Do data resources managed by EMBL-EBI and our collaborators make a difference to your work? Please take 10 minutes to fill in our annual user survey, and help us make the case for why sustaining open data resources is critical for life sciences research.
Johnson AG, Mayer ML, Schaefer SL, McNamara-Bordewick NK, Hummer G, Kranzusch PJ
Nature (2024) 628 pp. 657-663 [ DOI: 10.1038/s41586-024-07216-3 Pubmed: 38509367 ]
Image sets:
Structure of a bacterial gasdermin small oval pore assembly
Johnson AG, Mayer ML, Schaefer SL, McNamara-Bordewick NK, Hummer G, Kranzusch PJ
Nature (2024) 628 pp. 657-663 [ DOI: doi:10.1038/s41586-024-07216-3 Pubmed: 38509367 ]
Structure of a bacterial gasdermin medium oval pore assembly
Johnson AG, Mayer ML, Schaefer SL, McNamara-Bordewick NK, Hummer G, Kranzusch PJ
Nature (2024) 628 pp. 657-663 [ DOI: doi:10.1038/s41586-024-07216-3 Pubmed: 38509367 ]
Structure of a bacterial gasdermin large oval pore assembly
Johnson AG, Mayer ML, Schaefer SL, McNamara-Bordewick NK, Hummer G, Kranzusch PJ
Nature (2024) 628 pp. 657-663 [ DOI: doi:10.1038/s41586-024-07216-3 Pubmed: 38509367 ]
Structure of a bacterial gasdermin double pore assembly
Johnson AG, Mayer ML, Schaefer SL, McNamara-Bordewick NK, Hummer G, Kranzusch PJ
Nature (2024) 628 pp. 657-663 [ DOI: doi:10.1038/s41586-024-07216-3 Pubmed: 38509367 ]
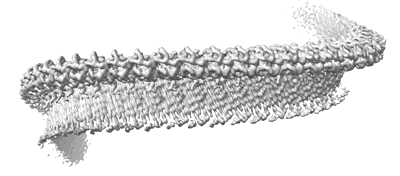
Structure of a bacterial gasdermin slinky-like oligomer from a heterogeneous sample
Johnson AG, Mayer ML, Schaefer SL, McNamara-Bordewick NK, Hummer G, Kranzusch PJ
Nature (2024) 628 pp. 657-663 [ DOI: doi:10.1038/s41586-024-07216-3 Pubmed: 38509367 ]
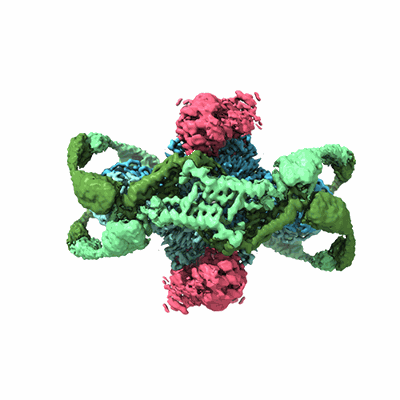
Structure of the phage immune evasion protein Gad1 bound to the Gabija GajAB complex
Resolution: 2.57 Å
EM Method: Single-particle
Fitted PDBs: 8u7i
Q-score: 0.491
Antine SP, Johnson AG, Mooney SE, Leavitt A, Mayer ML, Yirmiya E, Amitai G, Sorek R, Kranzusch PJ
Nature (2024) 625 pp. 360-365 [ DOI: doi:10.1038/s41586-023-06855-2 Pubmed: 37992757 ]
- Gabija gajab complex (Complex from Bacillus cereus VD045)
- Gabija protein gajb (57 kDa, Protein from Bacillus cereus VD045)
- Gad1 octamer (Complex from Bacillus phage phi3T)
- Phage phi3t gad1 bound to the bacillus cereus gabija gajab complex (280 kDa, Complex from Bacillus phage phi3T)
- Gabija anti-defense 1 (34 kDa, Protein from Bacillus phage phi3T)
- Endonuclease gaja (78 kDa, Protein from Bacillus cereus VD045)
Structure of a bacterial gasdermin slinky-like oligomer
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8sl0
Q-score: 0.444
Johnson AG, Mayer ML, Schaefer SL, McNamara-Bordewick NK, Hummer G, Kranzusch PJ
Nature (2024) 628 pp. 657-663 [ DOI: doi:10.1038/s41586-024-07216-3 Pubmed: 38509367 ]
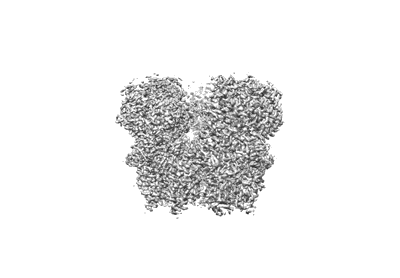
Structure of RdrA from Escherichia coli RADAR defense system
Resolution: 2.52 Å
EM Method: Single-particle
Fitted PDBs: 8fnt
Q-score: 0.486
Duncan-Lowey B, Tal N, Johnson AG, Rawson S, Mayer ML, Doron S, Millman A, Melamed S, Fedorenko T, Kacen A, Brandis A, Mehlman T, Amitai G, Sorek R, Kranzusch PJ
Cell (2023) 186 pp. 987- [ DOI: doi:10.1016/j.cell.2023.01.012 Pubmed: 36764290 ]
- E. coli rdra complex (Complex from Escherichia coli)
- Archaeal atpase (107 kDa, Protein from Escherichia coli)
Map of RdrA from Escherichia coli RADAR defense system in single-split conformation
Duncan-Lowey B, Tal N, Johnson AG, Rawson S, Mayer ML, Doron S, Millman A, Melamed S, Fedorenko T, Kacen A, Brandis A, Mehlman T, Amitai G, Sorek R, Kranzusch PJ
Cell (2023) 186 pp. 987-998.e15 [ DOI: doi:10.1016/j.cell.2023.01.012 Pubmed: 36764290 ]