Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 8sq9
Q-score: 0.528
Small GI, Fedorova O, Olinares PDB, Chandanani J, Banerjee A, Choi YJ, Molina H, Chait BT, Darst SA, Campbell EA
Mol Cell (2023) 83 pp. 3921-3930.e7 [ DOI: doi:10.1016/j.molcel.2023.10.001 Pubmed: 37890482 ]
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Primer rna (11 kDa, RNA from Severe acute respiratory syndrome coronavirus)
- Template rna (17 kDa, RNA from Severe acute respiratory syndrome coronavirus)
- 5'-o-[(s)-hydroxy{[(s)-hydroxy(phosphonooxy)phosphoryl]methyl}phosphoryl]uridine (482 Da, Ligand)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 replication-transcription complex bound to nsp9 and umpcpp, as a pre-catalytic nmpylation intermediate (Complex from Severe acute respiratory syndrome coronavirus 2)
Resolution: 3.06 Å
EM Method: Single-particle
Fitted PDBs: 8sqj
Q-score: 0.511
Small GI, Fedorova O, Olinares PDB, Chandanani J, Banerjee A, Choi YJ, Molina H, Chait BT, Darst SA, Campbell EA
Mol Cell (2023) 83 pp. 3921-3930.e7 [ DOI: doi:10.1016/j.molcel.2023.10.001 Pubmed: 37890482 ]
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Template rna (17 kDa, RNA from Severe acute respiratory syndrome coronavirus)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- 5'-o-[(r)-hydroxy(thiophosphonooxy)phosphoryl]guanosine (459 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Sars-cov-2 replication-transcription complex bound to rna-nsp9, as a noncatalytic rna-nsp9 binding mode (Complex from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 5' utr (6 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- Primer rna (11 kDa, RNA from Severe acute respiratory syndrome coronavirus)
- Rna-directed rna polymerase (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
Resolution: 3.01 Å
EM Method: Single-particle
Fitted PDBs: 8sqk
Q-score: 0.533
Small GI, Fedorova O, Olinares PDB, Chandanani J, Banerjee A, Choi YJ, Molina H, Chait BT, Darst SA, Campbell EA
Mol Cell (2023) 83 pp. 3921-3930.e7 [ DOI: doi:10.1016/j.molcel.2023.10.001 Pubmed: 37890482 ]
- Non-structural protein 9 (12 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Template rna (11 kDa, RNA from Severe acute respiratory syndrome coronavirus)
- Rna-directed rna polymerase nsp12 (106 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Non-structural protein 7 (9 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Water (18 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Sars-cov-2 5' utr (6 kDa, RNA from Severe acute respiratory syndrome coronavirus 2)
- 5'-o-[(r)-hydroxy(thiophosphonooxy)phosphoryl]guanosine (459 Da, Ligand)
- Non-structural protein 8 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Primer rna (10 kDa, RNA from Severe acute respiratory syndrome coronavirus)
- Magnesium ion (24 Da, Ligand)
- Sars-cov-2 replication-transcription complex bound to rna-nsp9 and gdp-betas, as a pre-catalytic dernaylation/mrna capping intermediate (Complex from Severe acute respiratory syndrome coronavirus 2)
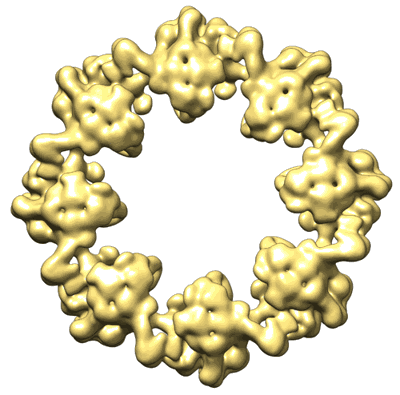
Composite multiscale structure of the yeast NPC
Akey CW, Echeverria I, Ouch C, Nudelman I, Shi Y, Wang J, Chait BT, Sali A, Fernandez-Martinez J, Rout MP
Mol Cell (2023) 83 pp. 3283-3302.e5 [ DOI: doi:10.1016/j.molcel.2023.08.025 Pubmed: 37738963 ]
Nuclear double outer ring of the yeast NPC
Akey CW, Echeverria I, Ouch C, Nudelman I, Shi Y, Wang J, Chait BT, Sali A, Fernandez-Martinez J, Rout MP
Mol Cell (2023) 83 pp. 3283-3302.e5 [ DOI: doi:10.1016/j.molcel.2023.08.025 Pubmed: 37738963 ]
- Nuclear pore complex (60 MDa, Complex from Saccharomyces cerevisiae)
Lumenal ring of the isolated yeast NPC
Akey CW, Echeverria I, Ouch C, Nudelman I, Shi Y, Wang J, Chait BT, Sali A, Fernandez-Martinez J, Rout MP
Mol Cell (2023) 83 pp. 3283-3302.e5 [ DOI: doi:10.1016/j.molcel.2023.08.025 Pubmed: 37738963 ]
Pom34-Pom152 membrane attachment site yeast NPC
Resolution: 7.0 Å
EM Method: Single-particle
Fitted PDBs: 8t9l
Q-score: 0.17
Akey CW, Echeverria I, Ouch C, Nudelman I, Shi Y, Wang J, Chait BT, Sali A, Fernandez-Martinez J, Rout MP
Mol Cell (2023) 83 pp. 3283-3302.e5 [ DOI: doi:10.1016/j.molcel.2023.08.025 Pubmed: 37738963 ]
- Pom34-pom152 inner ring anchor complex (Complex from Saccharomyces cerevisiae)
- Nucleoporin pom34 (34 kDa, Protein from Saccharomyces cerevisiae)
- Nucleoporin pom152 (151 kDa, Protein from Saccharomyces cerevisiae)
- Nuclear pore complex (Complex from Saccharomyces cerevisiae)
Cytoplasmic outer ring of the yeast NPC
Akey CW, Echeverria I, Ouch C, Nudelman I, Shi Y, Wang J, Chait BT, Sali A, Fernandez-Martinez J, Rout MP
Mol Cell (2023) 83 pp. 3283-3302.e5 [ DOI: doi:10.1016/j.molcel.2023.08.025 Pubmed: 37738963 ]
Central transporter/plug yeast NPC
Akey CW, Echeverria I, Ouch C, Nudelman I, Shi Y, Wang J, Chait BT, Sali A, Fernandez-Martinez J, Rout MP
Mol Cell (2023) 83 pp. 3283-3302.e5 [ DOI: doi:10.1016/j.molcel.2023.08.025 Pubmed: 37738963 ]
Double outer ring connectors of the yeast NPC
Akey CW, Echeverria I, Ouch C, Nudelman I, Shi Y, Wang J, Chait BT, Sali A, Fernandez-Martinez J, Rout MP
Mol Cell (2023) 83 pp. 3283-3302.e5 [ DOI: doi:10.1016/j.molcel.2023.08.025 Pubmed: 37738963 ]