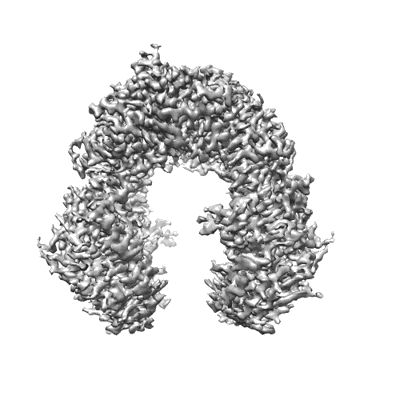Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
Structure of Gabija AB complex
Resolution: 2.79 Å
EM Method: Single-particle
Fitted PDBs: 8tjy
Q-score: 0.534
Yang XY, Shen Z, Xie J, Greenwald J, Marathe I, Lin Q, Xie WJ, Wysocki VH, Fu TM
Nat Struct Mol Biol (2024) [ Pubmed: 38627580 DOI: doi:10.1038/s41594-024-01283-w ]
- Tetramer of gabija protein a (Complex from Bacillus cereus)
- Gabija protein gajb (57 kDa, Protein from Bacillus cereus)
- Endonuclease gaja (67 kDa, Protein from Bacillus cereus)
Structure of Gabija AB complex
Resolution: 3.23 Å
EM Method: Single-particle
Fitted PDBs: 8tk0
Q-score: 0.496
Yang XY, Shen Z, Xie J, Greenwald J, Marathe I, Lin Q, Xie WJ, Wysocki VH, Fu TM
Nat Struct Mol Biol (2024) [ Pubmed: 38627580 DOI: doi:10.1038/s41594-024-01283-w ]
- Tetramer of gabija protein a (Complex from Bacillus cereus)
- Endonuclease gaja (67 kDa, Protein from Bacillus cereus)
Structure of Gabija AB complex 1
Resolution: 2.98 Å
EM Method: Single-particle
Fitted PDBs: 8tk1
Q-score: 0.453
Yang XY, Shen Z, Xie J, Greenwald J, Marathe I, Lin Q, Xie WJ, Wysocki VH, Fu TM
Nat Struct Mol Biol (2024) [ Pubmed: 38627580 DOI: doi:10.1038/s41594-024-01283-w ]
- Tetramer of gabija protein ab (4:4) (Complex from Bacillus cereus)
- Gabija protein gajb (57 kDa, Protein from Bacillus cereus)
- Endonuclease gaja (67 kDa, Protein from Bacillus cereus)
E. coli SIR2-HerA complex (hexamer HerA bound with dodecamer Sir2)
Resolution: 2.83 Å
EM Method: Single-particle
Fitted PDBs: 8su9
Q-score: 0.562
Shen Z, Lin Q, Yang XY, Fosuah E, Fu TM
Mol Cell (2023) 83 pp. 4586-4599.e5 [ DOI: doi:10.1016/j.molcel.2023.11.007 Pubmed: 38096827 ]
- E. coli sir2-hera complex (Complex from Escherichia coli)
- [(2r,3s,4r,5r)-5-(6-aminopurin-9-yl)-3,4-dihydroxy-oxolan-2-yl]methyl [hydroxy-[[(2r,3s,4r,5s)-3,4,5-trihydroxyoxolan-2-yl]methoxy]phosphoryl] hydrogen phosphate (559 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Sir2-like domain-containing protein (46 kDa, Protein from Escherichia coli)
- Nucleoside triphosphate hydrolase (68 kDa, Protein from Escherichia coli)
E. coli SIR2-HerA complex (dodecamer SIR2 bound 4 protomers of HerA)
Resolution: 3.15 Å
EM Method: Single-particle
Fitted PDBs: 8suw
Q-score: 0.439
Shen Z, Lin Q, Yang XY, Fosuah E, Fu TM
Mol Cell (2023) 83 pp. 4586-4599.e5 [ DOI: doi:10.1016/j.molcel.2023.11.007 Pubmed: 38096827 ]
- [(2r,3s,4r,5r)-5-(6-aminopurin-9-yl)-3,4-dihydroxy-oxolan-2-yl]methyl [hydroxy-[[(2r,3s,4r,5s)-3,4,5-trihydroxyoxolan-2-yl]methoxy]phosphoryl] hydrogen phosphate (559 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Sir2-like domain-containing protein (46 kDa, Protein from Escherichia coli)
- Nucleoside triphosphate hydrolase (68 kDa, Protein from Escherichia coli)
- Sir2-hera complex(dodecomaer sir2 bound with four protomers of hera) (Complex from Escherichia coli)
Structure of E. coli PtuA hexamer
Resolution: 2.93 Å
EM Method: Single-particle
Fitted PDBs: 8sux
Q-score: 0.5
Li Y, Shen Z, Zhang M, Yang XY, Cleary SP, Xie J, Marathe IA, Kostelic M, Greenwald J, Rish AD, Wysocki VH, Chen C, Chen Q, Fu TM, Yu Y
Nat Struct Mol Biol (2024) 31 pp. 413-423 [ Pubmed: 38177683 DOI: doi:10.1038/s41594-023-01172-8 ]
- Ptua (Complex from Escherichia coli)
- Ptua (53 kDa, Protein from Escherichia coli)
- Adenosine-5'-triphosphate (507 Da, Ligand)
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8ee4
Q-score: 0.576
Li Y, Shen Z, Zhang M, Yang XY, Cleary SP, Xie J, Marathe IA, Kostelic M, Greenwald J, Rish AD, Wysocki VH, Chen C, Chen Q, Fu TM, Yu Y
Nat Struct Mol Biol (2024) 31 pp. 413-423 [ Pubmed: 38177683 DOI: doi:10.1038/s41594-023-01172-8 ]
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Ptua (53 kDa, Protein from Escherichia coli)
- 6 ptua form as a complex (Complex from Escherichia coli)
Structure of focused PtuA(dimer) and PtuB(monomer) complex
Resolution: 2.72 Å
EM Method: Single-particle
Fitted PDBs: 8ee7
Q-score: 0.609
Li Y, Shen Z, Zhang M, Yang XY, Cleary SP, Xie J, Marathe IA, Kostelic M, Greenwald J, Rish AD, Wysocki VH, Chen C, Chen Q, Fu TM, Yu Y
Nat Struct Mol Biol (2024) 31 pp. 413-423 [ Pubmed: 38177683 DOI: doi:10.1038/s41594-023-01172-8 ]
- Ptub (28 kDa, Protein from Escherichia coli)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Ptua (53 kDa, Protein from Escherichia coli)
- 6 ptua form as a complex (Complex from Escherichia coli)
Structure of E.coli Septu (PtuAB) complex
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8eea
Q-score: 0.603
Li Y, Shen Z, Zhang M, Yang XY, Cleary SP, Xie J, Marathe IA, Kostelic M, Greenwald J, Rish AD, Wysocki VH, Chen C, Chen Q, Fu TM, Yu Y
Nat Struct Mol Biol (2024) 31 pp. 413-423 [ Pubmed: 38177683 DOI: doi:10.1038/s41594-023-01172-8 ]
- Ptub (28 kDa, Protein from Escherichia coli)
- 6 ptua and 2 ptub form as a complex (Complex from Escherichia coli)
- Ptua (53 kDa, Protein from Escherichia coli)
- Adenosine-5'-triphosphate (507 Da, Ligand)
E. coli SIR2-HerA complex (dodecamer SIR2 pentamer HerA)
Resolution: 2.89 Å
EM Method: Single-particle
Fitted PDBs: 8sub
Q-score: 0.515
Shen Z, Lin Q, Yang XY, Fosuah E, Fu TM
Mol Cell (2023) 83 pp. 4586-4599.e5 [ DOI: doi:10.1016/j.molcel.2023.11.007 Pubmed: 38096827 ]
- [(2r,3s,4r,5r)-5-(6-aminopurin-9-yl)-3,4-dihydroxy-oxolan-2-yl]methyl [hydroxy-[[(2r,3s,4r,5s)-3,4,5-trihydroxyoxolan-2-yl]methoxy]phosphoryl] hydrogen phosphate (559 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Sir2-hera complex 2(12:5) (Complex from Escherichia coli)
- Magnesium ion (24 Da, Ligand)
- Sir2-like domain-containing protein (46 kDa, Protein from Escherichia coli)
- Nucleoside triphosphate hydrolase (68 kDa, Protein from Escherichia coli)