 |
PDBsum entry 3ebn

|

|

|
|
 |
Contents |
 |
|
|

|

|
|
|
|
|
|
* Residue conservation analysis
|
|
|

|
|
PDB id:
|
 |
|
 |
| Name: |
 |
Hydrolase
|
 |
|
Title:
|
 |
A special dimerization of sars-cov main proteasE C-terminal domain due to domain-swapping
|
|
Structure:
|
 |
Replicase polyprotein 1ab. Chain: a, b, c, d. Fragment: unp residues 3429-3546. Synonym: pp1ab, orf1ab polyprotein, 3c-like proteinase, 3cl-pro, 3clp, nsp5. Engineered: yes
|
|
Source:
|
 |
Sars coronavirus. Sars-cov. Organism_taxid: 227859. Gene: rep, 1a-1b. Expressed in: escherichia coli. Expression_system_taxid: 562.
|
|
Resolution:
|
 |
|
2.40Å
|
R-factor:
|
0.208
|
R-free:
|
0.243
|
|
|
Authors:
|
 |
N.Zhong,S.Zhang,F.Xue,X.Kang,Z.Lou,B.Xia
|
|
Key ref:
|
 |
N.Zhong
et al.
(2009).
C-terminal domain of SARS-CoV main protease can form a 3D domain-swapped dimer.
Protein Sci,
18,
839-844.
PubMed id:
|
 |
|
Date:
|
 |
|
28-Aug-08
|
Release date:
|
19-May-09
|
|
|

|

|
|
PROCHECK
|

|

|
|
|
Headers
|
 |
|
|
References
|
|
|
|

|

|
|
P0C6X7
(R1AB_CVHSA) -
Replicase polyprotein 1ab from Severe acute respiratory syndrome coronavirus
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
7073 a.a.
100 a.a.
|

|

|

|

|
|

|

|

|
 |
 |

|
|
Key: |
 |
PfamA domain |
 |
 |
 |
Secondary structure |
 |
 |
CATH domain |
 |
|

|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 2:
|
 |
E.C.2.1.1.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
E.C.2.1.1.56
- mRNA (guanine-N(7))-methyltransferase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L- methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-homocysteine
|
 |
 |
 |
 |
 |
5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L- methionine
S-adenosyl-L- methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
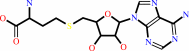 S-adenosyl-L-homocysteine
S-adenosyl-L-homocysteine
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 4:
|
 |
E.C.2.1.1.57
- methyltransferase cap1.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA + S-adenosyl-L-homocysteine + H+
|
 |
 |
 |
 |
 |
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L-methionine
S-adenosyl-L-methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA
|
+
|
 S-adenosyl-L-homocysteine
S-adenosyl-L-homocysteine
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 5:
|
 |
E.C.2.7.7.48
- RNA-directed Rna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
RNA(n) + a ribonucleoside 5'-triphosphate = RNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
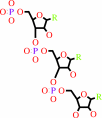 RNA(n)
RNA(n)
|
+
|
ribonucleoside 5'-triphosphate
|
=
|
RNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 6:
|
 |
E.C.2.7.7.50
- mRNA guanylyltransferase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end diphospho-ribonucleoside in mRNA + GTP + H+ = a 5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA + diphosphate
|
 |
 |
 |
 |
 |
5'-end diphospho-ribonucleoside in mRNA
|
+
|
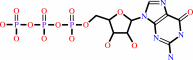 GTP
GTP
|
+
|
H(+)
|
=
|
5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 7:
|
 |
E.C.3.1.13.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 8:
|
 |
E.C.3.4.19.12
- ubiquitinyl hydrolase 1.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Thiol-dependent hydrolysis of ester, thiolester, amide, peptide and isopeptide bonds formed by the C-terminal Gly of ubiquitin (a 76-residue protein attached to proteins as an intracellular targeting signal).
|
 |
 |
 |
 |
 |
Enzyme class 9:
|
 |
E.C.3.4.22.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 10:
|
 |
E.C.3.4.22.69
- Sars coronavirus main proteinase.
|
|
 |
 |
 |
 |
 |
Enzyme class 11:
|
 |
E.C.3.6.4.12
- Dna helicase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
ATP + H2O = ADP + phosphate + H+
|
 |
 |
 |
 |
 |
 ATP
ATP
|
+
|
H2O
|
=
|
 ADP
ADP
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 12:
|
 |
E.C.3.6.4.13
- Rna helicase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
ATP + H2O = ADP + phosphate + H+
|
 |
 |
 |
 |
 |
 ATP
ATP
|
+
|
H2O
|
=
|
 ADP
ADP
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 13:
|
 |
E.C.4.6.1.-
- ?????
|
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
|
Protein Sci
18:839-844
(2009)
|
|
PubMed id:
|
|
|
|

|
| |
|
C-terminal domain of SARS-CoV main protease can form a 3D domain-swapped dimer.
|
|
N.Zhong,
S.Zhang,
F.Xue,
X.Kang,
P.Zou,
J.Chen,
C.Liang,
Z.Rao,
C.Jin,
Z.Lou,
B.Xia.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
SARS coronavirus main protease (M(pro)) plays an essential role in the extensive
proteolytic processing of the viral polyproteins (pp1a and pp1ab), and it is an
important target for anti-SARS drug development. We have reported that both the
M(pro) C-terminal domain alone (M(pro)-C) and the N-finger deletion mutant of
M(pro) (M(pro)-Delta7) exist as a stable dimer and a stable monomer (Zhong et
al., J Virol 2008; 82:4227-4234). Here, we report structures of both M(pro)-C
monomer and dimer. The structure of the M(pro)-C monomer is almost identical to
that of the C-terminal domain in the crystal structure of M(pro). Interestingly,
the M(pro)-C dimer structure is characterized by 3D domain-swapping, in which
the first helices of the two protomers are interchanged and each is enwrapped by
four other helices from the other protomer. Each folding subunit of the M(pro)-C
domain-swapped dimer still has the same general fold as that of the M(pro)-C
monomer. This special dimerization elucidates the structural basis for the
observation that there is no exchange between monomeric and dimeric forms of
M(pro)-C and M(pro)-Delta7.

|

|
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Literature references that cite this PDB file's key reference
|
|
 |
| |
PubMed id
|
 |
Reference
|
 |
|
|
|
 |
J.Shi,
N.Han,
L.Lim,
S.Lua,
J.Sivaraman,
L.Wang,
Y.Mu,
and
J.Song
(2011).
Dynamically-driven inactivation of the catalytic machinery of the SARS 3C-like protease by the N214A mutation on the extra domain.
|
| |
PLoS Comput Biol,
7,
e1001084.
|
 |
|
|
|
|
 |
M.Y.Tsai,
W.H.Chang,
J.Y.Liang,
L.L.Lin,
G.G.Chang,
and
H.P.Chang
(2010).
Essential covalent linkage between the chymotrypsin-like domain and the extra domain of the SARS-CoV main protease.
|
| |
J Biochem,
148,
349-358.
|
 |
|
|
|
|
 |
S.C.Cheng,
G.G.Chang,
and
C.Y.Chou
(2010).
Mutation of Glu-166 blocks the substrate-induced dimerization of SARS coronavirus main protease.
|
| |
Biophys J,
98,
1327-1336.
|
 |
|
 |
 |
|
The most recent references are shown first.
Citation data come partly from CiteXplore and partly
from an automated harvesting procedure. Note that this is likely to be
only a partial list as not all journals are covered by
either method. However, we are continually building up the citation data
so more and more references will be included with time.
|
 |

