Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
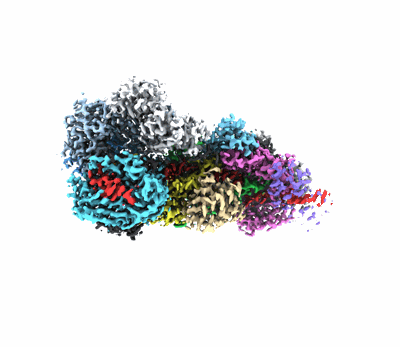
PSI-LHCI of the red alga Cyanidium caldarium RK-1 (NIES-2137)
Resolution: 1.92 Å
EM Method: Single-particle
Fitted PDBs: 8wey
Q-score: 0.662
Kato K, Hamaguchi T, Kumazawa M, Nakajima Y, Ifuku K, Hirooka S, Hirose Y, Miyagishima SY, Suzuki T, Kawakami K, Dohmae N, Yonekura K, Shen JR, Nagao R
PNAS (2024) 121 pp. e2319658121-e2319658121 [ Pubmed: 38442179 DOI: doi:10.1073/pnas.2319658121 ]
- Photosystem i reaction center subunit xi (15 kDa, Protein from Cyanidium caldarium)
- Photosystem i iron-sulfur center (8 kDa, Protein from Cyanidium caldarium)
- Chlorophyll a (893 Da, Ligand)
- (1~{r})-3,5,5-trimethyl-4-[(1~{e},3~{e},5~{e},7~{e},9~{e},11~{e},13~{e},15~{e},17~{e})-3,7,12,16-tetramethyl-18-[(4~{r} )-2,6,6-trimethyl-4-oxidanyl-cyclohexen-1-yl]octadeca-1,3,5,7,9,11,13,15,17-nonaenyl]cyclohex-3-en-1-ol (568 Da, Ligand)
- Lhcr3 (23 kDa, Protein from Cyanidium caldarium)
- Lhcr2 (24 kDa, Protein from Cyanidium caldarium)
- Dodecyl-beta-d-maltoside (510 Da, Ligand)
- Photosystem i p700 chlorophyll a apoprotein a1 (82 kDa, Protein from Cyanidium caldarium)
- Photosystem i reaction center subunit viii (3 kDa, Protein from Cyanidium caldarium)
- Chlorophyll a isomer (893 Da, Ligand)
- Digalactosyl diacyl glycerol (dgdg) (949 Da, Ligand)
- Photosystem i reaction center subunit xii (3 kDa, Protein from Cyanidium caldarium)
- Photosystem i reaction center subunit ii (15 kDa, Protein from Cyanidium caldarium)
- Iron/sulfur cluster (351 Da, Ligand)
- Photosystem i reaction center subunit ix (4 kDa, Protein from Cyanidium caldarium)
- Lhcr1 (24 kDa, Protein from Cyanidium caldarium)
- Psi-lhci (550 kDa, Complex from Cyanidium caldarium)
- Photosystem i reaction center subunit iv (6 kDa, Protein from Cyanidium caldarium)
- 1,2-dipalmitoyl-phosphatidyl-glycerole (722 Da, Ligand)
- Beta-carotene (536 Da, Ligand)
- Photosystem i subunit o (16 kDa, Protein from Cyanidium caldarium)
- Unknown ligand (Ligand)
- Photosystem i p700 chlorophyll a apoprotein a2 (82 kDa, Protein from Cyanidium caldarium)
- Photosystem i reaction center subunit x (7 kDa, Protein from Cyanidium caldarium)
- Water (18 Da, Ligand)
- Phylloquinone (450 Da, Ligand)
- Photosystem i reaction center subunit iii (20 kDa, Protein from Cyanidium caldarium)
Human ATAD2 Walker B mutant, ATP state
Resolution: 3.15 Å
EM Method: Single-particle
Fitted PDBs: 8h3h
Q-score: 0.571
Cho C, Ganser C, Uchihashi T, Kato K, Song JJ
Commun Biol (2023) 6 pp. 993-993 [ DOI: doi:10.1038/s42003-023-05373-1 Pubmed: 37770645 ]
- Atad2 (Complex from Homo sapiens)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Atpase family aaa domain-containing protein 2 (93 kDa, Protein from Homo sapiens)
Human ATAD2 Walker B mutant-H3/H4K5Q complex, ATP state
Resolution: 3.79 Å
EM Method: Single-particle
Fitted PDBs: 8juw
Q-score: 0.364
Cho C, Ganser C, Uchihashi T, Kato K, Song JJ
Commun Biol (2023) 6 pp. 993-993 [ DOI: doi:10.1038/s42003-023-05373-1 Pubmed: 37770645 ]
- Atad2 (Complex from Homo sapiens)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Atpase family aaa domain-containing protein 2 (93 kDa, Protein from Homo sapiens)
Human ATAD2 Walker B mutant-H3/H4K5Q complex, ATP state (Class II)
Resolution: 4.34 Å
EM Method: Single-particle
Fitted PDBs: 8juy
Q-score: 0.239
Cho C, Ganser C, Uchihashi T, Kato K, Song JJ
Commun Biol (2023) 6 pp. 993-993 [ DOI: doi:10.1038/s42003-023-05373-1 Pubmed: 37770645 ]
- Atad2 (Complex from Homo sapiens)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Atpase family aaa domain-containing protein 2 (93 kDa, Protein from Homo sapiens)
Human ATAD2 Walker B mutant-H3/H4K5Q complex, ATP state (Class III)
Resolution: 4.29 Å
EM Method: Single-particle
Fitted PDBs: 8juz
Q-score: 0.18
Cho C, Ganser C, Uchihashi T, Kato K, Song JJ
Commun Biol (2023) 6 pp. 993-993 [ DOI: doi:10.1038/s42003-023-05373-1 Pubmed: 37770645 ]
- Atad2 (Complex from Homo sapiens)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Atpase family aaa domain-containing protein 2 (93 kDa, Protein from Homo sapiens)
Conformation 1 of SARS-CoV-2 Omicron BA.1 Variant Spike protein complexed with MO1 Fab
Resolution: 2.48 Å
EM Method: Single-particle
Fitted PDBs: 8h3m
Q-score: 0.635
Ishimaru H, Nishimura M, Tjan LH, Sutandhio S, Marini MI, Effendi GB, Shigematsu H, Kato K, Hasegawa N, Aoki K, Kurahashi Y, Furukawa K, Shinohara M, Nakamura T, Arii J, Nagano T, Nakamura S, Sano S, Iwata S, Okamura S, Mori Y
J Virol (2023) 97 pp. e0028623-e0028623 [ Pubmed: 37191569 DOI: doi:10.1128/jvi.00286-23 ]
- Spike protein complexed with mo1 fab (500 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Mo1 heavy chain (48 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Spike glycoprotein (139 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
Conformation 2 of SARS-CoV-2 Omicron BA.1 Variant Spike protein complexed with MO1 Fab
Resolution: 2.73 Å
EM Method: Single-particle
Fitted PDBs: 8h3n
Q-score: 0.611
Ishimaru H, Nishimura M, Tjan LH, Sutandhio S, Marini MI, Effendi GB, Shigematsu H, Kato K, Hasegawa N, Aoki K, Kurahashi Y, Furukawa K, Shinohara M, Nakamura T, Arii J, Nagano T, Nakamura S, Sano S, Iwata S, Okamura S, Mori Y
J Virol (2023) 97 pp. e0028623-e0028623 [ Pubmed: 37191569 DOI: doi:10.1128/jvi.00286-23 ]
- Mo1 light chain (23 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Spike protein complexed with mo1 fab (500 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Mo1 heavy-chain (48 kDa, Protein from Homo sapiens)
- Spike glycoprotein (139 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
Structure of the Anabaena PSI-monomer-IsiA supercomplex
Resolution: 2.62 Å
EM Method: Single-particle
Fitted PDBs: 7y3f
Q-score: 0.526
Nagao R, Kato K, Hamaguchi T, Ueno Y, Tsuboshita N, Shimizu S, Furutani M, Ehira S, Nakajima Y, Kawakami K, Suzuki T, Dohmae N, Akimoto S, Yonekura K, Shen JR
Nat Commun (2023) 14 pp. 920-920 [ Pubmed: 36805598 DOI: doi:10.1038/s41467-023-36504-1 ]
- Chlorophyll a (893 Da, Ligand)
- Photosystem i p700 chlorophyll a apoprotein a1 (83 kDa, Protein from Nostoc sp.)
- Psam (4 kDa, Protein from Nostoc sp.)
- Photosystem i p700 chlorophyll a apoprotein a2 1 (83 kDa, Protein from Nostoc sp.)
- Iron stress-induced chlorophyll-binding protein (37 kDa, Protein from Nostoc sp.)
- Dodecyl-beta-d-maltoside (510 Da, Ligand)
- 1,2-distearoyl-monogalactosyl-diglyceride (787 Da, Ligand)
- Photosystem i reaction center subunit ix (5 kDa, Protein from Nostoc sp.)
- Photosystem i reaction center subunit viii (5 kDa, Protein from Nostoc sp.)
- Photosystem i reaction center subunit ii (15 kDa, Protein from Nostoc sp.)
- Photosystem i 4.8 kda protein (4 kDa, Protein from Nostoc sp.)
- Chlorophyll a isomer (893 Da, Ligand)
- Psi-monomer-isia (640 kDa, Complex from Nostoc sp.)
- Iron/sulfur cluster (351 Da, Ligand)
- Photosystem i reaction center subunit iv (7 kDa, Protein from Nostoc sp.)
- 1,2-dipalmitoyl-phosphatidyl-glycerole (722 Da, Ligand)
- Beta-carotene (536 Da, Ligand)
- Photosystem i iron-sulfur center (8 kDa, Protein from Nostoc sp.)
- Photosystem i reaction center subunit iii (17 kDa, Protein from Nostoc sp.)
- Photosystem i reaction center subunit xi (51 kDa, Protein from Nostoc sp.)
- Phylloquinone (450 Da, Ligand)
- Isia (27 kDa, Protein from Nostoc sp.)
- Water (18 Da, Ligand)
- Unknown (6 kDa, Protein from Nostoc sp.)
Structure of the OgeuIscB-omega RNA-target DNA complex
Resolution: 2.55 Å
EM Method: Single-particle
Fitted PDBs: 7xht
Q-score: 0.619
Kato K, Okazaki S, Kannan S, Altae-Tran H, Esra Demircioglu F, Isayama Y, Ishikawa J, Fukuda M, Macrae RK, Nishizawa T, Makarova KS, Koonin EV, Zhang F, Nishimasu H
Nat Commun (2022) 13 pp. 6719-6719 [ DOI: doi:10.1038/s41467-022-34378-3 Pubmed: 36344504 ]
- Iscb-omegarna (Complex from metagenome)
- Ogeuiscb (56 kDa, Protein from metagenome)
- Dna (5'-d(p*gp*ap*ap*gp*ap*ap*ap*ap*cp*cp*ap*t)-3') (4 kDa, DNA from synthetic construct)
- Structure of the iscb-omegarna ribonucleoprotein complex (Complex)
- Dna (49-mer) (15 kDa, DNA from synthetic construct)
- Lauryl dimethylamine-n-oxide (229 Da, Ligand)
- Rna (228-mer) (73 kDa, RNA from metagenome)
- Magnesium ion (24 Da, Ligand)
- Dna (Complex)
Structure of the Cas7-11-Csx29-guide RNA complex
Resolution: 2.49 Å
EM Method: Single-particle
Fitted PDBs: 7y9x
Q-score: 0.59
Kato K, Okazaki S, Schmitt-Ulms C, Jiang K, Zhou W, Ishikawa J, Isayama Y, Adachi S, Nishizawa T, Makarova KS, Koonin EV, Abudayyeh OO, Gootenberg JS, Nishimasu H
Science (2022) 378 pp. 882-889 [ Pubmed: 36423304 DOI: doi:10.1126/science.add7347 ]
- Cas7-11, csx29 (Complex from Desulfonema ishimotonii)
- Rna (Complex)
- Crispr-associated ramp family protein (185 kDa, Protein from Desulfonema ishimotonii)
- Crrna (12 kDa, RNA from Desulfonema ishimotonii)
- Cas7-11 in complex with a crrna and its target rna (Complex from Desulfonema ishimotonii)
- Zinc ion (65 Da, Ligand)
- Chat domain-containing protein (88 kDa, Protein from Desulfonema ishimotonii)