Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
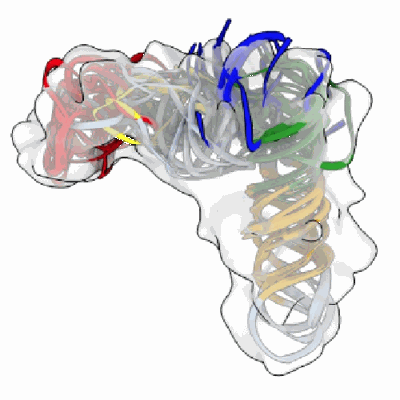
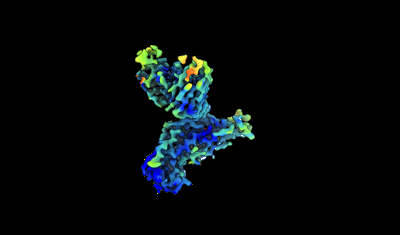
Model and map from local refinement of a CAB-A17 - Omicron Ba.1 spike complex
Resolution: 2.68 Å
EM Method: Single-particle
Fitted PDBs: 8v4f
Q-score: 0.513
Sheward DJ, Pushparaj P, Das H, Greaney AJ, Kim C, Kim S, Hanke L, Hyllner E, Dyrdak R, Lee J, Dopico XC, Dosenovic P, Peacock TP, McInerney GM, Albert J, Corcoran M, Bloom JD, Murrell B, Karlsson Hedestam GB, Hallberg BM
Cell Rep Med (2024) pp. 101577-101577 [ Pubmed: 38761799 DOI: doi:10.1016/j.xcrm.2024.101577 ]
- Complex between fab preparation of cab-a17 and omicron ba.1 spike (Complex from Homo sapiens)
- Spike protein s1 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Cab-a17 variable heavy-chain (12 kDa, Protein from Homo sapiens)
- Cab-a17 variable light chain (11 kDa, Protein from Homo sapiens)
The interface structure of Omicron RBD binding to 5817 Fab
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 8khd
Q-score: 0.393
Wang Y, Zhang Z, Yang M, Xiong X, Yan Q, Cao L, Wei P, Zhang Y, Zhang L, Lv K, Chen J, Liu X, Zhao X, Xiao J, Zhang S, Zhu A, Gan M, Zhang J, Cai R, Zhuo J, Zhang Y, Rao H, Qu B, Zhang Y, Chen L, Dai J, Cheng L, Hu Q, Chen Y, Lv H, So RTY, Peiris M, Zhao J, Liu X, Mok CKP, Wang X, Zhao J
Cell Rep (2024) 43 pp. 113653-113653 [ DOI: doi:10.1016/j.celrep.2023.113653 Pubmed: 38175758 ]
- The interface structure of omicron rbd binding to 5817 fab (Complex from Severe acute respiratory syndrome coronavirus 2)
- Light chain of 5817 (11 kDa, Protein from Homo sapiens)
- Heavy chain of 5817 (13 kDa, Protein from Homo sapiens)
- Spike protein s1 (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
de Moura TR, Purta E, Bernat A, Martin-Cuevas EM, Kurkowska M, Baulin EF, Mukherjee S, Nowak J, Biela AP, Rawski M, Glatt S, Moreno-Herrero F, Bujnicki JM
Nucleic Acids Res (2024) 52 pp. 3419-3432 [ DOI: doi:10.1093/nar/gkae144 Pubmed: 38426934 ]
XBB.1.5 RBD in complex with BD55-1205
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8xe9
Q-score: 0.473
To Be Published
- Xbb.1.5 rbd (Complex from Severe acute respiratory syndrome coronavirus 2)
- Spike protein s2' (22 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Bd55-1205 heavy chain (12 kDa, Protein from Homo sapiens)
- Bd55-1205 (Complex from Homo sapiens)
- Xbb.1.5 rbd in complex with bd55-1205 (Complex)
- Bd55-1205 light chain (11 kDa, Protein from Homo sapiens)
SARS-CoV-2 Frameshift Stimulatory Element with Upstream Multibranch Loop
Peterson JM, Becker ST, O'Leary CA, Juneja P, Yang Y, Moss WN
Biochemistry (2024) 63 pp. 1287-1296 [ Pubmed: 38727003 DOI: doi:10.1021/acs.biochem.3c00716 ]
SARS-CoV-2 5' proximal stem-loop 5
Kretsch RC, Xu L, Zheludev IN, Zhou X, Huang R, Nye G, Li S, Zhang K, Chiu W, Das R
PNAS (2024) 121 pp. e2320493121-e2320493121 [ Pubmed: 38427602 DOI: doi:10.1073/pnas.2320493121 ]
Local refined cryo-EM reconstruction of Omicron BA.5 RBD in complex with 8-9D Fab
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8j1t
Q-score: 0.455
Tai W, Yang K, Liu Y, Li R, Feng S, Chai B, Zhuang X, Qi S, Shi H, Liu Z, Lei J, Ma E, Wang W, Tian C, Le T, Wang J, Chen Y, Tian M, Xiang Y, Yu G, Cheng G
Nat Commun (2023) 14 pp. 8042-8042 [ DOI: doi:10.1038/s41467-023-43798-8 Pubmed: 38052844 ]
- 8-9d fab (Complex from Homo sapiens)
- Severe acute respiratory syndrome coronavirus 2 omicron ba.5 rbd bound 8-9d fab (22 kDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- 8-9d light chain (11 kDa, Protein from Homo sapiens)
- Sars-cov2 omicron ba.5 rbd (Complex)
- 8-9d heavy chain (12 kDa, Protein from Homo sapiens)
- Spike protein (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
Cryo-EM structure of SARS-CoV-2 BA.4/5 spike protein in complex with 1G11 (local refinement)
Resolution: 3.98 Å
EM Method: Single-particle
Fitted PDBs: 8ix3
Q-score: 0.451
Sun H, Wang Y, Chen X, Jiang Y, Wang S, Huang Y, Liu L, Li Y, Lan M, Guo H, Yuan Q, Zhang Y, Li T, Yu H, Gu Y, Zhang J, Li S, Zheng Z, Zheng Q, Xia N
J Virol (2023) 97 pp. e0113723-e0113723 [ DOI: doi:10.1128/jvi.01137-23 Pubmed: 37855619 ]
- Heavy chain of 1g11 (13 kDa, Protein from Homo sapiens)
- Light chain of 1g11 (11 kDa, Protein from Homo sapiens)
- Nab 1g11 (Complex from Homo sapiens)
- Ba.4/5 variant spike protein (23 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 ba.4/5 spike protein (Complex from Severe acute respiratory syndrome coronavirus 2)
- Sars-cov-2 ba.4/5 spike protein in complex with 1g11 (Complex)
XBB 1.0 RBD bound to P4J15 (Local)
Resolution: 3.85 Å
EM Method: Single-particle
Fitted PDBs: 8pq2
Q-score: 0.481
Fenwick C, Turelli P, Duhoo Y, Lau K, Herate C, Marlin R, Lamrayah M, Campos J, Esteves-Leuenberger L, Farina A, Raclot C, Genet V, Fiscalini F, Cesborn J, Perez L, Dereuddre-Bosquet N, Contreras V, Lheureux K, Relouzat F, Abdelnabi R, Leyssen P, Levy Y, Pojer F, Le Grand R, Trono D, Pantaleo G
J Infect (2023) 87 pp. 524-537 [ DOI: doi:10.1016/j.jinf.2023.10.008 Pubmed: 37852477 ]
- Spike xbb.1 rbd up - p4j15 fab (local) (72 MDa, Complex from Severe acute respiratory syndrome coronavirus 2)
- Spike protein s2' (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- P4j15 fragment antigen-binding heavy chain (13 kDa, Protein from Homo sapiens)
- P4j15 fragment antigen-binding light chain (11 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
cryoEM structure of a broadly neutralizing anti-SARS-CoV-2 antibody STI-9167
Resolution: 3.16 Å
EM Method: Single-particle
Fitted PDBs: 8eqf
Q-score: 0.518
Atanasoff KE, Brambilla L, Adelsberg DC, Kowdle S, Stevens CS, Slamanig S, Hung C-T, Fu Y, Lim R, Tran L, Allen R, Sun W, Duty JA, Bajic G, Lee B, Tortorella D
Mbio (2024) 15 pp. e0247723-e0247723 [ Pubmed: 38054729 DOI: doi:10.1128/mbio.02477-23 ]
- Sars-cov-2 spike rbd in complex with 10a3 fab (Complex from Homo sapiens)
- Fab light chain (11 kDa, Protein from Homo sapiens)
- Fab heavy chain (13 kDa, Protein from Homo sapiens)
- Spike protein s1 (21 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)