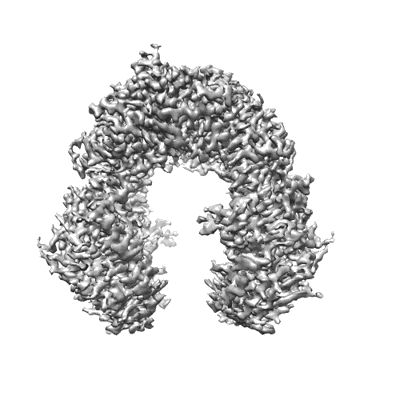Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research. We appreciate your responses prior to closing of the survey on Monday 17th June.
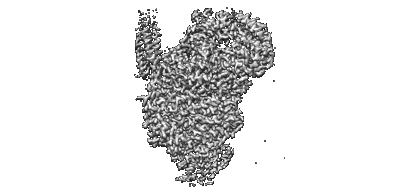
Cryo-EM structure of the human parainfluenza virus hPIV3 L-P polymerase in dimeric form
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 8kdb
Q-score: 0.577
Xie J, Ouizougun-Oubari M, Wang L, Zhai G, Wu D, Lin Z, Wang M, Ludeke B, Yan X, Nilsson T, Gao L, Huang X, Fearns R, Chen S
Nat Commun (2024) 15 pp. 3163-3163 [ DOI: doi:10.1038/s41467-024-47470-7 Pubmed: 38605025 ]
- Phosphoprotein (68 kDa, Protein from Human respirovirus 3)
- Human parainfluenza virus hpiv3 l-p polymerase in dimeric form (Complex from Human respirovirus 3)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase l (260 kDa, Protein from Human respirovirus 3)
- Zinc ion (65 Da, Ligand)
Cryo-EM structure of the human parainfluenza virus hPIV3 L-P polymerase in monomeric form
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8kdc
Q-score: 0.506
Xie J, Ouizougun-Oubari M, Wang L, Zhai G, Wu D, Lin Z, Wang M, Ludeke B, Yan X, Nilsson T, Gao L, Huang X, Fearns R, Chen S
Nat Commun (2024) 15 pp. 3163-3163 [ DOI: doi:10.1038/s41467-024-47470-7 Pubmed: 38605025 ]
- Human parainfluenza virus hpiv3 l-p polymerase in monomeric form (Complex from Human respirovirus 3)
- Phosphoprotein (68 kDa, Protein from Human respirovirus 3)
- Magnesium ion (24 Da, Ligand)
- Rna-directed rna polymerase l (260 kDa, Protein from Human respirovirus 3)
- Zinc ion (65 Da, Ligand)
Structure of Gabija AB complex
Resolution: 2.79 Å
EM Method: Single-particle
Fitted PDBs: 8tjy
Q-score: 0.534
Yang XY, Shen Z, Xie J, Greenwald J, Marathe I, Lin Q, Xie WJ, Wysocki VH, Fu TM
Nat Struct Mol Biol (2024) [ Pubmed: 38627580 DOI: doi:10.1038/s41594-024-01283-w ]
- Tetramer of gabija protein a (Complex from Bacillus cereus)
- Gabija protein gajb (57 kDa, Protein from Bacillus cereus)
- Endonuclease gaja (67 kDa, Protein from Bacillus cereus)
Structure of Gabija AB complex
Resolution: 3.23 Å
EM Method: Single-particle
Fitted PDBs: 8tk0
Q-score: 0.496
Yang XY, Shen Z, Xie J, Greenwald J, Marathe I, Lin Q, Xie WJ, Wysocki VH, Fu TM
Nat Struct Mol Biol (2024) [ Pubmed: 38627580 DOI: doi:10.1038/s41594-024-01283-w ]
- Tetramer of gabija protein a (Complex from Bacillus cereus)
- Endonuclease gaja (67 kDa, Protein from Bacillus cereus)
Structure of Gabija AB complex 1
Resolution: 2.98 Å
EM Method: Single-particle
Fitted PDBs: 8tk1
Q-score: 0.453
Yang XY, Shen Z, Xie J, Greenwald J, Marathe I, Lin Q, Xie WJ, Wysocki VH, Fu TM
Nat Struct Mol Biol (2024) [ Pubmed: 38627580 DOI: doi:10.1038/s41594-024-01283-w ]
- Tetramer of gabija protein ab (4:4) (Complex from Bacillus cereus)
- Gabija protein gajb (57 kDa, Protein from Bacillus cereus)
- Endonuclease gaja (67 kDa, Protein from Bacillus cereus)
Structure of E. coli PtuA hexamer
Resolution: 2.93 Å
EM Method: Single-particle
Fitted PDBs: 8sux
Q-score: 0.5
Li Y, Shen Z, Zhang M, Yang XY, Cleary SP, Xie J, Marathe IA, Kostelic M, Greenwald J, Rish AD, Wysocki VH, Chen C, Chen Q, Fu TM, Yu Y
Nat Struct Mol Biol (2024) 31 pp. 413-423 [ Pubmed: 38177683 DOI: doi:10.1038/s41594-023-01172-8 ]
- Ptua (Complex from Escherichia coli)
- Ptua (53 kDa, Protein from Escherichia coli)
- Adenosine-5'-triphosphate (507 Da, Ligand)
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8ee4
Q-score: 0.576
Li Y, Shen Z, Zhang M, Yang XY, Cleary SP, Xie J, Marathe IA, Kostelic M, Greenwald J, Rish AD, Wysocki VH, Chen C, Chen Q, Fu TM, Yu Y
Nat Struct Mol Biol (2024) 31 pp. 413-423 [ Pubmed: 38177683 DOI: doi:10.1038/s41594-023-01172-8 ]
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Ptua (53 kDa, Protein from Escherichia coli)
- 6 ptua form as a complex (Complex from Escherichia coli)
Structure of focused PtuA(dimer) and PtuB(monomer) complex
Resolution: 2.72 Å
EM Method: Single-particle
Fitted PDBs: 8ee7
Q-score: 0.609
Li Y, Shen Z, Zhang M, Yang XY, Cleary SP, Xie J, Marathe IA, Kostelic M, Greenwald J, Rish AD, Wysocki VH, Chen C, Chen Q, Fu TM, Yu Y
Nat Struct Mol Biol (2024) 31 pp. 413-423 [ Pubmed: 38177683 DOI: doi:10.1038/s41594-023-01172-8 ]
- Ptub (28 kDa, Protein from Escherichia coli)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Ptua (53 kDa, Protein from Escherichia coli)
- 6 ptua form as a complex (Complex from Escherichia coli)
Structure of E.coli Septu (PtuAB) complex
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8eea
Q-score: 0.603
Li Y, Shen Z, Zhang M, Yang XY, Cleary SP, Xie J, Marathe IA, Kostelic M, Greenwald J, Rish AD, Wysocki VH, Chen C, Chen Q, Fu TM, Yu Y
Nat Struct Mol Biol (2024) 31 pp. 413-423 [ Pubmed: 38177683 DOI: doi:10.1038/s41594-023-01172-8 ]
- Ptub (28 kDa, Protein from Escherichia coli)
- 6 ptua and 2 ptub form as a complex (Complex from Escherichia coli)
- Ptua (53 kDa, Protein from Escherichia coli)
- Adenosine-5'-triphosphate (507 Da, Ligand)
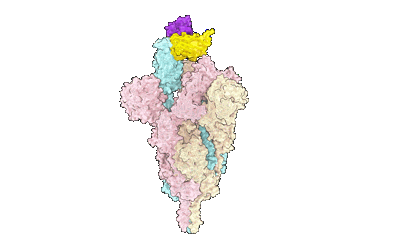
SARS-CoV-2 Omicron variant spike in complex with Fab 9A8 (State 1)
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7wt7
Q-score: 0.385
Liu S, Jia Z, Nie J, Liang Z, Xie J, Wang L, Zhang L, Wang X, Wang Y, Huang W
Cell Discov (2022) 8 pp. 81-81 [ Pubmed: 35977939 DOI: doi:10.1038/s41421-022-00443-w ]
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Sars-cov-2 omicron variant spike in complex with fab 9a8 (state 1) (Complex from Severe acute respiratory syndrome coronavirus 2)
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Light chain of fab 9a8 (11 kDa, Protein from Homo sapiens)
- Heavy chain of fab 9a8 (12 kDa, Protein from Homo sapiens)