 |
PDBsum entry 5tl7

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Signaling protein/hydrolase
|
PDB id
|
|

|

|
5tl7
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
|

|
|
PDB id:
|
 |
|
 |
| Name: |
 |
Signaling protein/hydrolase
|
 |
|
Title:
|
 |
Crystal structure of sars-cov papain-like protease in complex with c- terminal domain mouse isg15
|
|
Structure:
|
 |
Ubiquitin-like protein isg15. Chain: a, c. Fragment: c-terminal domain (unp residues 78-155. Synonym: interferon-induced 15 kda protein,interferon-induced 17 kda protein,ip17,ubiquitin cross-reactive protein. Engineered: yes. Replicase polyprotein 1ab. Chain: b, d. Fragment: unp residues 1541-1855.
|
|
Source:
|
 |
Mus musculus. Mouse. Organism_taxid: 10090. Gene: isg15, g1p2, ucrp. Expressed in: escherichia coli bl21(de3). Expression_system_taxid: 469008. Sars coronavirus. Sars-cov. Organism_taxid: 227859.
|
|
Resolution:
|
 |
|
2.44Å
|
R-factor:
|
0.196
|
R-free:
|
0.267
|
|
|
Authors:
|
 |
C.D.Daczkowski,J.V.Dzimianski,S.D.Pegan
|
|
Key ref:
|
 |
C.M.Daczkowski
et al.
(2017).
Structural Insights into the Interaction of Coronavirus Papain-Like Proteases and Interferon-Stimulated Gene Product 15 from Different Species.
J Mol Biol,
429,
1661-1683.
PubMed id:
|
 |
|
Date:
|
 |
|
10-Oct-16
|
Release date:
|
03-May-17
|
|
|

|

|
|
PROCHECK
|

|

|
|
|
Headers
|
 |
|
|
References
|
|
|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 2:
|
 |
Chains B, D:
E.C.2.1.1.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
Chains B, D:
E.C.2.1.1.56
- mRNA (guanine-N(7))-methyltransferase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L- methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-homocysteine
|
 |
 |
 |
 |
 |
5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L- methionine
S-adenosyl-L- methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
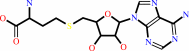 S-adenosyl-L-homocysteine
S-adenosyl-L-homocysteine
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 4:
|
 |
Chains B, D:
E.C.2.1.1.57
- methyltransferase cap1.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA + S-adenosyl-L-homocysteine + H+
|
 |
 |
 |
 |
 |
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L-methionine
S-adenosyl-L-methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA
|
+
|
 S-adenosyl-L-homocysteine
S-adenosyl-L-homocysteine
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 5:
|
 |
Chains B, D:
E.C.2.7.7.48
- RNA-directed Rna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
RNA(n) + a ribonucleoside 5'-triphosphate = RNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
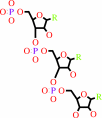 RNA(n)
RNA(n)
|
+
|
ribonucleoside 5'-triphosphate
|
=
|
RNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 6:
|
 |
Chains B, D:
E.C.2.7.7.50
- mRNA guanylyltransferase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end diphospho-ribonucleoside in mRNA + GTP + H+ = a 5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA + diphosphate
|
 |
 |
 |
 |
 |
5'-end diphospho-ribonucleoside in mRNA
|
+
|
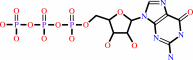 GTP
GTP
|
+
|
H(+)
|
=
|
5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 7:
|
 |
Chains B, D:
E.C.3.1.13.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 8:
|
 |
Chains B, D:
E.C.3.4.19.12
- ubiquitinyl hydrolase 1.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Thiol-dependent hydrolysis of ester, thiolester, amide, peptide and isopeptide bonds formed by the C-terminal Gly of ubiquitin (a 76-residue protein attached to proteins as an intracellular targeting signal).
|
 |
 |
 |
 |
 |
Enzyme class 9:
|
 |
Chains B, D:
E.C.3.4.22.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 10:
|
 |
Chains B, D:
E.C.3.4.22.69
- Sars coronavirus main proteinase.
|
|
 |
 |
 |
 |
 |
Enzyme class 11:
|
 |
Chains B, D:
E.C.3.6.4.12
- Dna helicase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
ATP + H2O = ADP + phosphate + H+
|
 |
 |
 |
 |
 |
 ATP
ATP
|
+
|
H2O
|
=
|
 ADP
ADP
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 12:
|
 |
Chains B, D:
E.C.3.6.4.13
- Rna helicase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
ATP + H2O = ADP + phosphate + H+
|
 |
 |
 |
 |
 |
 ATP
ATP
|
+
|
H2O
|
=
|
 ADP
ADP
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 13:
|
 |
Chains B, D:
E.C.4.6.1.-
- ?????
|
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
|
J Mol Biol
429:1661-1683
(2017)
|
|
PubMed id:
|
|
|
|

|
| |
|
Structural Insights into the Interaction of Coronavirus Papain-Like Proteases and Interferon-Stimulated Gene Product 15 from Different Species.
|
|
C.M.Daczkowski,
J.V.Dzimianski,
J.R.Clasman,
O.Goodwin,
A.D.Mesecar,
S.D.Pegan.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
Severe acute respiratory syndrome coronavirus (SARS-CoV) and Middle East
respiratory syndrome coronavirus (MERS-CoV) encode multifunctional papain-like
proteases (PLPs) that have the ability to process the viral polyprotein to
facilitate RNA replication and antagonize the host innate immune response. The
latter function involves reversing the post-translational modification of
cellular proteins conjugated with either ubiquitin (Ub) or Ub-like
interferon-stimulated gene product 15 (ISG15). Ub is known to be highly
conserved among eukaryotes, but surprisingly, ISG15 is highly divergent among
animals. The ramifications of this sequence divergence to the recognition of
ISG15 by coronavirus PLPs at a structural and biochemical level are poorly
understood. Therefore, the activity of PLPs from SARS-CoV, MERS-CoV, and mouse
hepatitis virus was evaluated against seven ISG15s originating from an
assortment of animal species susceptible, and not, to certain coronavirus
infections. Excitingly, our kinetic, thermodynamic, and structural analysis
revealed an array of different preferences among PLPs. Included in these studies
is the first insight into a coronavirus PLP's interface with ISG15 via SARS-CoV
PLpro in complex with the principle binding domain of human ISG15 (hISG15) and
mouse ISG15s (mISG15s). The first X-ray structure of the full-length mISG15
protein is also reported and highlights a unique, twisted hinge region of ISG15
that is not conserved in hISG15, suggesting a potential role in differential
recognition. Taken together, this new information provides a structural and
biochemical understanding of the distinct specificities among coronavirus PLPs
observed and addresses a critical gap of how PLPs can interact with ISG15s from
a wide variety of species.

|

|
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
|

