Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
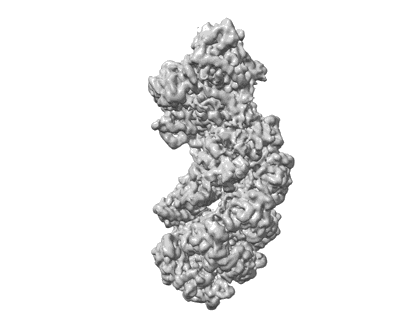
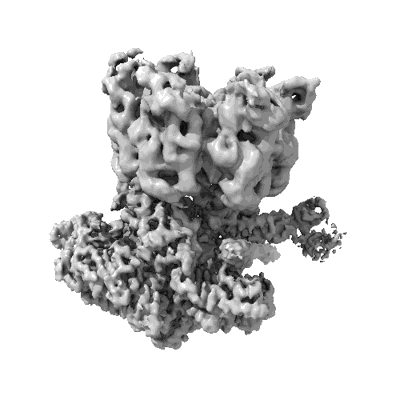
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 8ip0
Q-score: 0.326
Lu M, Yu C, Zhang Y, Ju W, Ye Z, Hua C, Mao J, Hu C, Yang Z, Xiao Y
Nat Commun (2024) 15 pp. 4126-4126 [ Pubmed: 38750051 DOI: doi:10.1038/s41467-024-48598-2 ]
- Dna (5'-d(p*ap*ap*ap*ap*ap*ap*ap*ap*ap*a)-3') (3 kDa, DNA from synthetic construct)
- Dna (41-mer) (12 kDa, DNA from synthetic construct)
- Cryo-em structure of synechocystis sp. pcc6714 cascade bound to a pam-containing dsdna target presented as full r-loop formed state at 3.6 angstrom resolution (Complex from Synechocystis sp. PCC 6714)
- Rna (44-mer) (13 kDa, RNA from synthetic construct)
- Dna (5'-d(p*tp*cp*cp*ap*tp*gp*tp*tp*tp*ap*tp*cp*ap*cp*c)-3') (4 kDa, DNA from synthetic construct)
- Type i-myxan crispr-associated protein cmx8 (63 kDa, Protein from Synechocystis sp. PCC 6714)
- Crispr associated protein cas11b (14 kDa, Protein from Synechocystis sp. PCC 6714)
- Fruiting body developmental protein r-like protein (33 kDa, Protein from Synechocystis sp. PCC 6714)
- Type i-myxan crispr-associated protein cas5/cmx5/devs (26 kDa, Protein from Synechocystis sp. PCC 6714)
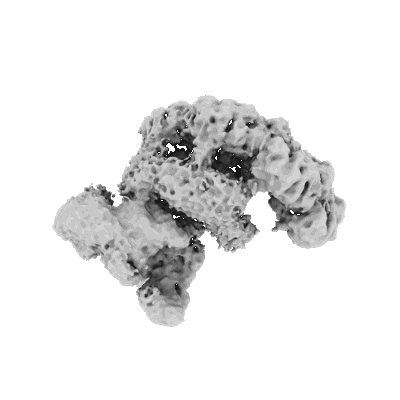
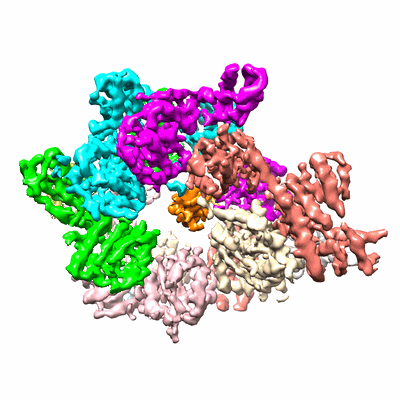
Cryo-EM structure of the bacterial replication origin opening basal unwinding system
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 8btg
Q-score: 0.298
Pelliciari S, Bodet-Lefevre S, Fenyk S, Stevens D, Winterhalter C, Schramm FD, Pintar S, Burnham DR, Merces G, Richardson TT, Tashiro Y, Hubbard J, Yardimci H, Ilangovan A, Murray H
Nat Commun (2023) 14 pp. 8339- [ DOI: doi:10.1093/nar/gkac1060 Pubmed: 38097584 ]
- Basal unwinding complex (Complex from Bacillus subtilis)
- Chromosomal replication initiator protein dnaa (50 kDa, Protein from Bacillus subtilis)
- Magnesium ion (24 Da, Ligand)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Dna (5'-d(p*ap*cp*tp*tp*ap*tp*cp*cp*ap*cp*ap*ap*ap*tp*cp*cp*ap*cp*ap*gp*gp*cp*cp*c)-3') (7 kDa, DNA from DNA molecule)
- Dna (41-mer) (12 kDa, DNA from DNA molecule)
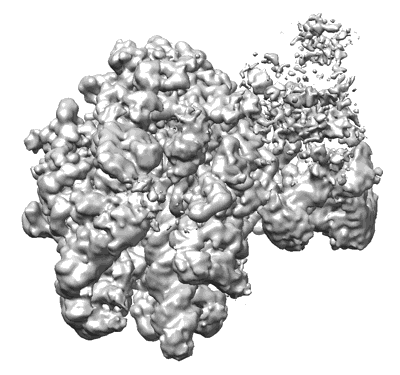
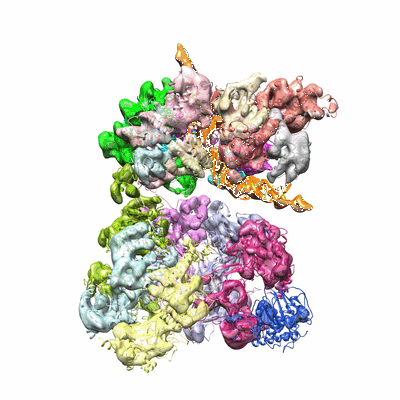
Structural insights into human co-transcriptional capping - structure 3
Resolution: 3.8 Å
EM Method: Single-particle
Fitted PDBs: 8p4c
Q-score: 0.297
Garg G, Dienemann C, Farnung L, Schwarz J, Linden A, Urlaub H, Cramer P
Mol Cell (2023) 83 pp. 2464-2477.e5 [ Pubmed: 37369200 DOI: doi:10.1016/j.molcel.2023.06.002 ]
- Dna-directed rna polymerase ii subunit rpb11-a (13 kDa, Protein from Sus scrofa)
- Dna-directed rna polymerase subunit beta (134 kDa, Protein from Sus scrofa)
- Dna-directed rna polymerase ii subunit rpb9 (14 kDa, Protein from Sus scrofa)
- Transcription elongation factor spt4 (13 kDa, Protein from Homo sapiens)
- Mrna-capping enzyme (68 kDa, Protein from Homo sapiens)
- Zinc ion (65 Da, Ligand)
- Dna-directed rna polymerases i, ii, and iii subunit rpabc5 (7 kDa, Protein from Sus scrofa)
- Dna-directed rna polymerase ii subunit e (24 kDa, Protein from Sus scrofa)
- Dna-directed rna polymerase ii subunit rpb7 (19 kDa, Protein from Sus scrofa)
- Rna polymerase ii subunit k (7 kDa, Protein from Sus scrofa)
- Rna polymerase ii subunit d (16 kDa, Protein from Sus scrofa)
- Dna-directed rna polymerase ii subunit rpb3 (31 kDa, Protein from Sus scrofa)
- Rna (5'-r(p*cp*gp*gp*ap*gp*ap*gp*gp*gp*ap*ap*cp*cp*cp*ap*cp*u)-3') (5 kDa, RNA from unidentified)
- Dna-directed rna polymerases i, ii, and iii subunit rpabc3 (17 kDa, Protein from Sus scrofa)
- Dna (32-mer) (9 kDa, DNA from unidentified)
- Dna-directed rna polymerase ii subunit f (14 kDa, Protein from Sus scrofa)
- Transcription elongation factor spt5 (121 kDa, Protein from Homo sapiens)
- Pol ii - tpase complex (Complex from Homo sapiens)
- Magnesium ion (24 Da, Ligand)
- Dna (41-mer) (12 kDa, DNA from unidentified)
- Dna-directed rna polymerase ii subunit rpb1 (217 kDa, Protein from Sus scrofa)
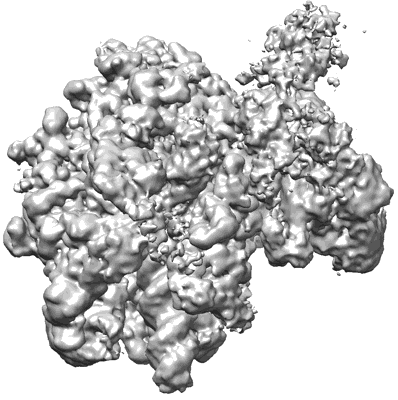
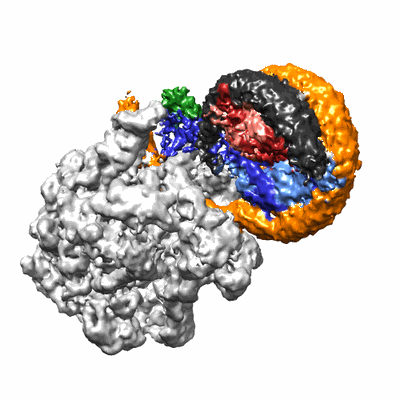
Structural insights into human co-transcriptional capping - structure 4
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 8p4d
Q-score: 0.348
Garg G, Dienemann C, Farnung L, Schwarz J, Linden A, Urlaub H, Cramer P
Mol Cell (2023) 83 pp. 2464-2477.e5 [ Pubmed: 37369200 DOI: doi:10.1016/j.molcel.2023.06.002 ]
- Dna-directed rna polymerase ii subunit rpb11-a (13 kDa, Protein from Sus scrofa)
- Dna-directed rna polymerase subunit beta (134 kDa, Protein from Sus scrofa)
- Dna-directed rna polymerase ii subunit rpb9 (14 kDa, Protein from Sus scrofa)
- Transcription elongation factor spt4 (13 kDa, Protein from Homo sapiens)
- Mrna-capping enzyme (68 kDa, Protein from Homo sapiens)
- Zinc ion (65 Da, Ligand)
- Dna-directed rna polymerases i, ii, and iii subunit rpabc5 (7 kDa, Protein from Sus scrofa)
- Dna-directed rna polymerase ii subunit e (24 kDa, Protein from Sus scrofa)
- Dna-directed rna polymerase ii subunit rpb7 (19 kDa, Protein from Sus scrofa)
- Rna polymerase ii subunit k (7 kDa, Protein from Sus scrofa)
- Rna polymerase ii subunit d (16 kDa, Protein from Sus scrofa)
- Dna-directed rna polymerase ii subunit rpb3 (31 kDa, Protein from Sus scrofa)
- Rna (5'-r(p*ap*cp*cp*gp*gp*ap*gp*ap*gp*gp*gp*ap*ap*cp*cp*cp*ap*cp*u)-3') (6 kDa, RNA from unidentified)
- Dna-directed rna polymerases i, ii, and iii subunit rpabc3 (17 kDa, Protein from Sus scrofa)
- Dna (32-mer) (9 kDa, DNA from unidentified)
- Dna-directed rna polymerase ii subunit f (14 kDa, Protein from Sus scrofa)
- Transcription elongation factor spt5 (121 kDa, Protein from Homo sapiens)
- Pol ii - tpase complex (Complex from Homo sapiens)
- Dna (41-mer) (12 kDa, DNA from unidentified)
- Dna-directed rna polymerase ii subunit rpb1 (217 kDa, Protein from Sus scrofa)
Thermus thermophilus Rho-engaged RNAP elongation complex
Resolution: 4.0 Å
EM Method: Single-particle
Fitted PDBs: 8hsr
Q-score: 0.317
Murayama Y, Ehara H, Aoki M, Goto M, Yokoyama T, Sekine SI
Sci Adv (2023) 9 pp. eade7093-eade7093 [ DOI: doi:10.1126/sciadv.ade7093 Pubmed: 36753546 ]
- Dna-directed rna polymerase subunit beta' (171 kDa, Protein from Thermus thermophilus HB8)
- Beryllium trifluoride ion (66 Da, Ligand)
- Rho-engaged rna polymerase elongation complex (Complex from Thermus thermophilus HB8)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Transcription termination factor rho (47 kDa, Protein from Thermus thermophilus HB8)
- Magnesium ion (24 Da, Ligand)
- Dna (31-mer) (54 kDa, DNA from synthetic construct)
- Dna (41-mer) (57 kDa, DNA from synthetic construct)
- Dna-directed rna polymerase subunit omega (11 kDa, Protein from Thermus thermophilus HB8)
- Rna (29-mer) (39 kDa, RNA from synthetic construct)
- Dna-directed rna polymerase subunit beta (125 kDa, Protein from Thermus thermophilus HB8)
- Zinc ion (65 Da, Ligand)
- Dna-directed rna polymerase subunit alpha (35 kDa, Protein from Thermus thermophilus HB8)
Cryo-EM map of semi-attached mutant OCCM-DNA complex (ORC-Cdc6-Cdt1-Mcm2-7 with Mcm6 WHD truncation)
Resolution: 4.3 Å
EM Method: Single-particle
Fitted PDBs: 6wgc
Q-score: 0.301
Yuan Z, Schneider S, Dodd T, Riera A, Bai L, Yan C, Magdalou I, Ivanov I, Stillman B, Li H, Speck C
PNAS (2020) 117 pp. 17747-17756 [ DOI: doi:10.1073/pnas.2006231117 Pubmed: 32669428 ]
- Origin recognition complex subunit 5 (55 kDa, Protein from Saccharomyces cerevisiae)
- Phosphothiophosphoric acid-adenylate ester (523 Da, Ligand)
- Origin recognition complex subunit 1 (104 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm3 (107 kDa, Protein from Saccharomyces cerevisiae)
- Dna (41-mer) (12 kDa, DNA from Saccharomyces cerevisiae)
- Origin recognition complex subunit 3 (72 kDa, Protein from Saccharomyces cerevisiae)
- Orc-cdc6-mcm3-mcm7-dsdna (Complex from Saccharomyces cerevisiae)
- Origin recognition complex subunit 2 (71 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 6 (50 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm7 (95 kDa, Protein from Saccharomyces cerevisiae)
- Dna (41-mer) (12 kDa, DNA from Saccharomyces cerevisiae)
- Cell division control protein 6 (58 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 4 (60 kDa, Protein from Saccharomyces cerevisiae)
Cryo-EM map of pre-insertion mutant OCCM-DNA complex (ORC-Cdc6-Cdt1-Mcm2-7 with Mcm6 WHD truncation)
Resolution: 8.1 Å
EM Method: Single-particle
Fitted PDBs: 6wgg
Q-score: 0.104
Yuan Z, Schneider S, Dodd T, Riera A, Bai L, Yan C, Magdalou I, Ivanov I, Stillman B, Li H, Speck C
PNAS (2020) 117 pp. 17747-17756 [ DOI: doi:10.1073/pnas.2006231117 Pubmed: 32669428 ]
- Origin recognition complex subunit 5 (55 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm2 (98 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 1 (104 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm4 (105 kDa, Protein from Saccharomyces cerevisiae)
- Phosphothiophosphoric acid-adenylate ester (523 Da, Ligand)
- Dna (41-mer) (12 kDa, DNA from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm3 (107 kDa, Protein from Saccharomyces cerevisiae)
- Minichromosome maintenance protein 5 (86 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 3 (72 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 2 (71 kDa, Protein from Saccharomyces cerevisiae)
- Cell division cycle protein cdt1 (68 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 6 (50 kDa, Protein from Saccharomyces cerevisiae)
- Dna (41-mer) (12 kDa, DNA from Saccharomyces cerevisiae)
- Orc-cdc6-mcm2-7-dsdna (Complex from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm7 (95 kDa, Protein from Saccharomyces cerevisiae)
- Cell division control protein 6 (58 kDa, Protein from Saccharomyces cerevisiae)
- Dna replication licensing factor mcm6 (113 kDa, Protein from Saccharomyces cerevisiae)
- Origin recognition complex subunit 4 (60 kDa, Protein from Saccharomyces cerevisiae)
Resolution: 4.7 Å
EM Method: Single-particle
Fitted PDBs: 6j50
Q-score: 0.195
Ehara H, Kujirai T, Fujino Y, Shirouzu M, Kurumizaka H, Sekine SI
Science (2019) 363 pp. 744-747 [ DOI: doi:10.1126/science.aav8912 Pubmed: 30733384 ]
- Dna (41-mer) (12 kDa, DNA from synthetic construct)
- Dna-directed rna polymerase subunit beta (139 kDa, Protein from Komagataella phaffii (strain GS115 / ATCC 20864))
- Histone h4 (11 kDa, Protein from Homo sapiens)
- Dna (198-mer) (61 kDa, DNA from synthetic construct)
- Rna (5'-r(p*gp*cp*cp*up*gp*gp*up*gp*up*cp*up*up*gp*gp*gp*u)-3') (5 kDa, RNA from synthetic construct)
- Zinc ion (65 Da, Ligand)
- Histone h2a type 1-b/e (14 kDa, Protein from Homo sapiens)
- Rna polymerase ii third largest subunit b44, part of central core (34 kDa, Protein from Komagataella phaffii (strain GS115 / ATCC 20864))
- Rna polymerase ii subunit b32 (20 kDa, Protein from Komagataella phaffii (strain GS115 / ATCC 20864))
- Dna (41-mer) (12 kDa, DNA from synthetic construct)
- Protein that forms a complex with spt4p (101 kDa, Protein from Komagataella phaffii (strain GS115 / ATCC 20864))
- Dna (198-mer) (60 kDa, DNA from synthetic construct)
- Histone h3.3 (15 kDa, Protein from Homo sapiens)
- Rna polymerase subunit abc10-beta, common to rna polymerases i, ii, and iii (8 kDa, Protein from Komagataella phaffii (strain GS115 / ATCC 20864))
- Rna polymerase subunit abc27, common to rna polymerases i, ii, and iii (24 kDa, Protein from Komagataella phaffii (strain GS115 / ATCC 20864))
- Dna-directed rna polymerase subunit (13 kDa, Protein from Komagataella phaffii (strain ATCC 76273 / CBS 7435 / CECT 11047 / NRRL Y-11430 / Wegner 21-1))
- Rna polymerase subunit abc23, common to rna polymerases i, ii, and iii (17 kDa, Protein from Komagataella phaffii (strain GS115 / ATCC 20864))
- Rna polymerase ii subunit (18 kDa, Protein from Komagataella phaffii (strain GS115 / ATCC 20864))
- Dna-directed rna polymerase subunit (194 kDa, Protein from Komagataella phaffii (strain GS115 / ATCC 20864))
- Histone h2b type 1-j (14 kDa, Protein from Homo sapiens)
- Rna polymerase subunit abc10-alpha (7 kDa, Protein from Komagataella phaffii (strain ATCC 76273 / CBS 7435 / CECT 11047 / NRRL Y-11430 / Wegner 21-1))
- Magnesium ion (24 Da, Ligand)
- Rna polymerase ii elongation complex bound with spt4/5 and foreign dna, stalled at shl(-1) of the nucleosome (tilted conformation) (Complex from Komagataella phaffii)
- Transcription elongation factor spt4 (12 kDa, Protein from Komagataella phaffii (strain GS115 / ATCC 20864))
- Dna-directed rna polymerases i, ii, and iii subunit rpabc3 (16 kDa, Protein from Komagataella phaffii (strain GS115 / ATCC 20864))
- Rna polymerase ii subunit b12.5 (13 kDa, Protein from Komagataella phaffii (strain GS115 / ATCC 20864))
Resolution: 3.17 Å
EM Method: Single-particle
Fitted PDBs: 6cg0
Q-score: 0.452
Kim MS, Chuenchor W, Chen X, Cui Y, Zhang X, Zhou ZH, Gellert M, Yang W
Mol Cell (2018) 70 pp. 358-370.e4 [ DOI: doi:10.1016/j.molcel.2018.03.008 Pubmed: 29628308 ]
- Dna (46-mer) (14 kDa, DNA from Escherichia coli K-12)
- Dna (30-mer) (9 kDa, DNA from Escherichia coli K-12)
- V(d)j recombination-activating protein 1 (88 kDa, Protein from Mus musculus)
- Rag1/2 in complex with nicked dnas (Complex from Mus musculus)
- Dna (5'-d(*gp*ap*tp*cp*tp*gp*gp*cp*cp*tp*gp*tp*cp*tp*tp*a)-3') (4 kDa, DNA from Homo sapiens)
- Calcium ion (40 Da, Ligand)
- Dna (60-mer) (18 kDa, DNA from Escherichia coli K-12)
- High mobility group protein b1 (14 kDa, Protein from Homo sapiens)
- Dna (41-mer) (12 kDa, DNA from Escherichia coli K-12)
- Dna (5'-d(p*cp*tp*gp*gp*ap*tp*cp*tp*gp*gp*cp*cp*tp*gp*tp*cp*tp*tp*a)-3') (5 kDa, DNA from Escherichia coli K-12)
- Zinc ion (65 Da, Ligand)
- V(d)j recombination-activating protein 2 (58 kDa, Protein from Mus musculus)
Cryo-EM structure of mouse RAG1/2 HFC complex containing partial HMGB1 linker(3.9 A)
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 6cij
Q-score: 0.409
Kim MS, Chuenchor W, Chen X, Cui Y, Zhang X, Zhou ZH, Gellert M, Yang W
Mol Cell (2018) 70 pp. 358-370.e4 [ DOI: doi:10.1016/j.molcel.2018.03.008 Pubmed: 29628308 ]
- Dna (60-mer) (18 kDa, DNA from Homo sapiens)
- Rag1/2 in complex with nicked dnas (Complex from Mus musculus)
- High mobility group protein b1 (18 kDa, Protein from Homo sapiens)
- Dna (5'-d(*gp*ap*tp*cp*tp*gp*gp*cp*cp*tp*gp*tp*cp*tp*tp*a)-3') (4 kDa, DNA from Homo sapiens)
- Dna (30-mer) (9 kDa, DNA from Homo sapiens)
- Calcium ion (40 Da, Ligand)
- V(d)j recombination-activating protein 1 (88 kDa, Protein from Mus musculus)
- Dna (41-mer) (12 kDa, DNA from Homo sapiens)
- Dna (5'-d(p*cp*tp*gp*gp*ap*tp*cp*tp*gp*gp*cp*cp*tp*gp*tp*cp*tp*tp*a)-3') (6 kDa, DNA from Homo sapiens)
- Dna (46-mer) (14 kDa, DNA from Homo sapiens)
- Zinc ion (65 Da, Ligand)
- V(d)j recombination-activating protein 2 (58 kDa, Protein from Mus musculus)