38 in complex with ftl004 (Complex from Homo sapiens)
- Light chain (23 kDa, Protein from Homo sapiens)
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 2.66 Å
EM Method: Single-particle
Fitted PDBs: 8gwe
Q-score: 0.019
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ DOI: doi:10.1016/j.cell.2022.09.037 Pubmed: 36335936 ]
- Replicase polyprotein 1a (8 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (5'-r(p*ap*up*up*a)-3') (1 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_rna-nsp9_gmppnp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Phosphoaminophosphonic acid-guanylate ester (522 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Helicase nsp13 (65 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Template (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analogue inhibitors
Resolution: 3.39 Å
EM Method: Single-particle
Fitted PDBs: 8gwf
Q-score: 0.29
Yan L, Huang Y, Ge J, Liu Z, Lu P, Huang B, Gao S, Wang J, Tan L, Ye S, Yu F, Lan W, Xu S, Zhou F, Shi L, Guddat LW, Gao Y, Rao Z, Lou Z
Cell (2022) 185 pp. 4347-4360.e17 [ DOI: doi:10.1016/j.cell.2022.09.037 Pubmed: 36335936 ]
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Helicase (66 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Rna (Complex)
- Water (18 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- E-rtc_rna-nsp9_gtp (Complex from Severe acute respiratory syndrome coronavirus 2)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Template (10 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Zinc ion (65 Da, Ligand)
- Primer (8 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Nsp (Complex)