Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
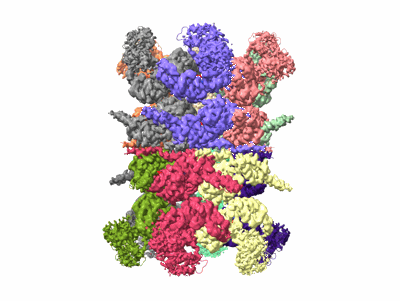
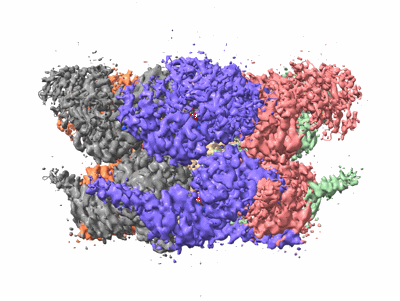
Human p97 double hexamer conformer II with ATPgammaS bound
Resolution: 3.32 Å
EM Method: Single-particle
Fitted PDBs: 7vcs
Q-score: 0.401
Gao H, Li F, Ji Z, Shi Z, Li Y, Yu H
Cell Discov (2022) 8 pp. 19-19 [ DOI: doi:10.1038/s41421-022-00379-1 Pubmed: 35190543 ]
- Transitional endoplasmic reticulum atpase (90 kDa, Protein from Homo sapiens)
- Magnesium ion (24 Da, Ligand)
- Phosphothiophosphoric acid-adenylate ester (523 Da, Ligand)
- Human p97 double hexamer conformer ii with atpgammas bound (1 MDa, Complex from Homo sapiens)
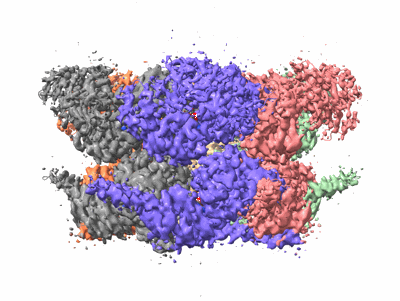
Human p97 single hexamer conformer III with D1-ATPgammaS and D2-ADP bound
Resolution: 3.21 Å
EM Method: Single-particle
Fitted PDBs: 7vct
Q-score: 0.452
Gao H, Li F, Ji Z, Shi Z, Li Y, Yu H
Cell Discov (2022) 8 pp. 19-19 [ DOI: doi:10.1038/s41421-022-00379-1 Pubmed: 35190543 ]
- Phosphothiophosphoric acid-adenylate ester (523 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Transitional endoplasmic reticulum atpase (90 kDa, Protein from Homo sapiens)
- Magnesium ion (24 Da, Ligand)
- Human p97 single hexamer conformer iii with d1-atpgammas and d2-adp bound (1 MDa, Complex from Homo sapiens)
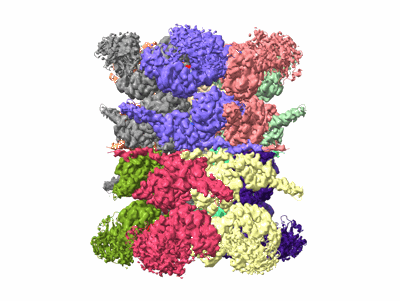
Human p97 double hexamer conformer I with D1-ATPgammaS and D2-ADP bound
Resolution: 3.15 Å
EM Method: Single-particle
Fitted PDBs: 7vcu
Q-score: 0.414
Gao H, Li F, Ji Z, Shi Z, Li Y, Yu H
Cell Discov (2022) 8 pp. 19-19 [ DOI: doi:10.1038/s41421-022-00379-1 Pubmed: 35190543 ]
- Phosphothiophosphoric acid-adenylate ester (523 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Transitional endoplasmic reticulum atpase (90 kDa, Protein from Homo sapiens)
- Magnesium ion (24 Da, Ligand)
- Human p97 double hexamer conformer i with d1-atpgammas and d2-adp bound (1 MDa, Complex from Homo sapiens)
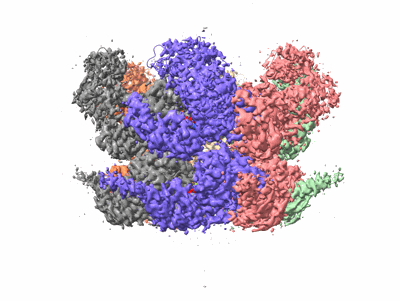
Human p97 single hexamer conformer I with ATPgammaS bound
Resolution: 3.21 Å
EM Method: Single-particle
Fitted PDBs: 7vcv
Q-score: 0.463
Gao H, Li F, Ji Z, Shi Z, Li Y, Yu H
Cell Discov (2022) 8 pp. 19-19 [ DOI: doi:10.1038/s41421-022-00379-1 Pubmed: 35190543 ]
- Transitional endoplasmic reticulum atpase (90 kDa, Protein from Homo sapiens)
- Magnesium ion (24 Da, Ligand)
- Phosphothiophosphoric acid-adenylate ester (523 Da, Ligand)
- Human p97 double hexamer conformer ii with atpgammas bound (1 MDa, Complex from Homo sapiens)
Human p97 single hexamer conformer II with ATPgammaS bound
Resolution: 3.24 Å
EM Method: Single-particle
Fitted PDBs: 7vcx
Q-score: 0.493
Gao H, Li F, Ji Z, Shi Z, Li Y, Yu H
Cell Discov (2022) 8 pp. 19-19 [ DOI: doi:10.1038/s41421-022-00379-1 Pubmed: 35190543 ]
- Transitional endoplasmic reticulum atpase (90 kDa, Protein from Homo sapiens)
- Magnesium ion (24 Da, Ligand)
- Phosphothiophosphoric acid-adenylate ester (523 Da, Ligand)
- Human p97 double hexamer conformer ii with atpgammas bound (1 MDa, Complex from Homo sapiens)
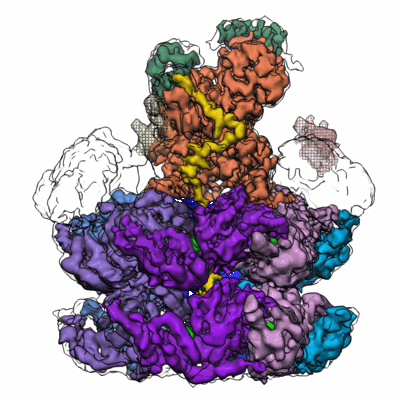
Cdc48-Ufd1/Npl4 complex processing poly-ubiquitinated substrate in the presence of ATP
Resolution: 3.9 Å
EM Method: Single-particle
Fitted PDBs: 6oa9
Q-score: 0.282
Twomey EC, Ji Z, Wales TE, Bodnar NO, Ficarro SB, Marto JA, Engen JR, Rapoport TA
Science (2019) 365 [ Pubmed: 31249135 DOI: doi:10.1126/science.aax1033 ]
- Nuclear protein localization protein 4 (65 kDa, Protein from Saccharomyces cerevisiae)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Ubiquitin (8 kDa, Protein from Saccharomyces cerevisiae)
- Cdc48-npl4 complex processing poly-ubiquitinated substrate in the presence of atp (Complex from Saccharomyces cerevisiae)
- Zinc ion (65 Da, Ligand)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Cell division control protein 48 (92 kDa, Protein from Saccharomyces cerevisiae)
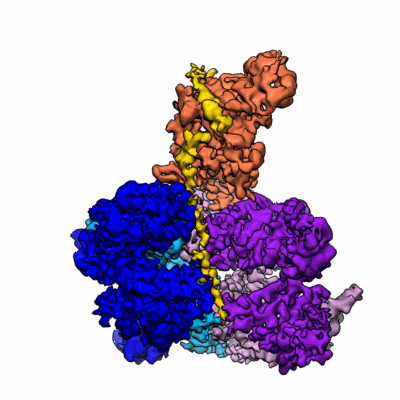
Cdc48-Ufd1/Npl4 complex processing poly-ubiquitinated substrate in the presence of ADP-BeFx, state 1
Resolution: 4.1 Å
EM Method: Single-particle
Fitted PDBs: 6oaa
Q-score: 0.295
Twomey EC, Ji Z, Wales TE, Bodnar NO, Ficarro SB, Marto JA, Engen JR, Rapoport TA
Science (2019) 365 [ Pubmed: 31249135 DOI: doi:10.1126/science.aax1033 ]
- Beryllium trifluoride ion (66 Da, Ligand)
- Nuclear protein localization protein 4 (65 kDa, Protein from Saccharomyces cerevisiae)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Ubiquitin (8 kDa, Protein from Saccharomyces cerevisiae)
- Cell division control protein 48 (92 kDa, Protein from Saccharomyces cerevisiae S288C)
- Cdc48-npl4 complex processing poly-ubiquitinated substrate in the presence of adp-befx, state 1 (Complex from Saccharomyces cerevisiae)
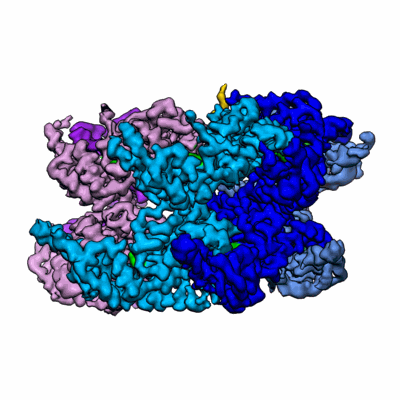
Cdc48-Ufd1/Npl4 complex processing poly-ubiquitinated substrate in the presence of ADP-BeFx, state 2
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 6oab
Q-score: 0.306
Twomey EC, Ji Z, Wales TE, Bodnar NO, Ficarro SB, Marto JA, Engen JR, Rapoport TA
Science (2019) 365 [ Pubmed: 31249135 DOI: doi:10.1126/science.aax1033 ]
- Beryllium trifluoride ion (66 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Cdc48-npl4 complex processing poly-ubiquitinated substrate in the presence of adp-befx, state 2 (Complex from Saccharomyces cerevisiae)
- Cell division control protein 48 (92 kDa, Protein from Saccharomyces cerevisiae)
- Poly(alanine) substrate (1 kDa, Protein from Saccharomyces cerevisiae)