 |
PDBsum entry 4wy3

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Hydrolase/hydrolase inhibitor
|
PDB id
|
|

|

|
4wy3
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
|

|
|
PDB id:
|
 |
|
 |
| Name: |
 |
Hydrolase/hydrolase inhibitor
|
 |
|
Title:
|
 |
Structure of sars-3cl protease complex with a phenylbenzoyl (r,s)-n- decalin type inhibitor
|
|
Structure:
|
 |
3c-like proteinase. Chain: a. Synonym: 3cl-pro, 3clp, nsp5. Engineered: yes
|
|
Source:
|
 |
Sars coronavirus. Sars-cov. Organism_taxid: 227859. Gene: rep, 1a-1b. Expressed in: escherichia coli. Expression_system_taxid: 469008.
|
|
Resolution:
|
 |
|
1.89Å
|
R-factor:
|
0.275
|
R-free:
|
0.299
|
|
|
Authors:
|
 |
K.Akaji,K.Teruya,Y.Shimamoto,A.Sanjho,E.Yamashita,A.Nakagawa
|
|
Key ref:
|
 |
Y.Shimamoto
et al.
(2015).
Fused-ring structure of decahydroisoquinolin as a novel scaffold for SARS 3CL protease inhibitors.
Bioorg Med Chem Lett,
23,
876-890.
PubMed id:
DOI:
|
 |
|
Date:
|
 |
|
15-Nov-14
|
Release date:
|
18-Feb-15
|
|
|
Supersedes:
|

|
|
|

|

|
|
PROCHECK
|

|

|
|
|
Headers
|
 |
|
|
References
|
|
|
|

|

|
|
P0C6X7
(R1AB_CVHSA) -
Replicase polyprotein 1ab from Severe acute respiratory syndrome coronavirus
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
7073 a.a.
306 a.a.*
|

|

|

|

|
|

|

|

|
 |
 |

|
|
Key: |
 |
PfamA domain |
 |
 |
 |
Secondary structure |
 |
 |
CATH domain |
 |
|
*
PDB and UniProt seqs differ
at 1 residue position (black
cross)
|
|

|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 2:
|
 |
E.C.2.1.1.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
E.C.2.1.1.56
- mRNA (guanine-N(7))-methyltransferase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L- methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-homocysteine
|
 |
 |
 |
 |
 |
5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L- methionine
S-adenosyl-L- methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
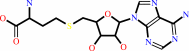 S-adenosyl-L-homocysteine
S-adenosyl-L-homocysteine
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 4:
|
 |
E.C.2.1.1.57
- methyltransferase cap1.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA + S-adenosyl-L-homocysteine + H+
|
 |
 |
 |
 |
 |
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L-methionine
S-adenosyl-L-methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA
|
+
|
 S-adenosyl-L-homocysteine
S-adenosyl-L-homocysteine
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 5:
|
 |
E.C.2.7.7.48
- RNA-directed Rna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
RNA(n) + a ribonucleoside 5'-triphosphate = RNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
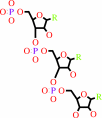 RNA(n)
RNA(n)
|
+
|
ribonucleoside 5'-triphosphate
|
=
|
RNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 6:
|
 |
E.C.2.7.7.50
- mRNA guanylyltransferase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end diphospho-ribonucleoside in mRNA + GTP + H+ = a 5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA + diphosphate
|
 |
 |
 |
 |
 |
5'-end diphospho-ribonucleoside in mRNA
|
+
|
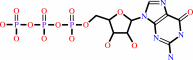 GTP
GTP
|
+
|
H(+)
|
=
|
5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 7:
|
 |
E.C.3.1.13.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 8:
|
 |
E.C.3.4.19.12
- ubiquitinyl hydrolase 1.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Thiol-dependent hydrolysis of ester, thiolester, amide, peptide and isopeptide bonds formed by the C-terminal Gly of ubiquitin (a 76-residue protein attached to proteins as an intracellular targeting signal).
|
 |
 |
 |
 |
 |
Enzyme class 9:
|
 |
E.C.3.4.22.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 10:
|
 |
E.C.3.4.22.69
- Sars coronavirus main proteinase.
|
|
 |
 |
 |
 |
 |
Enzyme class 11:
|
 |
E.C.3.6.4.12
- Dna helicase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
ATP + H2O = ADP + phosphate + H+
|
 |
 |
 |
 |
 |
 ATP
ATP
|
+
|
H2O
|
=
|
 ADP
ADP
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 12:
|
 |
E.C.3.6.4.13
- Rna helicase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
ATP + H2O = ADP + phosphate + H+
|
 |
 |
 |
 |
 |
 ATP
ATP
|
+
|
H2O
|
=
|
 ADP
ADP
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 13:
|
 |
E.C.4.6.1.-
- ?????
|
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
DOI no:
|
Bioorg Med Chem Lett
23:876-890
(2015)
|
|
PubMed id:
|
|
|
|

|
| |
|
Fused-ring structure of decahydroisoquinolin as a novel scaffold for SARS 3CL protease inhibitors.
|
|
Y.Shimamoto,
Y.Hattori,
K.Kobayashi,
K.Teruya,
A.Sanjoh,
A.Nakagawa,
E.Yamashita,
K.Akaji.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
The design and evaluation of a novel decahydroisoquinolin scaffold as an
inhibitor for severe acute respiratory syndrome (SARS) chymotrypsin-like
protease (3CL(pro)) are described. Focusing on hydrophobic interactions at the
S2 site, the decahydroisoquinolin scaffold was designed by connecting the P2
site cyclohexyl group of the substrate-based inhibitor to the main-chain at the
α-nitrogen atom of the P2 position via a methylene linker. Starting from a
cyclohexene enantiomer obtained by salt resolution, trans-decahydroisoquinolin
derivatives were synthesized. All decahydroisoquinolin inhibitors synthesized
showed moderate but clear inhibitory activities for SARS 3CL(pro), which
confirmed the fused ring structure of the decahydroisoquinolin functions as a
novel scaffold for SARS 3CL(pro) inhibitor. X-ray crystallographic analyses of
the SARS 3CL(pro) in a complex with the decahydroisoquinolin inhibitor revealed
the expected interactions at the S1 and S2 sites, as well as additional
interactions at the N-substituent of the inhibitor.

|

|
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

