Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
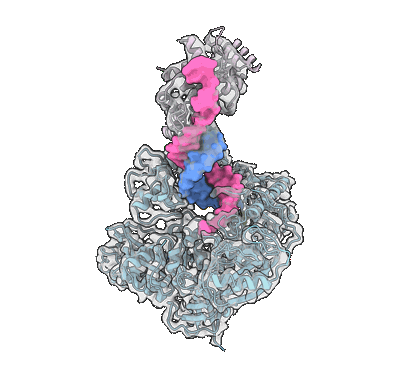
Human DNA polymerase alpha/primase elongation complex I bound to primer/template
Resolution: 3.35 Å
EM Method: Single-particle
Fitted PDBs: 8d96
Q-score: 0.426
He Q, Baranovskiy AG, Morstadt LM, Lisova AE, Babayeva ND, Lusk BL, Lim CJ, Tahirov TH
Nat Struct Mol Biol (2023) 30 pp. 579-583 [ DOI: doi:10.1038/s41594-023-00971-3 Pubmed: 37069376 ]
- Dna (5'-d(*ap*tp*ap*ap*tp*gp*gp*tp*cp*gp*tp*gp*cp*cp*gp*cp*cp*ap*ap*tp*ap*a)-3') (6 kDa, DNA from Homo sapiens)
- Dna primase large subunit (58 kDa, Protein from Homo sapiens)
- Dna/rna (5'-gtp)-r(p*gp*cp*gp*gp*cp*ap*cp*g)-d(p*ap*cp*c)-3') (3 kDa, Macromolecule from Homo sapiens)
- Iron/sulfur cluster (351 Da, Ligand)
- 2'-deoxyadenosine 5'-triphosphate (491 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Dna polymerase alpha catalytic subunit (166 kDa, Protein from Homo sapiens)
- Elongation complex i of human dna polymerase alpha/primase bound to rna-dna primer and dna template (340 kDa, Complex from Homo sapiens)
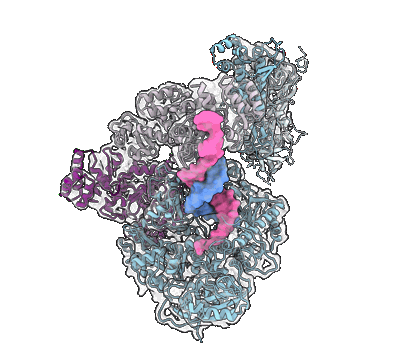
Human DNA polymerase-alpha/primase elongation complex II bound to primer/template
Resolution: 3.59 Å
EM Method: Single-particle
Fitted PDBs: 8d9d
Q-score: 0.347
He Q, Baranovskiy AG, Morstadt LM, Lisova AE, Babayeva ND, Lusk BL, Lim CJ, Tahirov TH
Nat Struct Mol Biol (2023) 30 pp. 579-583 [ DOI: doi:10.1038/s41594-023-00971-3 Pubmed: 37069376 ]
- Dna primase small subunit (49 kDa, Protein from Homo sapiens)
- Dna primase large subunit (58 kDa, Protein from Homo sapiens)
- Dna (5'-d(*ap*tp*gp*gp*tp*cp*gp*tp*gp*cp*cp*gp*cp*cp*ap*ap*tp*ap*a)-3') (5 kDa, DNA from Homo sapiens)
- Iron/sulfur cluster (351 Da, Ligand)
- Dna/rna (5'-d(*(gtp))-r(p*gp*cp*gp*gp*cp*ap*cp*g)-d(p*ap*cp*c)-3') (3 kDa, Macromolecule from Homo sapiens)
- Zinc ion (65 Da, Ligand)
- Elongation complex ii of human dna polymerase alpha/primase bound to rna-dna primer and dna template (340 kDa, Complex from Homo sapiens)
- 2'-deoxyadenosine 5'-triphosphate (491 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Dna polymerase alpha catalytic subunit (166 kDa, Protein from Homo sapiens)
- Dna polymerase alpha subunit b (49 kDa, Protein from Homo sapiens)
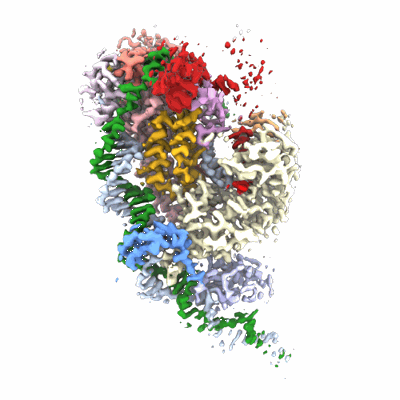
Partial auto-inhibitory complex of Xenopus laevis DNA polymerase alpha-primase
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 8g99
Q-score: 0.551
Mullins EA, Salay LE, Durie CL, Bradley NP, Jackman JE, Ohi MD, Chazin WJ, Eichman BF
Nat Struct Mol Biol (2024) [ Pubmed: 38491139 DOI: doi:10.1038/s41594-024-01227-4 ]
- Dna polymerase alpha catalytic subunit (129 kDa, Protein from Xenopus laevis)
- Dna primase large subunit (59 kDa, Protein from Xenopus laevis)
- Dna polymerase alpha subunit b (67 kDa, Protein from Xenopus laevis)
- Primase (Complex from Xenopus laevis)
- Polymerase alpha (200 kDa, Complex from Xenopus laevis)
- Polymerase alpha-primase (110 kDa, Complex)
- Zinc ion (65 Da, Ligand)
- Iron/sulfur cluster (351 Da, Ligand)
Complete auto-inhibitory complex of Xenopus laevis DNA polymerase alpha-primase
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 8g9f
Q-score: 0.48
Mullins EA, Salay LE, Durie CL, Bradley NP, Jackman JE, Ohi MD, Chazin WJ, Eichman BF
Nat Struct Mol Biol (2024) [ Pubmed: 38491139 DOI: doi:10.1038/s41594-024-01227-4 ]
- Dna primase (49 kDa, Protein from Xenopus laevis)
- Dna polymerase alpha catalytic subunit (129 kDa, Protein from Xenopus laevis)
- Dna primase large subunit (59 kDa, Protein from Xenopus laevis)
- Dna polymerase alpha subunit b (67 kDa, Protein from Xenopus laevis)
- Primase (Complex from Xenopus laevis)
- Polymerase alpha (200 kDa, Complex from Xenopus laevis)
- Polymerase alpha-primase (110 kDa, Complex)
- Zinc ion (65 Da, Ligand)
- Iron/sulfur cluster (351 Da, Ligand)
DNA initiation subcomplex of Xenopus laevis DNA polymerase alpha-primase
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8g9l
Q-score: 0.453
Mullins EA, Salay LE, Durie CL, Bradley NP, Jackman JE, Ohi MD, Chazin WJ, Eichman BF
Nat Struct Mol Biol (2024) [ Pubmed: 38491139 DOI: doi:10.1038/s41594-024-01227-4 ]
- Dna template (15 kDa, DNA from synthetic construct)
- Dna polymerase alpha catalytic subunit (129 kDa, Protein from Xenopus laevis)
- Dna primase large subunit (59 kDa, Protein from Xenopus laevis)
- Primase (Complex from Xenopus laevis)
- Rna primer (2 kDa, RNA from synthetic construct)
- Polymerase alpha-primase with a dna initiation substrate (110 kDa, Complex)
- Polymerase alpha (Complex from Xenopus laevis)
- Magnesium ion (24 Da, Ligand)
- Dna initiation substrate (20 kDa, Complex from synthetic construct)
- 2'-deoxyguanosine-5'-triphosphate (507 Da, Ligand)
- Iron/sulfur cluster (351 Da, Ligand)
Partial DNA elongation subcomplex of Xenopus laevis DNA polymerase alpha-primase
Resolution: 3.5 Å
EM Method: Single-particle
Fitted PDBs: 8g9n
Q-score: 0.415
Mullins EA, Salay LE, Durie CL, Bradley NP, Jackman JE, Ohi MD, Chazin WJ, Eichman BF
Nat Struct Mol Biol (2024) [ Pubmed: 38491139 DOI: doi:10.1038/s41594-024-01227-4 ]
- Dna template (15 kDa, DNA from synthetic construct)
- Dna polymerase alpha catalytic subunit (129 kDa, Protein from Xenopus laevis)
- Primase (Complex from Xenopus laevis)
- Rna-dna primer (6 kDa, DNA from synthetic construct)
- Polymerase alpha (Complex from Xenopus laevis)
- Magnesium ion (24 Da, Ligand)
- Polymerase alpha-primase with a dna elongation substrate (20 kDa, Complex)
- Dna elongation substrate (Complex from synthetic construct)
- 2'-deoxyguanosine-5'-triphosphate (507 Da, Ligand)
Complete DNA elongation subcomplex of Xenopus laevis DNA polymerase alpha-primase
Resolution: 4.4 Å
EM Method: Single-particle
Fitted PDBs: 8g9o
Q-score: 0.275
Mullins EA, Salay LE, Durie CL, Bradley NP, Jackman JE, Ohi MD, Chazin WJ, Eichman BF
Nat Struct Mol Biol (2024) [ Pubmed: 38491139 DOI: doi:10.1038/s41594-024-01227-4 ]
- Dna template (15 kDa, DNA from synthetic construct)
- Dna polymerase alpha catalytic subunit (129 kDa, Protein from Xenopus laevis)
- Dna primase large subunit (59 kDa, Protein from Xenopus laevis)
- Iron/sulfur cluster (351 Da, Ligand)
- Primase (Complex from Xenopus laevis)
- Rna-dna primer (6 kDa, Macromolecule from synthetic construct)
- Polymerase alpha (Complex from Xenopus laevis)
- Dna elongation substrate (20 kDa, Complex from synthetic construct)
- Magnesium ion (24 Da, Ligand)
- 2'-deoxyguanosine-5'-triphosphate (507 Da, Ligand)
- Polymerase alpha-primase with a dna elongation substrate (110 kDa, Complex)
In situ structure of polymerase complex of mammalian reovirus in the elongation state
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 7yed
Q-score: 0.498
Bao K, Zhang X, Li D, Sun W, Sun Z, Wang J, Zhu P
PNAS (2022) 119 pp. e2203054119-e2203054119 [ Pubmed: 36469786 DOI: doi:10.1073/pnas.2203054119 ]
- Mammalian orthoreovirus 3 (Complex)
- Rna (36-mer) (11 kDa, RNA from Mammalian orthoreovirus 3)
- Rna-directed rna polymerase (142 kDa, Protein from Mammalian orthoreovirus 3)
- Mu-2 protein (83 kDa, Protein from Mammalian orthoreovirus 3)
- Uridine 5'-triphosphate (484 Da, Ligand)
- Mammalian orthoreovirus 3 (Virus from Mammalian orthoreovirus 3)
- Lambda-2 protein (143 kDa, Protein from Mammalian orthoreovirus 3)
- Rna (Complex)
- Magnesium ion (24 Da, Ligand)
- S-adenosylmethionine (398 Da, Ligand)
- Rna helicase (141 kDa, Protein from Mammalian orthoreovirus 3)
- Rna (5'-r(p*up*up*up*ap*ap*ap*ap*ap*up*up*up*up*ap*ap*ap*ap*up*ap*ap*up*u)-3') (6 kDa, RNA from Mammalian orthoreovirus 3)
- Zinc ion (65 Da, Ligand)
- Guanosine-5'-triphosphate (523 Da, Ligand)
Bombyx mori R2 retrotransposon initiating target-primed reverse transcription
Resolution: 3.08 Å
EM Method: Single-particle
Fitted PDBs: 8gh6
Q-score: 0.49
Wilkinson ME, Frangieh CJ, Macrae RK, Zhang F
Science (2023) 380 pp. 301-308 [ DOI: doi:10.1126/science.adg7883 Pubmed: 37023171 ]
- Thymidine-5'-triphosphate (482 Da, Ligand)
- 28s dna bottom strand, 5' side (priming strand) (7 kDa, DNA from Bombyx mori)
- Reverse transcriptase-like protein (123 kDa, Protein from Bombyx mori)
- Zinc ion (65 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- 28s dna top strand (21 kDa, DNA from Bombyx mori)
- Bombyx mori r2 retrotransposon initiating target-primed reverse transcription (230 kDa, Complex from Bombyx mori)
- 28s dna bottom strand, 3' side (21 kDa, DNA from Bombyx mori)
- R2bm 3'utr rna (81 kDa, RNA from Bombyx mori)
Porcine epidemic diarrhea virus core polymerase complex
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8g6r
Q-score: 0.435
Anderson TK, Hoferle PJ, Lee KW, Coon JJ, Kirchdoerfer RN
bioRxiv (2023) [ DOI: doi:10.1101/2023.03.15.532841 Pubmed: 36993498 ]
- Core replication complex of porcine epidemic diarrhea virus composed of nsp12, nsp7, nsp8, and a short rna substrate (185 kDa, Complex from Porcine epidemic diarrhea virus)
- Nsp7 (6 kDa, Protein from Porcine epidemic diarrhea virus)
- Nsp8 (12 kDa, Protein from Porcine epidemic diarrhea virus)
- Zinc ion (65 Da, Ligand)
- Rna (5'-r(p*ap*ap*gp*ap*ap*gp*cp*up*ap*up*up*ap*ap*ap*ap*up*cp*ap*cp*a)-3') (6 kDa, RNA from synthetic construct)
- Rna (5'-r(p*gp*gp*up*up*gp*up*gp*ap*up*up*up*up*ap*ap*up*ap*gp*cp*up*u)-3') (6 kDa, RNA from synthetic construct)
- Nsp12 (104 kDa, Protein from Porcine epidemic diarrhea virus)