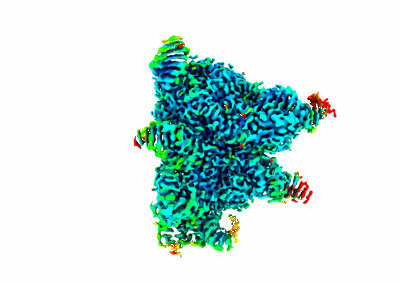
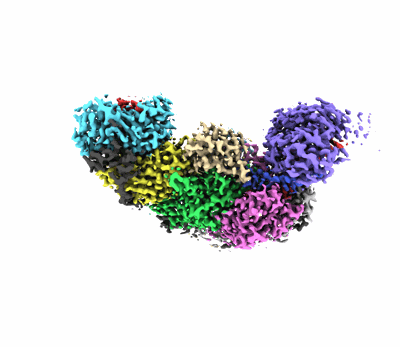Conformation 1 of SARS-CoV-2 Omicron BA.1 Variant Spike protein complexed with MO1 Fab
Resolution: 2.48 Å
EM Method: Single-particle
Fitted PDBs: 8h3m
Q-score: 0.635
Ishimaru H, Nishimura M, Tjan LH, Sutandhio S, Marini MI, Effendi GB, Shigematsu H, Kato K, Hasegawa N, Aoki K, Kurahashi Y, Furukawa K, Shinohara M, Nakamura T, Arii J, Nagano T, Nakamura S, Sano S, Iwata S, Okamura S, Mori Y
J Virol (2023) 97 pp. e0028623-e0028623 [ DOI: doi:10.1128/jvi.00286-23 Pubmed: 37191569 ]
- Mo1 heavy chain (48 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Spike glycoprotein (139 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Spike protein complexed with mo1 fab (500 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
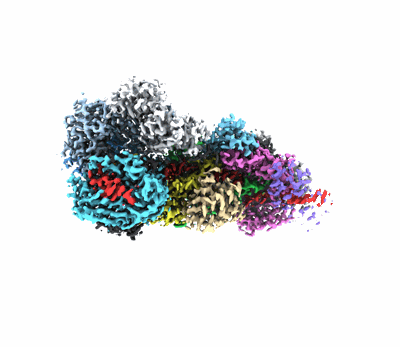
Conformation 2 of SARS-CoV-2 Omicron BA.1 Variant Spike protein complexed with MO1 Fab
Resolution: 2.73 Å
EM Method: Single-particle
Fitted PDBs: 8h3n
Q-score: 0.611
Ishimaru H, Nishimura M, Tjan LH, Sutandhio S, Marini MI, Effendi GB, Shigematsu H, Kato K, Hasegawa N, Aoki K, Kurahashi Y, Furukawa K, Shinohara M, Nakamura T, Arii J, Nagano T, Nakamura S, Sano S, Iwata S, Okamura S, Mori Y
J Virol (2023) 97 pp. e0028623-e0028623 [ DOI: doi:10.1128/jvi.00286-23 Pubmed: 37191569 ]
- Mo1 heavy-chain (48 kDa, Protein from Homo sapiens)
- Mo1 light chain (23 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Spike protein complexed with mo1 fab (500 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Spike glycoprotein (139 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
Structure of the Anabaena PSI-monomer-IsiA supercomplex
Resolution: 2.62 Å
EM Method: Single-particle
Fitted PDBs: 7y3f
Q-score: 0.526
Nagao R, Kato K, Hamaguchi T, Ueno Y, Tsuboshita N, Shimizu S, Furutani M, Ehira S, Nakajima Y, Kawakami K, Suzuki T, Dohmae N, Akimoto S, Yonekura K, Shen JR
Nat Commun (2023) 14 pp. 920-920 [ Pubmed: 36805598 DOI: doi:10.1038/s41467-023-36504-1 ]
- Photosystem i reaction center subunit ix (5 kDa, Protein from Nostoc sp.)
- Iron stress-induced chlorophyll-binding protein (37 kDa, Protein from Nostoc sp.)
- Phylloquinone (450 Da, Ligand)
- Psam (4 kDa, Protein from Nostoc sp.)
- Photosystem i iron-sulfur center (8 kDa, Protein from Nostoc sp.)
- Chlorophyll a isomer (893 Da, Ligand)
- Photosystem i reaction center subunit ii (15 kDa, Protein from Nostoc sp.)
- Chlorophyll a (893 Da, Ligand)
- 1,2-distearoyl-monogalactosyl-diglyceride (787 Da, Ligand)
- Isia (27 kDa, Protein from Nostoc sp.)
- 1,2-dipalmitoyl-phosphatidyl-glycerole (722 Da, Ligand)
- Beta-carotene (536 Da, Ligand)
- Photosystem i p700 chlorophyll a apoprotein a1 (83 kDa, Protein from Nostoc sp.)
- Photosystem i reaction center subunit viii (5 kDa, Protein from Nostoc sp.)
- Photosystem i reaction center subunit iii (17 kDa, Protein from Nostoc sp.)
- Photosystem i reaction center subunit xi (51 kDa, Protein from Nostoc sp.)
- Unknown (6 kDa, Protein from Nostoc sp.)
- Photosystem i reaction center subunit iv (7 kDa, Protein from Nostoc sp.)
- Water (18 Da, Ligand)
- Photosystem i p700 chlorophyll a apoprotein a2 1 (83 kDa, Protein from Nostoc sp.)
- Psi-monomer-isia (640 kDa, Complex from Nostoc sp.)
- Iron/sulfur cluster (351 Da, Ligand)
- Photosystem i 4.8 kda protein (4 kDa, Protein from Nostoc sp.)
- Dodecyl-beta-d-maltoside (510 Da, Ligand)
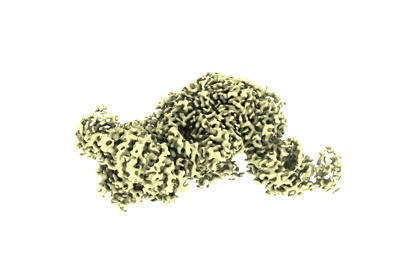
Structure of the OgeuIscB-omega RNA-target DNA complex
Resolution: 2.55 Å
EM Method: Single-particle
Fitted PDBs: 7xht
Q-score: 0.619
Kato K, Okazaki S, Kannan S, Altae-Tran H, Esra Demircioglu F, Isayama Y, Ishikawa J, Fukuda M, Macrae RK, Nishizawa T, Makarova KS, Koonin EV, Zhang F, Nishimasu H
Nat Commun (2022) 13 pp. 6719-6719 [ Pubmed: 36344504 DOI: doi:10.1038/s41467-022-34378-3 ]
- Rna (228-mer) (73 kDa, RNA from metagenome)
- Structure of the iscb-omegarna ribonucleoprotein complex (Complex)
- Lauryl dimethylamine-n-oxide (229 Da, Ligand)
- Dna (5'-d(p*gp*ap*ap*gp*ap*ap*ap*ap*cp*cp*ap*t)-3') (4 kDa, DNA from synthetic construct)
- Ogeuiscb (56 kDa, Protein from metagenome)
- Dna (49-mer) (15 kDa, DNA from synthetic construct)
- Iscb-omegarna (Complex from metagenome)
- Magnesium ion (24 Da, Ligand)
- Dna (Complex)
Structure of the Cas7-11-Csx29-guide RNA complex
Resolution: 2.49 Å
EM Method: Single-particle
Fitted PDBs: 7y9x
Q-score: 0.59
Kato K, Okazaki S, Schmitt-Ulms C, Jiang K, Zhou W, Ishikawa J, Isayama Y, Adachi S, Nishizawa T, Makarova KS, Koonin EV, Abudayyeh OO, Gootenberg JS, Nishimasu H
Science (2022) 378 pp. 882-889 [ Pubmed: 36423304 DOI: doi:10.1126/science.add7347 ]
- Cas7-11 in complex with a crrna and its target rna (Complex from Desulfonema ishimotonii)
- Chat domain-containing protein (88 kDa, Protein from Desulfonema ishimotonii)
- Rna (Complex)
- Crrna (12 kDa, RNA from Desulfonema ishimotonii)
- Zinc ion (65 Da, Ligand)
- Crispr-associated ramp family protein (185 kDa, Protein from Desulfonema ishimotonii)
- Cas7-11, csx29 (Complex from Desulfonema ishimotonii)
Structure of the Cas7-11-Csx29-guide RNA-target RNA (no PFS) complex
Resolution: 2.77 Å
EM Method: Single-particle
Fitted PDBs: 7y9y
Q-score: 0.564
Kato K, Okazaki S, Schmitt-Ulms C, Jiang K, Zhou W, Ishikawa J, Isayama Y, Adachi S, Nishizawa T, Makarova KS, Koonin EV, Abudayyeh OO, Gootenberg JS, Nishimasu H
Science (2022) 378 pp. 882-889 [ Pubmed: 36423304 DOI: doi:10.1126/science.add7347 ]
- Crispr-associated ramp family protein (185 kDa, Protein from Desulfonema ishimotonii)
- Chat domain-containing protein (88 kDa, Protein from Desulfonema ishimotonii)
- Cas7-11 in complex with a crrna and its target rna (Complex from Desulfonema ishimotonii)
- Rna (27-mer) (8 kDa, RNA from Desulfonema ishimotonii)
- Crrna, target rna (Complex from Desulfonema ishimotonii)
- Rna (38-mer) (12 kDa, RNA from Desulfonema ishimotonii)
- Cas7-11, csx29 (Complex)
- Zinc ion (65 Da, Ligand)
Structure of the Cas7-11-Csx29-guide RNA-target RNA (non-matching PFS) complex
Resolution: 2.84 Å
EM Method: Single-particle
Fitted PDBs: 8gs2
Q-score: 0.571
Kato K, Okazaki S, Schmitt-Ulms C, Jiang K, Zhou W, Ishikawa J, Isayama Y, Adachi S, Nishizawa T, Makarova KS, Koonin EV, Abudayyeh OO, Gootenberg JS, Nishimasu H
Science (2022) 378 pp. 882-889 [ Pubmed: 36423304 DOI: doi:10.1126/science.add7347 ]
- Adenosine monophosphate (347 Da, Ligand)
- Crrna (30 kDa, RNA from Desulfonema ishimotonii)
- Crispr-associated ramp family protein (185 kDa, Protein from Desulfonema ishimotonii)
- Chat domain-containing protein (88 kDa, Protein from Desulfonema ishimotonii)
- Crrna (Complex)
- Cas7-11 in complex with a crrna and its target rna (Complex from Desulfonema ishimotonii)
- Target rna with 3'-tag (Complex from Desulfonema ishimotonii)
- Cytidine-5'-monophosphate (323 Da, Ligand)
- Cas7-11, csx29 (Complex)
- Zinc ion (65 Da, Ligand)
- Target rna with non-matching pfs (10 kDa, RNA from Desulfonema ishimotonii)
Structure of Cas7-11 in complex with guide RNA and target RNA
Resolution: 2.45 Å
EM Method: Single-particle
Fitted PDBs: 7wah
Q-score: 0.561
Kato K, Zhou W, Okazaki S, Isayama Y, Nishizawa T, Gootenberg JS, Abudayyeh OO, Nishimasu H
Cell (2022) 185 pp. 2324-2337.e16 [ Pubmed: 35643083 DOI: doi:10.1016/j.cell.2022.05.003 ]
- Tgrna (5'-r(p*up*ap*cp*cp*cp*ap*up*gp*up*cp*gp*ap*ap*gp*ap*cp*ap*ap*cp*ap*ap*ap*g)-3') (7 kDa, RNA from Escherichia coli)
- Cas7-11 (Complex)
- Crrna (39-mer) (12 kDa, RNA from Escherichia coli)
- Crrna and target rna (Complex from Desulfonema ishimotonii)
- Cas7-11 in complex with a crrna and its target rna (Complex from Escherichia coli)
- Zinc ion (65 Da, Ligand)
- Crispr-associated ramp family protein (185 kDa, Protein from Desulfonema ishimotonii)
Cryo-EM structure of a primordial cyanobacterial photosystem I
Resolution: 2.04 Å
EM Method: Single-particle
Fitted PDBs: 7f4v
Q-score: 0.752
Kato K, Hamaguchi T, Nagao R, Kawakami K, Ueno Y, Suzuki T, Uchida H, Murakami A, Nakajima Y, Yokono M, Akimoto S, Dohmae N, Yonekura K, Shen JR
bioRxiv (2022) [ DOI: doi:10.1101/2021.09.29.462462 ]
- Photosystem i reaction center subunit iv (7 kDa, Protein from Gloeobacter violaceus (strain ATCC 29082 / PCC 7421))
- Photosystem i reaction center subunit iii (19 kDa, Protein from Gloeobacter violaceus (strain ATCC 29082 / PCC 7421))
- Chlorophyll a isomer (893 Da, Ligand)
- Chlorophyll a (893 Da, Ligand)
- Photosystem i reaction center subunit xii (3 kDa, Protein from Gloeobacter violaceus (strain ATCC 29082 / PCC 7421))
- 1,2-distearoyl-monogalactosyl-diglyceride (787 Da, Ligand)
- 1,2-dipalmitoyl-phosphatidyl-glycerole (722 Da, Ligand)
- Beta-carotene (536 Da, Ligand)
- Menaquinone-4 (444 Da, Ligand)
- Photosystem i reaction center subunit xi (15 kDa, Protein from Gloeobacter violaceus (strain ATCC 29082 / PCC 7421))
- Psi trimer (1 MDa, Complex from Gloeobacter violaceus PCC 7421)
- Photosystem i reaction center subunit z (3 kDa, Protein from Gloeobacter violaceus (strain ATCC 29082 / PCC 7421))
- Photosystem i p700 chlorophyll a apoprotein a2 (96 kDa, Protein from Gloeobacter violaceus (strain ATCC 29082 / PCC 7421))
- Water (18 Da, Ligand)
- Photosystem i iron-sulfur center (8 kDa, Protein from Gloeobacter violaceus (strain ATCC 29082 / PCC 7421))
- Iron/sulfur cluster (351 Da, Ligand)
- Photosystem i p700 chlorophyll a apoprotein a1 (86 kDa, Protein from Gloeobacter violaceus (strain ATCC 29082 / PCC 7421))
- Unknown protein (2 kDa, Protein from Gloeobacter violaceus (strain ATCC 29082 / PCC 7421))
- Photosystem i reaction center subunit ii (15 kDa, Protein from Gloeobacter violaceus (strain ATCC 29082 / PCC 7421))
Structure of C2S2M2-type PSII-FCPII supercomplex from diatom
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 7vd5
Q-score: 0.543
Nagao R, Kato K, Kumazawa M, Ifuku K, Yokono M, Suzuki T, Dohmae N, Akita F, Akimoto S, Miyazaki N, Shen JR
Nat Commun (2022) 13 pp. 1764- [ DOI: doi:10.1038/s41467-022-29294-5 ]
- Fcpb5, fucoxanthin chlorophyll a/c-binding protein (24 kDa, Protein from Chaetoceros gracilis)
- Fe (ii) ion (55 Da, Ligand)
- Photosystem ii cp43 reaction center protein (49 kDa, Protein from Chaetoceros gracilis)
- Photosystem ii reaction center protein h (7 kDa, Protein from Chaetoceros gracilis)
- Photosystem ii cp47 reaction center protein (53 kDa, Protein from Chaetoceros gracilis)
- Cytochrome b559 subunit alpha (8 kDa, Protein from Chaetoceros gracilis)
- Photosystem ii reaction center protein l (4 kDa, Protein from Chaetoceros gracilis)
- Unknown protein 1 (2 kDa, Protein from Chaetoceros gracilis)
- Photosystem ii subunit (4 kDa, Protein from Chaetoceros gracilis)
- Extrinsic protein in photosystem ii (16 kDa, Protein from Chaetoceros gracilis)
- Cytochrome b559 subunit beta (3 kDa, Protein from Chaetoceros gracilis)
- Photosystem ii reaction center protein z (6 kDa, Protein from Chaetoceros gracilis)
- Fcpb2, fucoxanthin chlorophyll a/c-binding protein (18 kDa, Protein from Chaetoceros gracilis)
- Chlorophyll a/b-binding protein (19 kDa, Protein from Chaetoceros gracilis)
- Photosystem ii reaction center protein t (3 kDa, Protein from Chaetoceros gracilis)
- Chlorophyll a (893 Da, Ligand)
- Photosystem ii d2 protein (37 kDa, Protein from Chaetoceros gracilis)
- Photosystem ii protein d1 (37 kDa, Protein from Chaetoceros gracilis)
- 1,2-distearoyl-monogalactosyl-diglyceride (787 Da, Ligand)
- 2,3-dimethyl-5-(3,7,11,15,19,23,27,31,35-nonamethyl-2,6,10,14,18,22,26,30,34-hexatriacontanonaenyl-2,5-cyclohexadiene-1,4-dione-2,3-dimethyl-5-solanesyl-1,4-benzoquinone (749 Da, Ligand)
- 1,2-dipalmitoyl-phosphatidyl-glycerole (722 Da, Ligand)
- Beta-carotene (536 Da, Ligand)
- Extrinsic protein in photosystem ii (26 kDa, Protein from Chaetoceros gracilis)
- Fcpb4, fucoxanthin chlorophyll a/c-binding protein (19 kDa, Protein from Chaetoceros gracilis)
- Pheophytin a (871 Da, Ligand)
- Extrinsic protein in photosystem ii (10 kDa, Protein from Chaetoceros gracilis)
- Unknown protein 0 (2 kDa, Protein from Chaetoceros gracilis)
- Ca-mn4-o5 cluster (339 Da, Ligand)
- (3s,3'r,5r,6s,7cis)-7',8'-didehydro-5,6-dihydro-5,6-epoxy-beta,beta-carotene-3,3'-diol (582 Da, Ligand)
- Chlorophyll a/b-binding protein (18 kDa, Protein from Chaetoceros gracilis)
- Chlorophyll c1 (610 Da, Ligand)
- Chlorophyll a/b-binding protein (18 kDa, Protein from Chaetoceros gracilis)
- Water (18 Da, Ligand)
- Chlorophyll a/b-binding protein (18 kDa, Protein from Chaetoceros gracilis)
- 1,2-di-o-acyl-3-o-[6-deoxy-6-sulfo-alpha-d-glucopyranosyl]-sn-glycerol (795 Da, Ligand)
- Cytochrome c-550 (14 kDa, Protein from Chaetoceros gracilis)
- (3s,3's,5r,5'r,6s,6'r,8'r)-3,5'-dihydroxy-8-oxo-6',7'-didehydro-5,5',6,6',7,8-hexahydro-5,6-epoxy-beta,beta-caroten-3'- yl acetate (658 Da, Ligand)
- Dodecyl-alpha-d-maltoside (510 Da, Ligand)
- Fcpb6, fucoxanthin chlorophyll a/c-binding protein (16 kDa, Protein from Chaetoceros gracilis)
- Photosystem ii reaction center protein ycf12 (3 kDa, Protein from Chaetoceros gracilis)
- Photosystem ii reaction center protein w (5 kDa, Protein from Chaetoceros gracilis)
- Digalactosyl diacyl glycerol (dgdg) (949 Da, Ligand)
- C2s2m2-type psii-fcpii supercomplex (1 MDa, Complex from Chaetoceros gracilis)
- Protoporphyrin ix containing fe (616 Da, Ligand)
- Unknown ligand (Ligand)
- Chlorophyll a/b-binding protein (18 kDa, Protein from Chaetoceros gracilis)
- Photosystem ii reaction center protein i (4 kDa, Protein from Chaetoceros gracilis)
- Bicarbonate ion (61 Da, Ligand)
- Photosystem ii reaction center x protein (3 kDa, Protein from Chaetoceros gracilis)
- Fcpb7, fucoxanthin chlorophyll a/c-binding protein (17 kDa, Protein from Chaetoceros gracilis)
- Photosystem ii reaction center protein k (4 kDa, Protein from Chaetoceros gracilis)
- Photosystem ii reaction center protein j (3 kDa, Protein from Chaetoceros gracilis)
- Fcpb3, fucoxanthin chlorophyll a/c-binding protein (19 kDa, Protein from Chaetoceros gracilis)