Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
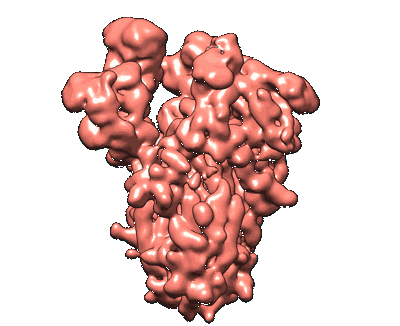
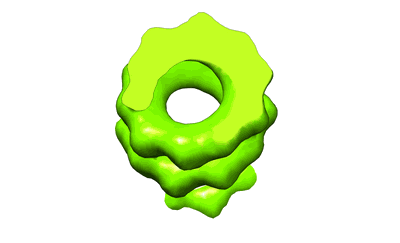
SARS-CoV-2 S protein S:A222V + S:D614G mutant 1-up
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7qdg
Q-score: 0.31
Ginex T, Marco-Marin C, Wieczor M, Mata CP, Krieger J, Ruiz-Rodriguez P, Lopez-Redondo ML, Frances-Gomez C, Melero R, Sanchez-Sorzano CO, Martinez M, Gougeard N, Forcada-Nadal A, Zamora-Caballero S, Gozalbo-Rovira R, Sanz-Frasquet C, Arranz R, Bravo J, Rubio V, Marina A, Geller R, Comas I, Gil C, Coscolla M, Orozco M, Llacer JL, Carazo JM
PLoS Pathog (2022) 18 pp. e1010631-e1010631 [ DOI: doi:10.1371/journal.ppat.1010631 Pubmed: 35816514 ]
SARS-CoV-2 S protein S:A222V + S:D614G mutant 2-up
Ginex T, Marco-Marin C, Wieczor M, Mata CP, Krieger J, Ruiz-Rodriguez P, Lopez-Redondo ML, Frances-Gomez C, Melero R, Sanchez-Sorzano CO, Martinez M, Gougeard N, Forcada-Nadal A, Zamora-Caballero S, Gozalbo-Rovira R, Sanz-Frasquet C, Arranz R, Bravo J, Rubio V, Marina A, Geller R, Comas I, Gil C, Coscolla M, Orozco M, Llacer JL, Carazo JM
PLoS Pathog (2022) 18 pp. e1010631-e1010631 [ DOI: doi:10.1371/journal.ppat.1010631 Pubmed: 35816514 ]
SARS-CoV-2 S protein S:A222V + S:D614G mutant 3-down
Ginex T, Marco-Marin C, Wieczor M, Mata CP, Krieger J, Ruiz-Rodriguez P, Lopez-Redondo ML, Frances-Gomez C, Melero R, Sanchez-Sorzano CO, Martinez M, Gougeard N, Forcada-Nadal A, Zamora-Caballero S, Gozalbo-Rovira R, Sanz-Frasquet C, Arranz R, Bravo J, Rubio V, Marina A, Geller R, Comas I, Gil C, Coscolla M, Orozco M, Llacer JL, Carazo JM
PLoS Pathog (2022) 18 pp. e1010631-e1010631 [ DOI: doi:10.1371/journal.ppat.1010631 Pubmed: 35816514 ]
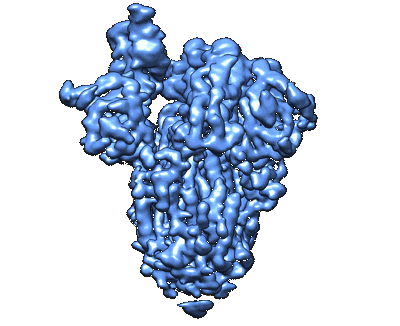
SARS-CoV-2 S protein S:D614G mutant 1-up
Resolution: 4.2 Å
EM Method: Single-particle
Fitted PDBs: 7qdh
Q-score: 0.079
Ginex T, Marco-Marin C, Wieczor M, Mata CP, Krieger J, Ruiz-Rodriguez P, Lopez-Redondo ML, Frances-Gomez C, Melero R, Sanchez-Sorzano CO, Martinez M, Gougeard N, Forcada-Nadal A, Zamora-Caballero S, Gozalbo-Rovira R, Sanz-Frasquet C, Arranz R, Bravo J, Rubio V, Marina A, Geller R, Comas I, Gil C, Coscolla M, Orozco M, Llacer JL, Carazo JM
PLoS Pathog (2022) 18 pp. e1010631-e1010631 [ DOI: doi:10.1371/journal.ppat.1010631 Pubmed: 35816514 ]
- Spike glycoprotein,fibritin (138 kDa, Protein from Escherichia virus T4)
- Sars-cov-2 s protein s:d614g mutant (340 kDa, Complex from Severe acute respiratory syndrome-related coronavirus)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
SARS-CoV-2 S protein S:D614G mutant 2-up
Ginex T, Marco-Marin C, Wieczor M, Mata CP, Krieger J, Ruiz-Rodriguez P, Lopez-Redondo ML, Frances-Gomez C, Melero R, Sanchez-Sorzano CO, Martinez M, Gougeard N, Forcada-Nadal A, Zamora-Caballero S, Gozalbo-Rovira R, Sanz-Frasquet C, Arranz R, Bravo J, Rubio V, Marina A, Geller R, Comas I, Gil C, Coscolla M, Orozco M, Llacer JL, Carazo JM
PLoS Pathog (2022) 18 pp. e1010631-e1010631 [ DOI: doi:10.1371/journal.ppat.1010631 Pubmed: 35816514 ]
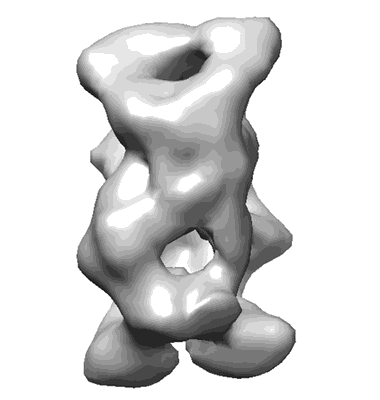
Putative tetramer of human CPAP (residues 897-1338)
Alvarez-Cabrera AL, Delgado S, Gil-Carton D, Mortuza GB, Montoya G, Sorzano CO, Tang TK, Carazo JM
Front Mol Biosci (2017) 4 pp. 17-17 [ DOI: doi:10.3389/fmolb.2017.00017 Pubmed: 28396859 ]
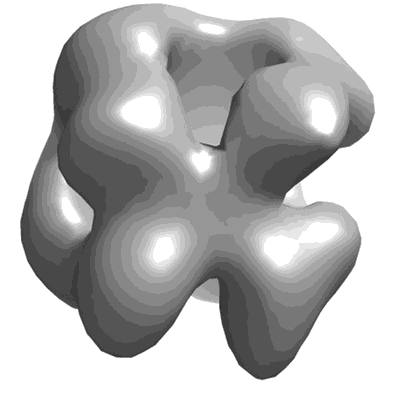
Putative dimer of human CPAP (residues 897-1338)
Alvarez-Cabrera AL, Delgado S, Gil-Carton D, Mortuza GB, Montoya G, Sorzano CO, Tang TK, Carazo JM
Front Mol Biosci (2017) 4 pp. 17-17 [ DOI: doi:10.3389/fmolb.2017.00017 Pubmed: 28396859 ]
- Putative homodimer of human cpap (residues 897-1338) (108 kDa, Complex from Homo sapiens)
Resolution: 20.0 Å
EM Method: Helical reconstruction
Fitted PDBs: 4blf
Q-score: 0.044
Bharat TA, Zbaida D, Eisenstein M, Frankenstein Z, Mehlman T, Weiner L, Sorzano CO, Barak Y, Albeck S, Briggs JA, Wolf SG, Elbaum M
Structure (2013) 21 pp. 1158-1167 [ Pubmed: 23769668 DOI: doi:10.1016/j.str.2013.04.027 ]
Electron cryo-microscopy of ABC BmrA in 12-fold symmetry rings
Fribourg PF, Chami M, Sorzano CO, Gubellini F, Marabini R, Marco S, Jault JM, Levy D
J Mol Biol (2014) 426 pp. 2059-2069 [ DOI: doi:10.1016/j.jmb.2014.03.002 Pubmed: 24630999 ]
- Bmra (3 MDa, Protein from Bacillus subtilis)
- Bmra reconstituted at high density into lipid bilayer. (3 MDa, Sample)
Electron cryo-microscopy of ABC BmrA in 13-fold symmetry rings
Fribourg PF, Chami M, Sorzano CO, Gubellini F, Marabini R, Marco S, Jault JM, Levy D
J Mol Biol (2014) 426 pp. 2059-2069 [ DOI: doi:10.1016/j.jmb.2014.03.002 Pubmed: 24630999 ]
- Bmra (5 MDa, Protein from Bacillus subtilis)
- Bmra reconstituted at high density into lipid bilayer. (5 MDa, Sample)