 |
PDBsum entry 2z3d

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Hydrolase/hydrolase inhibitor
|
PDB id
|
|

|

|
2z3d
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
 |
Contents |
 |
|
|

|

|
|
|
|
|
|
|
|
* Residue conservation analysis
|
|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 2:
|
 |
E.C.2.1.1.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
E.C.2.1.1.56
- mRNA (guanine-N(7))-methyltransferase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L- methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-homocysteine
|
 |
 |
 |
 |
 |
5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L- methionine
S-adenosyl-L- methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
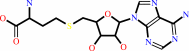 S-adenosyl-L-homocysteine
S-adenosyl-L-homocysteine
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 4:
|
 |
E.C.2.1.1.57
- methyltransferase cap1.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA + S-adenosyl-L-homocysteine + H+
|
 |
 |
 |
 |
 |
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L-methionine
S-adenosyl-L-methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA
|
+
|
 S-adenosyl-L-homocysteine
S-adenosyl-L-homocysteine
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 5:
|
 |
E.C.2.7.7.48
- RNA-directed Rna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
RNA(n) + a ribonucleoside 5'-triphosphate = RNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
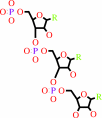 RNA(n)
RNA(n)
|
+
|
ribonucleoside 5'-triphosphate
|
=
|
RNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 6:
|
 |
E.C.2.7.7.50
- mRNA guanylyltransferase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end diphospho-ribonucleoside in mRNA + GTP + H+ = a 5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA + diphosphate
|
 |
 |
 |
 |
 |
5'-end diphospho-ribonucleoside in mRNA
|
+
|
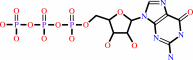 GTP
GTP
|
+
|
H(+)
|
=
|
5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 7:
|
 |
E.C.3.1.13.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 8:
|
 |
E.C.3.4.19.12
- ubiquitinyl hydrolase 1.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Thiol-dependent hydrolysis of ester, thiolester, amide, peptide and isopeptide bonds formed by the C-terminal Gly of ubiquitin (a 76-residue protein attached to proteins as an intracellular targeting signal).
|
 |
 |
 |
 |
 |
Enzyme class 9:
|
 |
E.C.3.4.22.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 10:
|
 |
E.C.3.4.22.69
- Sars coronavirus main proteinase.
|
|
 |
 |
 |
 |
 |
Enzyme class 11:
|
 |
E.C.3.6.4.12
- Dna helicase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
ATP + H2O = ADP + phosphate + H+
|
 |
 |
 |
 |
 |
 ATP
ATP
|
+
|
H2O
|
=
|
 ADP
ADP
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 12:
|
 |
E.C.3.6.4.13
- Rna helicase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
ATP + H2O = ADP + phosphate + H+
|
 |
 |
 |
 |
 |
 ATP
ATP
|
+
|
H2O
|
=
|
 ADP
ADP
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 13:
|
 |
E.C.4.6.1.-
- ?????
|
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
DOI no:
|
J Mol Biol
371:1060-1074
(2007)
|
|
PubMed id:
|
|
|
|

|
| |
|
A mechanistic view of enzyme inhibition and peptide hydrolysis in the active site of the SARS-CoV 3C-like peptidase.
|
|
J.Yin,
C.Niu,
M.M.Cherney,
J.Zhang,
C.Huitema,
L.D.Eltis,
J.C.Vederas,
M.N.James.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
The 3C-like main peptidase 3CL(pro) is a viral polyprotein processing enzyme
essential for the viability of the Severe Acute Respiratory Syndrome coronavirus
(SARS-CoV). While it is generalized that 3CL(pro) and the structurally related
3C(pro) viral peptidases cleave their substrates via a mechanism similar to that
underlying the peptide hydrolysis by chymotrypsin-like serine proteinases
(CLSPs), some of the hypothesized key intermediates have not been structurally
characterized. Here, we present three crystal structures of SARS 3CL(pro) in
complex with each of two members of a new class of peptide-based phthalhydrazide
inhibitors. Both inhibitors form an unusual thiiranium ring with the
nucleophilic sulfur atom of Cys145, trapping the enzyme's catalytic residues in
configurations similar to the intermediate states proposed to exist during the
hydrolysis of native substrates. Most significantly, our crystallographic data
are consistent with a scenario in which a water molecule, possibly via indirect
coordination from the carbonyl oxygen of Thr26, has initiated nucleophilic
attack on the enzyme-bound inhibitor. Our data suggest that this structure
resembles that of the proposed tetrahedral intermediate during the deacylation
step of normal peptidyl cleavage.

|

|
|
|

|
| |
Selected figure(s)
|
|

|
| |
 |
 |

|
 |
Figure 1.
Figure 1. The chemical formulae for the inhibitors used in
this study. Chiral centers are indicated where necessary.
|
 |
|
|

|
| |
The above figure is
reprinted
by permission from Elsevier:
J Mol Biol
(2007,
371,
1060-1074)
copyright 2007.
|
|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Literature references that cite this PDB file's key reference
|
|
 |
| |
PubMed id
|
 |
Reference
|
 |
|
|
|
 |
M.Shokhen,
N.Khazanov,
and
A.Albeck
(2011).
The mechanism of papain inhibition by peptidyl aldehydes.
|
| |
Proteins,
79,
975-985.
|
 |
|
|
|
|
 |
C.Niu,
J.Yin,
J.Zhang,
J.C.Vederas,
and
M.N.James
(2008).
Molecular docking identifies the binding of 3-chloropyridine moieties specifically to the S1 pocket of SARS-CoV Mpro.
|
| |
Bioorg Med Chem,
16,
293-302.
|
 |
|
 |
 |
|
The most recent references are shown first.
Citation data come partly from CiteXplore and partly
from an automated harvesting procedure. Note that this is likely to be
only a partial list as not all journals are covered by
either method. However, we are continually building up the citation data
so more and more references will be included with time.
|
 |

