Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
Resolution: 4.2 Å
EM Method: Helical reconstruction
Fitted PDBs: 8orh
Q-score: 0.571
Alempic JM, Bisio H, Villalta A, Santini S, Lartigue A, Schmitt A, Bugnot C, Notaro A, Belmudes L, Adrait A, Poirot O, Ptchelkine D, De Castro C, Coute Y, Abergel C
Microlife (2024) 5 pp. uqae006-uqae006 [ Pubmed: 38659623 DOI: doi:10.1093/femsml/uqae006 ]
- Putative gmc-type oxidoreductase (77 kDa, Protein from Mimivirus reunion)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- Mimivirus reunion (Virus from Mimivirus reunion)
Resolution: 4.3 Å
EM Method: Helical reconstruction
Fitted PDBs: 8ors
Q-score: 0.484
Alempic JM, Bisio H, Villalta A, Santini S, Lartigue A, Schmitt A, Bugnot C, Notaro A, Belmudes L, Adrait A, Poirot O, Ptchelkine D, De Castro C, Coute Y, Abergel C
Microlife (2024) 5 pp. uqae006-uqae006 [ Pubmed: 38659623 DOI: doi:10.1093/femsml/uqae006 ]
- Putative gmc-type oxidoreductase (77 kDa, Protein from Mimivirus reunion)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- Mimivirus reunion (Virus from Mimivirus reunion)
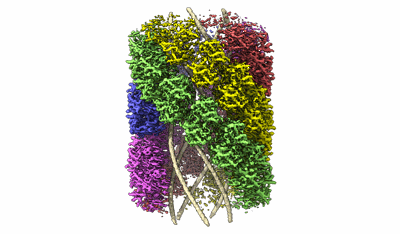
Tomogram of the Mimivirus genomic fiber
Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C
eLife (2022) 11 [ Pubmed: 35900198 DOI: doi:10.7554/eLife.77607 ]
Tomogram of the Mimivirus genomic fiber
Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C
eLife (2022) 11 [ Pubmed: 35900198 DOI: doi:10.7554/eLife.77607 ]
Tomogram of the Mimivirus genomic fiber
Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C
eLife (2022) 11 [ Pubmed: 35900198 DOI: doi:10.7554/eLife.77607 ]
Tomogram of the Mimivirus genomic fiber
Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C
eLife (2022) 11 [ Pubmed: 35900198 DOI: doi:10.7554/eLife.77607 ]
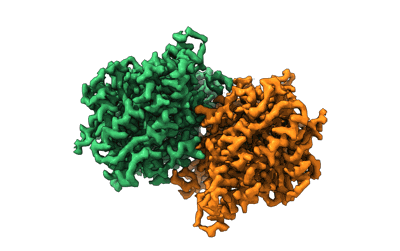
Structure of the Mimivirus genomic fibre asymmetric unit
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 7ptv
Q-score: 0.579
Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C
eLife (2022) 11 [ Pubmed: 35900198 DOI: doi:10.7554/eLife.77607 ]
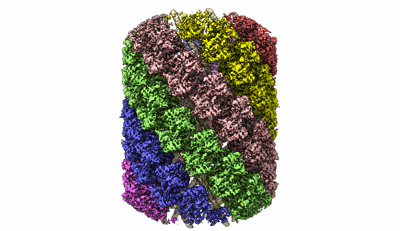
Structure of the Mimivirus genomic fibre in its compact 6-start helix form
Resolution: 4.0 Å
EM Method: Helical reconstruction
Fitted PDBs: 7yx3
Q-score: 0.458
Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C
eLife (2022) 11 [ Pubmed: 35900198 DOI: doi:10.7554/eLife.77607 ]
Structure of the Mimivirus genomic fibre in its compact 5-start helix form
Resolution: 3.7 Å
EM Method: Helical reconstruction
Fitted PDBs: 7yx4
Q-score: 0.481
Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C
eLife (2022) 11 [ Pubmed: 35900198 DOI: doi:10.7554/eLife.77607 ]
Structure of the Mimivirus genomic fibre in its relaxed 5-start helix form
Resolution: 3.7 Å
EM Method: Helical reconstruction
Fitted PDBs: 7yx5
Q-score: 0.468
Villalta A, Schmitt A, Estrozi LF, Quemin ERJ, Alempic JM, Lartigue A, Prazak V, Belmudes L, Vasishtan D, Colmant AMG, Honore FA, Coute Y, Grunewald K, Abergel C
eLife (2022) 11 [ Pubmed: 35900198 DOI: doi:10.7554/eLife.77607 ]