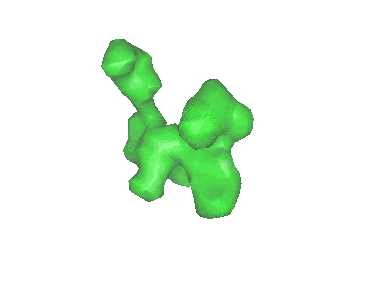Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
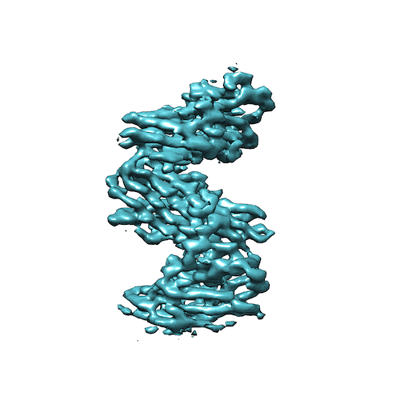
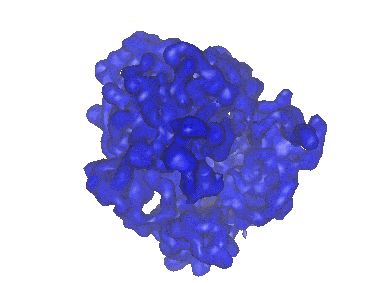
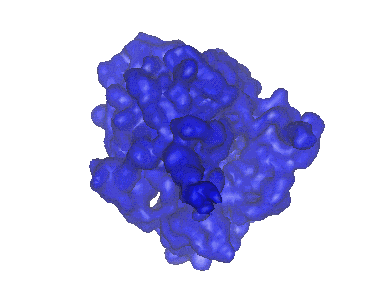
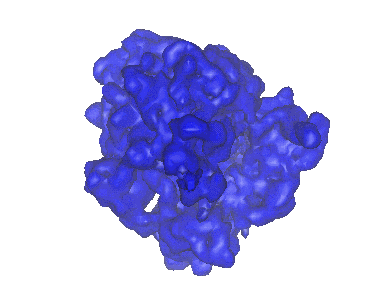
Cryo-EM structure of archaic chaperone-usher Csu pilus of Acinetobacter baumannii
Resolution: 3.45 Å
EM Method: Helical reconstruction
Fitted PDBs: 7zl4
Q-score: 0.412
Pakharukova N, Malmi H, Tuittila M, Dahlberg T, Ghosal D, Chang YW, Myint SL, Paavilainen S, Knight SD, Lamminmaki U, Uhlin BE, Andersson M, Jensen G, Zavialov AV
Nature (2022) 609 pp. 335-340 [ Pubmed: 35853476 DOI: doi:10.1038/s41586-022-05095-0 ]
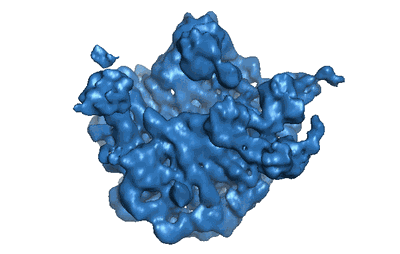
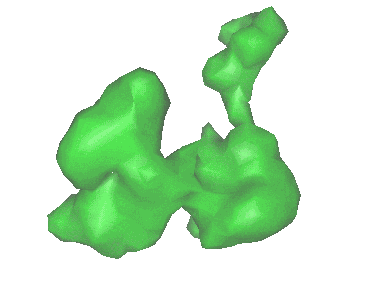
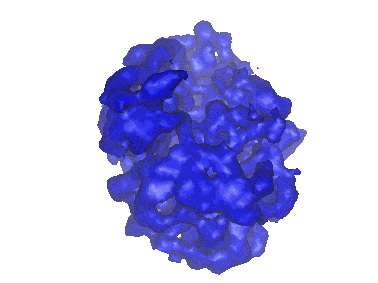
Resolution: 9.1 Å
EM Method: Single-particle
Fitted PDBs: 2rdo
Q-score: 0.039
Gao N, Zavialov AV, Ehrenberg M, Frank J
J Mol Biol (2007) 374 pp. 1345-1358 [ DOI: doi:10.1016/j.jmb.2007.10.021 Pubmed: 17996252 ]
- 50s subunit with ef-g and rrf bound (Sample)
- Ef-g (Protein from Escherichia coli)
- 50s subunit (Complex from Escherichia coli)
- Rrf (Protein from Escherichia coli)
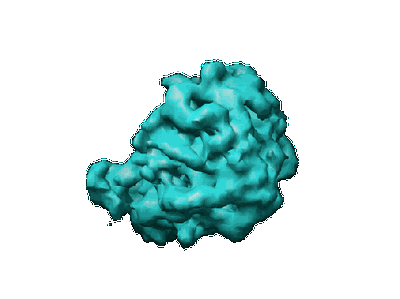
Mechanism for the disassembly of the posttermination complex inferred from cryo-EM studies.
Resolution: 15.5 Å
EM Method: Single-particle
Fitted PDBs: 1zn0, 1zn1
Q-score: -0.009
Gao N, Zavialov AV, Li W, Sengupta J, Valle M, Gursky RP, Ehrenberg M, Frank J
Mol Cell (2005) 18 pp. 663-674 [ DOI: doi:10.1016/j.molcel.2005.05.005 Pubmed: 15949441 ]
- 50s subunit (Complex from Escherichia coli)
- Ef-g (Protein from Escherichia coli)
- 50s.ef-g.gdpnp.rrf complex from e.coli (Sample)
- Rrf (Protein from Escherichia coli)
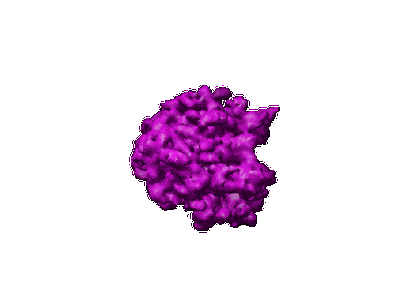
Mechanism for the disassembly of the posttermination complex inferred from cryo-EM studies.
Resolution: 14.1 Å
EM Method: Single-particle
Fitted PDBs: 1zn0, 1zn1
Q-score: 0.0285
Gao N, Zavialov AV, Li W, Sengupta J, Valle M, Gursky RP, Ehrenberg M, Frank J
Mol Cell (2005) 18 pp. 663-674 [ DOI: doi:10.1016/j.molcel.2005.05.005 Pubmed: 15949441 ]
- Rrf (Protein)
- Mrna (Macromolecule)
- 70s.trna-ile.mrna.puromycin.rrf from e.coli ribosome (Sample)
- 70s (Complex)
- Trna-ile (RNA from Escherichia coli)
A cryo-electron microscopic study of ribosome-bound termination factor RF2.
Resolution: 11.3 Å
EM Method: Single-particle
Fitted PDBs: 1mi6, 1mvr
Q-score: 0.013
Rawat UB, Zavialov AV, Sengupta J, Valle M, Grassucci RA, Linde J, Vestergaard B, Ehrenberg M, Frank J
Nature (2003) 421 pp. 87-90 [ Pubmed: 12511960 DOI: doi:10.1038/nature01224 ]
- Mfti-mrna (Ligand from Escherichia coli)
- E.coli 70s ribosome (2 MDa, Sample)
- E-trna (Ligand from Escherichia coli)
- E.coli 70s ribosome (2 MDa, Complex from Escherichia coli)
- Peptidyl p-trna (Ligand from Escherichia coli)
A cryo-electron microscopic study of ribosome-bound termination factor RF2.
Resolution: 12.9 Å
EM Method: Single-particle
Fitted PDBs: 1mi6, 1mvr
Q-score: 0.046
Rawat UB, Zavialov AV, Sengupta J, Valle M, Grassucci RA, Linde J, Vestergaard B, Ehrenberg M, Frank J
Nature (2003) 421 pp. 87-90 [ Pubmed: 12511960 DOI: doi:10.1038/nature01224 ]
- Mfti-mrna (Ligand from Escherichia coli)
- Rf2 (ggq) wild type (Ligand from Escherichia coli)
- E-trna (Ligand from Escherichia coli)
- P-trna (Ligand from Escherichia coli)
- E.coli 70s ribosome (2 MDa, Complex from Escherichia coli)
- E.coli 70s ribosome-rf2(wild type) complex (2 MDa, Sample)
A cryo-electron microscopic study of ribosome-bound termination factor RF2.
Resolution: 10.9 Å
EM Method: Single-particle
Fitted PDBs: 1mi6, 1mvr
Q-score: 0.054
Rawat UB, Zavialov AV, Sengupta J, Valle M, Grassucci RA, Linde J, Vestergaard B, Ehrenberg M, Frank J
Nature (2003) 421 pp. 87-90 [ Pubmed: 12511960 DOI: doi:10.1038/nature01224 ]
- Mfti-mrna (Ligand from Escherichia coli)
- E.coli 70s ribosome (Complex from Escherichia coli)
- E-trna (Ligand from Escherichia coli)
- E.coli 70s ribosome-rf2(mutant) complex (2 MDa, Sample)
- P-trna (Ligand from Escherichia coli)
- Rf2 (gaq) mutant (Ligand from Escherichia coli)
A cryo-electron microscopic study of ribosome-bound termination factor RF2.
Resolution: 10.9 Å
EM Method: Single-particle
Fitted PDBs: 1mi6, 1mvr
Q-score: 0.027
Rawat UB, Zavialov AV, Sengupta J, Valle M, Grassucci RA, Linde J, Vestergaard B, Ehrenberg M, Frank J
Nature (2003) 421 pp. 87-90 [ Pubmed: 12511960 DOI: doi:10.1038/nature01224 ]
A cryo-electron microscopic study of ribosome-bound termination factor RF2.
Resolution: 12.9 Å
EM Method: Single-particle
Fitted PDBs: 1mi6, 1mvr
Q-score: 0.0335
Rawat UB, Zavialov AV, Sengupta J, Valle M, Grassucci RA, Linde J, Vestergaard B, Ehrenberg M, Frank J
Nature (2003) 421 pp. 87-90 [ Pubmed: 12511960 DOI: doi:10.1038/nature01224 ]
Structure of the Escherichia coli ribosomal termination complex with release factor 2.
Resolution: 14.0 Å
EM Method: Single-particle
Fitted PDBs: 1ml5
Q-score: 0.013
Klaholz BP, Pape T, Zavialov AV, Myasnikov AG, Orlova EV, Vestergaard B, Ehrenberg M, van Heel M
Nature (2003) 421 pp. 90-94 [ Pubmed: 12511961 DOI: doi:10.1038/nature01225 ]
- Release factor 2 (40 kDa, Protein from Escherichia coli BL21(DE3))
- Ribosome (2 MDa, Complex from Escherichia coli)
- Release factor rf2 bound to e. coli ribosomes (2 MDa, Sample)