Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
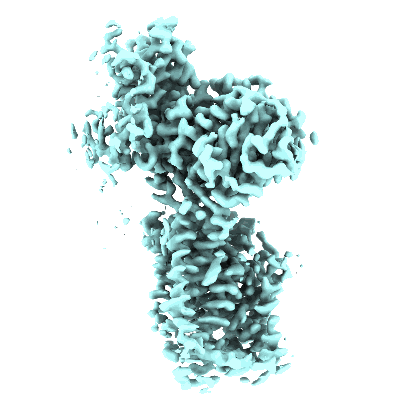
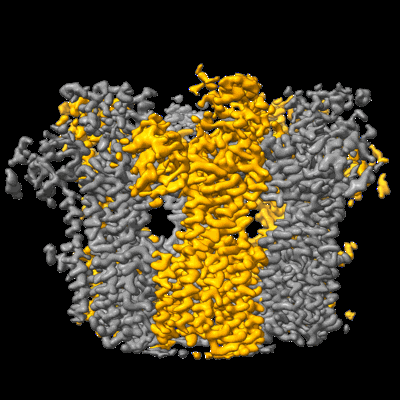
Resolution: 3.05 Å
EM Method: Single-particle
Fitted PDBs: 7nf6
Q-score: 0.558
Lee Y, Wiriyasermkul P, Kongpracha P, Moriyama S, Mills DJ, Kuhlbrandt W, Nagamori S
Nat Commun (2022) 13 pp. 2708-2708 [ DOI: doi:10.1038/s41467-022-30293-9 Pubmed: 35577790 ]
- Calcium ion (40 Da, Ligand)
- 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (760 Da, Ligand)
- Cholesterol (386 Da, Ligand)
- B(0,+)-type amino acid transporter 1 (56 kDa, Protein from Ovis aries)
- Neutral and basic amino acid transport protein rbat (78 kDa, Protein from Ovis aries)
- B0,+at-rbat heterodimer (135 MDa, Complex from Ovis aries)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
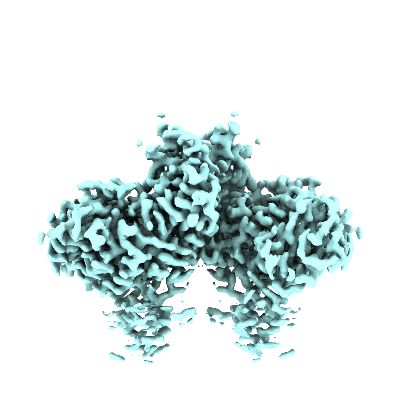
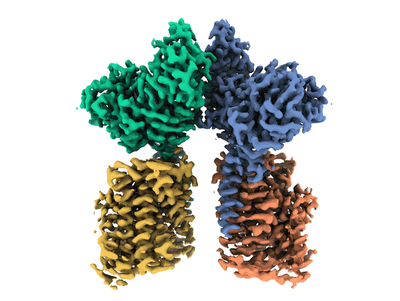
Ovine rBAT ectodomain homodimer, C2-symmetry
Resolution: 2.68 Å
EM Method: Single-particle
Fitted PDBs: 7nf7
Q-score: 0.642
Lee Y, Wiriyasermkul P, Kongpracha P, Moriyama S, Mills DJ, Kuhlbrandt W, Nagamori S
Nat Commun (2022) 13 pp. 2708-2708 [ DOI: doi:10.1038/s41467-022-30293-9 Pubmed: 35577790 ]
- Ovine rbat ectodomain homodimer (79 MDa, Complex from Ovis aries)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Neutral and basic amino acid transport protein rbat (78 kDa, Protein from Ovis aries)
- Calcium ion (40 Da, Ligand)
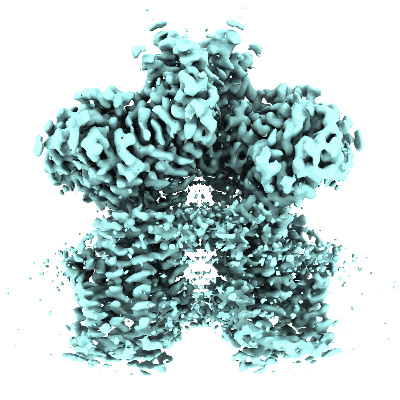
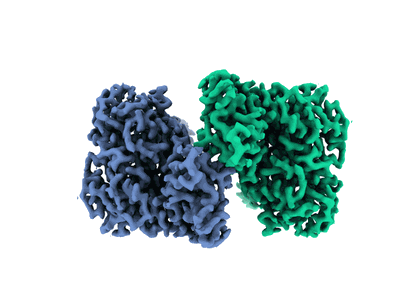
Ovine (b0,+AT-rBAT)2 hetero-tetramer, consensus refinement, C2-symmetry
Resolution: 2.83 Å
EM Method: Single-particle
Fitted PDBs: 7nf8
Q-score: 0.496
Lee Y, Wiriyasermkul P, Kongpracha P, Moriyama S, Mills DJ, Kuhlbrandt W, Nagamori S
Nat Commun (2022) 13 pp. 2708-2708 [ DOI: doi:10.1038/s41467-022-30293-9 Pubmed: 35577790 ]
- Calcium ion (40 Da, Ligand)
- 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (760 Da, Ligand)
- Cholesterol (386 Da, Ligand)
- (b0,+at-rbat) hetero-tetramer (270 MDa, Complex from Ovis aries)
- B(0,+)-type amino acid transporter 1 (56 kDa, Protein from Ovis aries)
- Neutral and basic amino acid transport protein rbat (78 kDa, Protein from Ovis aries)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
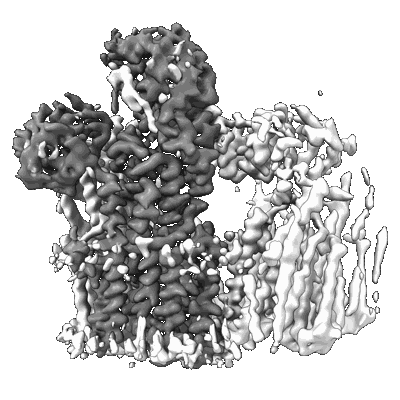
Structure of the fungal plasma membrane proton pump Pma1 in its auto-inhibited state - monomer unit
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7nxf
Q-score: 0.439
Heit S, Geurts MMG, Murphy BJ, Corey RA, Mills DJ, Kuhlbrandt W, Bublitz M
Sci Adv (2021) 7 pp. eabj5255-eabj5255 [ Pubmed: 34757782 DOI: doi:10.1126/sciadv.abj5255 ]
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Monomeric subunit of the hexameric fungal plasma membrane proton pump in its auto-inhibited state (Cellular component from Neurospora crassa)
- Plasma membrane atpase (99 kDa, Protein from Neurospora crassa)
- Magnesium ion (24 Da, Ligand)
- Potassium ion (39 Da, Ligand)
Resolution: 3.26 Å
EM Method: Single-particle
Fitted PDBs: 7ny1
Q-score: 0.357
Heit S, Geurts MMG, Murphy BJ, Corey RA, Mills DJ, Kuhlbrandt W, Bublitz M
Sci Adv (2021) 7 pp. eabj5255-eabj5255 [ Pubmed: 34757782 DOI: doi:10.1126/sciadv.abj5255 ]
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Plasma membrane atpase (99 kDa, Protein from Neurospora crassa)
- Magnesium ion (24 Da, Ligand)
- Potassium ion (39 Da, Ligand)
- Hexameric assembly of the fungal plasma membrane proton pump in its auto-inhibited state (Complex from Neurospora crassa)
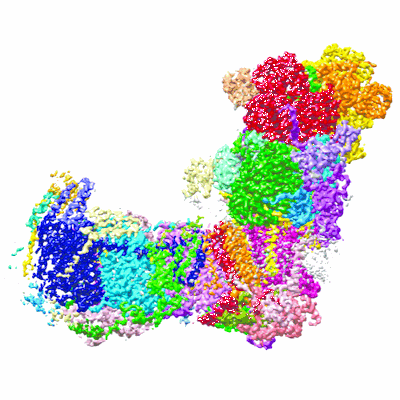
Cryo-EM structure of respiratory complex I under turnover
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7o6y
Q-score: 0.468
Parey K, Lasham J, Mills DJ, Djurabekova A, Haapanen O, Yoga EG, Xie H, Kuhlbrandt W, Sharma V, Vonck J, Zickermann V
Sci Adv (2021) 7 pp. eabj3221-eabj3221 [ Pubmed: 34767441 DOI: doi:10.1126/sciadv.abj3221 ]
- Subunit nuam of nadh:ubiquinone oxidoreductase (complex i) (79 kDa, Protein from Candida lipolytica)
- Subunit nugm of nadh:ubiquinone oxidoreductase (complex i) (32 kDa, Protein from Candida lipolytica)
- Subunit nuzm of nadh:ubiquinone oxidoreductase (complex i) (19 kDa, Protein from Yarrowia lipolytica)
- Subunit nipm of nadh:ubiquinone oxidoreductase (complex i) (10 kDa, Protein from Candida lipolytica)
- Subunit numm of protein nadh:ubiquinone oxidoreductase (complex i) (15 kDa, Protein from Candida lipolytica)
- Zinc ion (65 Da, Ligand)
- Subunit nebm of nadh:ubiquinone oxidoreductase (complex i) (8 kDa, Protein from Yarrowia lipolytica)
- Mitochondrial nadh:ubiquinone oxidoreductase (1 MDa, Complex from Yarrowia lipolytica)
- Subunit nb8m of nadh:ubiquinone oxidoreductase (complex i) (11 kDa, Protein from Candida lipolytica)
- Nadh-ubiquinone oxidoreductase chain 3 (14 kDa, Protein from Yarrowia lipolytica)
- Subunit nujm of nadh:ubiquinone oxidoreductase (complex i) (20 kDa, Protein from Yarrowia lipolytica)
- Subunit nb4m of protein nadh:ubiquinone oxidoreductase (complex i) (14 kDa, Protein from Candida lipolytica)
- Subunit niam of nadh:ubiquinone oxidoreductase (complex i) (17 kDa, Protein from Yarrowia lipolytica)
- 1-palmitoyl-2-linoleoyl-sn-glycero-3-phosphocholine (758 Da, Ligand)
- Cardiolipin (1 kDa, Ligand)
- Ubiquinone-9 (795 Da, Ligand)
- Subunit ni2m of nadh:ubiquinone oxidoreductase (complex i) (12 kDa, Protein from Candida lipolytica)
- Phosphatidylinositol (887 Da, Ligand)
- Subunit nuym of nadh:ubiquinone oxidoreductase (complex i) (18 kDa, Protein from Candida lipolytica)
- Subunit nu4m of nadh:ubiquinone oxidoreductase (complex i) (54 kDa, Protein from Yarrowia lipolytica)
- Subunit nuvm of nadh:ubiquinone oxidoreductase (complex i) (7 kDa, Protein from Yarrowia lipolytica)
- Nucm protein (52 kDa, Protein from Yarrowia lipolytica)
- Nadh-ubiquinone oxidoreductase chain 1 (38 kDa, Protein from Yarrowia lipolytica)
- Acyl carrier protein acpm2 of nadh:ubiquinone oxidoreductase (complex i) (14 kDa, Protein from Candida lipolytica)
- Flavin mononucleotide (456 Da, Ligand)
- Iron/sulfur cluster (351 Da, Ligand)
- Subunit nimm of nadh:ubiquinone oxidoreductase (complex i) (9 kDa, Protein from Candida lipolytica)
- Subunit nb5m of nadh:ubiquinone oxidoreductase (complex i) (10 kDa, Protein from Candida lipolytica)
- Subunit nesm of nadh:ubiquinone oxidoreductase (complex i) (28 kDa, Protein from Yarrowia lipolytica)
- Nadh dehydrogenase [ubiquinone] flavoprotein 1, mitochondrial (53 kDa, Protein from Candida lipolytica)
- 1,4-dihydronicotinamide adenine dinucleotide (665 Da, Ligand)
- Lauryl maltose neopentyl glycol (1 kDa, Ligand)
- Subunit nuxm of nadh:ubiquinone oxidoreductase (complex i) (18 kDa, Protein from Candida lipolytica)
- Acyl carrier protein acpm1 of nadh:ubiquinone oxidoreductase (complex i) (12 kDa, Protein from Candida lipolytica)
- Nadh dehydrogenase subunit 2 (53 kDa, Protein from Yarrowia lipolytica)
- Subunit nukm of nadh:ubiquinone oxidoreductase (complex i) (23 kDa, Protein from Candida lipolytica)
- Subunit nuem of nadh:ubiquinone oxidoreductase (complex i) (42 kDa, Protein from Yarrowia lipolytica)
- Subunit nuhm of nadh:ubiquinone oxidoreductase (complex i) (27 kDa, Protein from Candida lipolytica)
- 1,2-distearoyl-sn-glycerophosphoethanolamine (748 Da, Ligand)
- Subunit ni9m of protein nadh:ubiquinone oxidoreductase (complex i) (8 kDa, Protein from Yarrowia lipolytica)
- Subunit nuim of nadh:ubiquinone oxidoreductase (complex i) (25 kDa, Protein from Candida lipolytica)
- Subunit nupm of nadh:ubiquinone oxidoreductase (complex i) (19 kDa, Protein from Yarrowia lipolytica)
- Subunit nuum of nadh:ubiquinone oxidoreductase (complex i) (9 kDa, Protein from Yarrowia lipolytica)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Subunit nu5m of nadh:ubiquinone oxidoreductase (complex i) (73 kDa, Protein from Yarrowia lipolytica)
- Subunit nunm of nadh:ubiquinone oxidoreductase (complex i) (15 kDa, Protein from Candida lipolytica)
- Nadh-ubiquinone oxidoreductase chain 6 (20 kDa, Protein from Candida lipolytica)
- Subunit nufm of nadh:ubiquinone oxidoreductase (complex i) (16 kDa, Protein from Yarrowia lipolytica)
- Diundecyl phosphatidyl choline (622 Da, Ligand)
- S-[2-({n-[(2s)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-beta-alanyl}amino)ethyl] tetradecanethioate (568 Da, Ligand)
- Subunit nb2m of nadh:ubiquinone oxidoreductase (complex i) (6 kDa, Protein from Yarrowia lipolytica)
- Subunit nb6m of nadh:ubiquinone oxidoreductase (complex i) (14 kDa, Protein from Candida lipolytica)
- Subunit n7bm of nadh:ubiquinone oxidoreductase (complex i) (16 kDa, Protein from Yarrowia lipolytica)
- Subunit nidm of nadh:ubiquinone oxidoreductase (complex i) (11 kDa, Protein from Candida lipolytica)
- Subunit ni8m of nadh:ubiquinone oxidoreductase (complex i) (9 kDa, Protein from Candida lipolytica)
- Nadph dihydro-nicotinamide-adenine-dinucleotide phosphate (745 Da, Ligand)
- Nadh-ubiquinone oxidoreductase chain 4l (9 kDa, Protein from Yarrowia lipolytica)
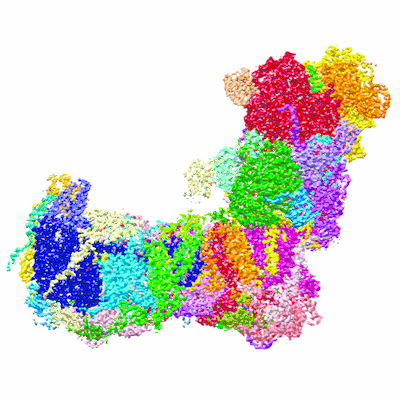
Cryo-EM structure of a respiratory complex I
Resolution: 2.4 Å
EM Method: Single-particle
Fitted PDBs: 7o71
Q-score: 0.644
Parey K, Lasham J, Mills DJ, Djurabekova A, Haapanen O, Yoga EG, Xie H, Kuhlbrandt W, Sharma V, Vonck J, Zickermann V
Sci Adv (2021) 7 pp. eabj3221-eabj3221 [ Pubmed: 34767441 DOI: doi:10.1126/sciadv.abj3221 ]
- Subunit nugm of nadh:ubiquinone oxidoreductase (complex i) (32 kDa, Protein from Candida lipolytica)
- Subunit nuzm of nadh:ubiquinone oxidoreductase (complex i) (19 kDa, Protein from Yarrowia lipolytica)
- Subunit nipm of nadh:ubiquinone oxidoreductase (complex i) (10 kDa, Protein from Candida lipolytica)
- Subunit numm of protein nadh:ubiquinone oxidoreductase (complex i) (15 kDa, Protein from Candida lipolytica)
- Nadh-ubiquinone oxidoreductase chain 4 (54 kDa, Protein from Yarrowia lipolytica)
- Subunit nunm of nadh:ubiquinone oxidoreductase (complex i) (13 kDa, Protein from Candida lipolytica)
- Zinc ion (65 Da, Ligand)
- Subunit nebm of nadh:ubiquinone oxidoreductase (complex i) (8 kDa, Protein from Yarrowia lipolytica)
- Mitochondrial nadh:ubiquinone oxidoreductase (1 MDa, Complex from Yarrowia lipolytica)
- Subunit nb8m of nadh:ubiquinone oxidoreductase (complex i) (11 kDa, Protein from Candida lipolytica)
- Nadh-ubiquinone oxidoreductase chain 3 (14 kDa, Protein from Yarrowia lipolytica)
- Subunit nujm of nadh:ubiquinone oxidoreductase (complex i) (20 kDa, Protein from Yarrowia lipolytica)
- Subunit nb4m of protein nadh:ubiquinone oxidoreductase (complex i) (14 kDa, Protein from Candida lipolytica)
- Subunit niam of nadh:ubiquinone oxidoreductase (complex i) (17 kDa, Protein from Yarrowia lipolytica)
- 1-palmitoyl-2-linoleoyl-sn-glycero-3-phosphocholine (758 Da, Ligand)
- Cardiolipin (1 kDa, Ligand)
- Subunit ni2m of nadh:ubiquinone oxidoreductase (complex i) (12 kDa, Protein from Candida lipolytica)
- Phosphatidylinositol (887 Da, Ligand)
- Subunit nuym of nadh:ubiquinone oxidoreductase (complex i) (18 kDa, Protein from Candida lipolytica)
- Subunit nuvm of nadh:ubiquinone oxidoreductase (complex i) (7 kDa, Protein from Yarrowia lipolytica)
- Nucm protein (52 kDa, Protein from Yarrowia lipolytica)
- Nadh-ubiquinone oxidoreductase chain 1 (38 kDa, Protein from Yarrowia lipolytica)
- Acyl carrier protein acpm2 of nadh:ubiquinone oxidoreductase (complex i) (14 kDa, Protein from Candida lipolytica)
- Flavin mononucleotide (456 Da, Ligand)
- Iron/sulfur cluster (351 Da, Ligand)
- Water (18 Da, Ligand)
- Subunit nimm of nadh:ubiquinone oxidoreductase (complex i) (9 kDa, Protein from Candida lipolytica)
- Subunit nb5m of nadh:ubiquinone oxidoreductase (complex i) (10 kDa, Protein from Candida lipolytica)
- Subunit nesm of nadh:ubiquinone oxidoreductase (complex i) (28 kDa, Protein from Yarrowia lipolytica)
- Nadh dehydrogenase [ubiquinone] flavoprotein 1, mitochondrial (53 kDa, Protein from Candida lipolytica)
- Lauryl maltose neopentyl glycol (1 kDa, Ligand)
- Subunit nuxm of nadh:ubiquinone oxidoreductase (complex i) (18 kDa, Protein from Candida lipolytica)
- Acyl carrier protein acpm1 of nadh:ubiquinone oxidoreductase (complex i) (12 kDa, Protein from Candida lipolytica)
- Nadh dehydrogenase subunit 2 (53 kDa, Protein from Yarrowia lipolytica)
- Subunit nukm of nadh:ubiquinone oxidoreductase (complex i) (23 kDa, Protein from Candida lipolytica)
- Nadh-ubiquinone oxidoreductase 78 kda subunit (79 kDa, Protein from Candida lipolytica)
- Nadh-ubiquinone oxidoreductase chain 5 (73 kDa, Protein from Yarrowia lipolytica)
- Subunit nuhm of nadh:ubiquinone oxidoreductase (complex i) (27 kDa, Protein from Candida lipolytica)
- 1,2-distearoyl-sn-glycerophosphoethanolamine (748 Da, Ligand)
- Subunit ni9m of protein nadh:ubiquinone oxidoreductase (complex i) (8 kDa, Protein from Yarrowia lipolytica)
- Subunit nuim of nadh:ubiquinone oxidoreductase (complex i) (25 kDa, Protein from Candida lipolytica)
- Subunit nupm of nadh:ubiquinone oxidoreductase (complex i) (19 kDa, Protein from Yarrowia lipolytica)
- Subunit nuum of nadh:ubiquinone oxidoreductase (complex i) (9 kDa, Protein from Yarrowia lipolytica)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Nadh-ubiquinone oxidoreductase chain 6 (20 kDa, Protein from Candida lipolytica)
- Subunit nufm of nadh:ubiquinone oxidoreductase (complex i) (16 kDa, Protein from Yarrowia lipolytica)
- Diundecyl phosphatidyl choline (622 Da, Ligand)
- S-[2-({n-[(2s)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-beta-alanyl}amino)ethyl] tetradecanethioate (568 Da, Ligand)
- Subunit nb2m of nadh:ubiquinone oxidoreductase (complex i) (6 kDa, Protein from Yarrowia lipolytica)
- Subunit nb6m of nadh:ubiquinone oxidoreductase (complex i) (14 kDa, Protein from Candida lipolytica)
- Subunit n7bm of nadh:ubiquinone oxidoreductase (complex i) (16 kDa, Protein from Yarrowia lipolytica)
- Nadh-ubiquinone oxidoreductase 40 kda subunit (42 kDa, Protein from Candida lipolytica)
- Subunit nidm of nadh:ubiquinone oxidoreductase (complex i) (11 kDa, Protein from Candida lipolytica)
- Subunit ni8m of nadh:ubiquinone oxidoreductase (complex i) (9 kDa, Protein from Candida lipolytica)
- Nadph dihydro-nicotinamide-adenine-dinucleotide phosphate (745 Da, Ligand)
- Nadh-ubiquinone oxidoreductase chain 4l (9 kDa, Protein from Yarrowia lipolytica)
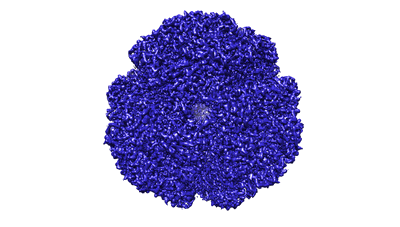
Helicobacter pylori urease with BME bound in the active site
Resolution: 2.4 Å
EM Method: Single-particle
Fitted PDBs: 6qsu
Q-score: 0.656
Cunha ES, Chen X, Sanz-Gaitero M, Mills DJ, Luecke H
Nat Commun (2021) 12 pp. 230-230 [ Pubmed: 33431861 DOI: doi:10.1038/s41467-020-20485-6 ]
- Urease subunit alpha (26 kDa, Protein from Helicobacter pylori)
- Nickel (ii) ion (58 Da, Ligand)
- Beta-mercaptoethanol (78 Da, Ligand)
- 1.1 mda helicobacter pylori urease (1 MDa, Complex from Helicobacter pylori)
- Water (18 Da, Ligand)
- Urease subunit beta (61 kDa, Protein from Helicobacter pylori)
Heterotetrameric structure of the rBAT-b(0,+)AT1 complex
Resolution: 2.9 Å
EM Method: Single-particle
Fitted PDBs: 6yup
Q-score: 0.491
Wu D, Grund TN, Welsch S, Mills DJ, Michel M, Safarian S, Michel H
PNAS (2020) 117 pp. 21281-21287 [ DOI: doi:10.1073/pnas.2008111117 Pubmed: 32817565 ]
- Calcium ion (40 Da, Ligand)
- B(0,+)-type amino acid transporter 1 (53 kDa, Protein from Homo sapiens)
- Heterotetrameric complex of rbat and b(0,+)at1 (264 MDa, Complex from Homo sapiens)
- Neutral and basic amino acid transport protein rbat (78 kDa, Protein from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
Homodimeric structure of the rBAT complex
Resolution: 2.8 Å
EM Method: Single-particle
Fitted PDBs: 6yuz
Q-score: 0.56
Wu D, Grund TN, Welsch S, Mills DJ, Michel M, Safarian S, Michel H
PNAS (2020) 117 pp. 21281-21287 [ DOI: doi:10.1073/pnas.2008111117 Pubmed: 32817565 ]
- Neutral and basic amino acid transport protein rbat (78 kDa, Protein from Homo sapiens)
- Homodimeric complex of rbat (157 MDa, Complex from Homo sapiens)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)
- Calcium ion (40 Da, Ligand)