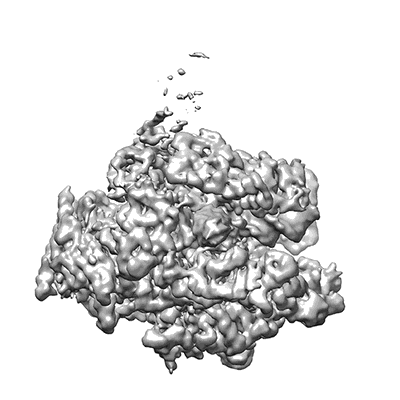Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
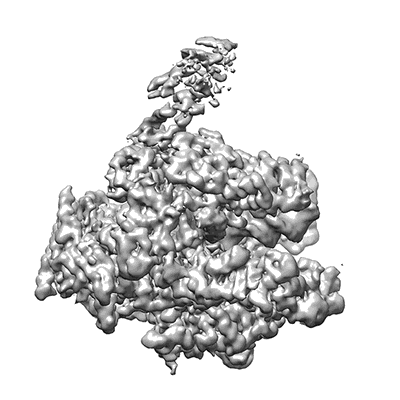
Subtomogram average of 80S ribosomes obtained using the VPP
Khoshouei M, Pfeffer S, Baumeister W, Foerster F, Danev R
J Struct Biol (2017) 197 pp. 94-101 [ DOI: doi:10.1016/j.jsb.2016.05.009 Pubmed: 27235783 ]
Subtomogram average of 80S ribosomes obtained using CTEM at one defocus
Khoshouei M, Pfeffer S, Baumeister W, Foerster F, Danev R
J Struct Biol (2017) 197 pp. 94-101 [ DOI: doi:10.1016/j.jsb.2016.05.009 Pubmed: 27235783 ]
Subtomogram average of 80S ribosomes obtained using CTEM with a range of defocus values
Khoshouei M, Pfeffer S, Baumeister W, Foerster F, Danev R
J Struct Biol (2017) 197 pp. 94-101 [ DOI: doi:10.1016/j.jsb.2016.05.009 Pubmed: 27235783 ]
Volta phase plate 3.2A resolution reconstruction of the T20S proteasome
eLife (2016) 5 [ Pubmed: 26949259 DOI: doi:10.7554/eLife.13046 ]
- 20s proteasome (700 kDa, Protein from Thermoplasma acidophilum)
- Thermoplasma acidophilum 20s proteasome (700 kDa, Sample)
cryo-EM structure of T20S proteasome
eLife (2016) 5 [ Pubmed: 26949259 DOI: doi:10.7554/eLife.13046 ]
- 20s proteasome (700 kDa, Protein from Thermoplasma acidophilum)
- Thermoplasma acidophilum 20s proteasome (700 kDa, Sample)
Volta phase plate cryo-EM of the small protein complex Prx3
Khoshouei M, Radjainia M, Phillips AJ, Gerrard JA, Mitra AK, Plitzko JM, Baumeister W, Danev R
Nat Commun (2016) 7 pp. 10534- [ DOI: doi:10.1038/ncomms10534 Pubmed: 26817416 ]
- Human peroxiredoxin 3 (257 kDa, Sample)
- Peroxiredoxin 3 (257 kDa, Protein from Homo sapiens)
Structure of transcribing mammalian RNA polymerase II (EC2)
Bernecky C, Herzog F, Baumeister W, Plitzko JM, Cramer P
Nature (2016) 529 pp. 551-554 [ Pubmed: 26789250 DOI: doi:10.1038/nature16482 ]
- Dna-rna synthetic construct (30 kDa, Macromolecule from unidentified)
- Bovine pol ii elongation complex (590 kDa, Sample)
- Dna-directed rna polymerase ii (520 kDa, Protein from Bos taurus)
- Dna-directed rna polymerase ii subunit grinl1a (40 kDa, Protein from Homo sapiens)
Structure of transcribing mammalian RNA polymerase II (EC3)
Bernecky C, Herzog F, Baumeister W, Plitzko JM, Cramer P
Nature (2016) 529 pp. 551-554 [ Pubmed: 26789250 DOI: doi:10.1038/nature16482 ]
- Dna-rna synthetic construct (30 kDa, Macromolecule from unidentified)
- Bovine pol ii elongation complex (590 kDa, Sample)
- Dna-directed rna polymerase ii (520 kDa, Protein from Bos taurus)
- Dna-directed rna polymerase ii subunit grinl1a (40 kDa, Protein from Homo sapiens)
Structure of transcribing mammalian RNA polymerase II (EC1)
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 5flm
Q-score: 0.442
Bernecky C, Herzog F, Baumeister W, Plitzko JM, Cramer P
Nature (2016) 529 pp. 551-554 [ Pubmed: 26789250 DOI: doi:10.1038/nature16482 ]
- Dna-rna synthetic construct (30 kDa, Macromolecule from unidentified)
- Bovine pol ii elongation complex (590 kDa, Sample)
- Dna-directed rna polymerase ii (520 kDa, Protein from Bos taurus)
- Dna-directed rna polymerase ii subunit grinl1a (40 kDa, Protein from Homo sapiens)
Cryo-electron tomography of a Golgi intracisternal protein array
Engel BD, Schaffer M, Albert S, Asano S, Plitzko JM, Baumeister W
PNAS (2015) 112 pp. 11264-11269 [ Pubmed: 26311849 DOI: doi:10.1073/pnas.1515337112 ]